BLAST program selection guide - Cromatina
BLAST program selection guide - Cromatina
BLAST program selection guide - Cromatina
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
<strong>BLAST</strong> <strong>program</strong> <strong>selection</strong> <strong>guide</strong><br />
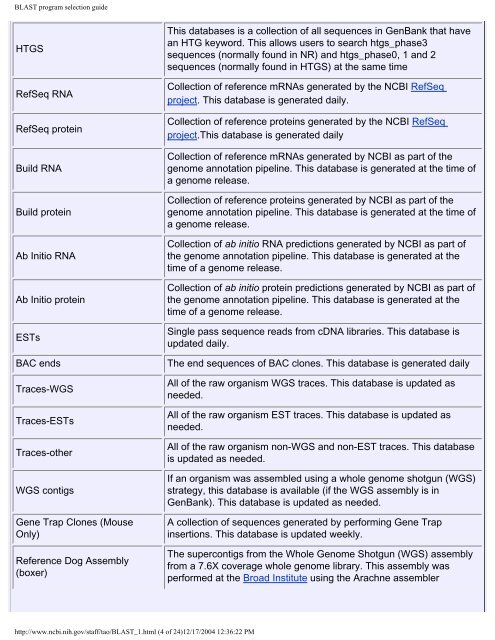
HTGS<br />
RefSeq RNA<br />
RefSeq protein<br />
Build RNA<br />
Build protein<br />
Ab Initio RNA<br />
Ab Initio protein<br />
ESTs<br />
BAC ends<br />
Traces-WGS<br />
Traces-ESTs<br />
Traces-other<br />
WGS contigs<br />
Gene Trap Clones (Mouse<br />
Only)<br />
Reference Dog Assembly<br />
(boxer)<br />
This databases is a collection of all sequences in GenBank that have<br />
an HTG keyword. This allows users to search htgs_phase3<br />
sequences (normally found in NR) and htgs_phase0, 1 and 2<br />
sequences (normally found in HTGS) at the same time<br />
Collection of reference mRNAs generated by the NCBI RefSeq<br />
project. This database is generated daily.<br />
Collection of reference proteins generated by the NCBI RefSeq<br />
project.This database is generated daily<br />
Collection of reference mRNAs generated by NCBI as part of the<br />
genome annotation pipeline. This database is generated at the time of<br />
a genome release.<br />
Collection of reference proteins generated by NCBI as part of the<br />
genome annotation pipeline. This database is generated at the time of<br />
a genome release.<br />
Collection of ab initio RNA predictions generated by NCBI as part of<br />
the genome annotation pipeline. This database is generated at the<br />
time of a genome release.<br />
Collection of ab initio protein predictions generated by NCBI as part of<br />
the genome annotation pipeline. This database is generated at the<br />
time of a genome release.<br />
Single pass sequence reads from cDNA libraries. This database is<br />
updated daily.<br />
The end sequences of BAC clones. This database is generated daily<br />
All of the raw organism WGS traces. This database is updated as<br />
needed.<br />
All of the raw organism EST traces. This database is updated as<br />
needed.<br />
All of the raw organism non-WGS and non-EST traces. This database<br />
is updated as needed.<br />
If an organism was assembled using a whole genome shotgun (WGS)<br />
strategy, this database is available (if the WGS assembly is in<br />
GenBank). This database is updated as needed.<br />
A collection of sequences generated by performing Gene Trap<br />
insertions. This database is updated weekly.<br />
The supercontigs from the Whole Genome Shotgun (WGS) assembly<br />
from a 7.6X coverage whole genome library. This assembly was<br />
performed at the Broad Institute using the Arachne assembler<br />
http://www.ncbi.nih.gov/staff/tao/<strong>BLAST</strong>_1.html (4 of 24)12/17/2004 12:36:22 PM