Brassica campestris L. ssp. chinensis M
Brassica campestris L. ssp. chinensis M
Brassica campestris L. ssp. chinensis M
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
450<br />
the BcMF5 transgenic plants of flowering Chinese<br />
cabbage.<br />
Integration and transcription confirmation of<br />
the hpRNA of the BcMF5 gene in transgenic<br />
plants of pBIA9-RMF5<br />
To confirm the hpRNA of the BcMF5 gene in the<br />
regenerated plants of flowering Chinese cabbage,<br />
19 T0 plants of pBIA9-RMF5 were subjected to PCR<br />
analysis with the primers specific for the uidA gene.<br />
Both PCR (Figure 5A) and the two transgenic lines<br />
TB6 and TB13 for DNA gel blot analysis (Figure 5B)<br />
confirmed that the hpRNA of the BcMF5 gene had<br />
integrated into the genome of the flowering<br />
Chinese cabbage transgenic seedling of pBIA9-<br />
RMF5. Data from the DNA hybridization showed<br />
that it was a single copy in the assayed transgenic<br />
lines TB6 and TB13 (Figure 5B). In addition, the RNA<br />
hybridization results for both TB6 and TB13<br />
indicated that the expression of the BcMF5 gene<br />
decreased seriously in the transgenic line of pBIA9-<br />
RMF5 (Figure 5C), thus confirming that the normal<br />
expression of BcMF5 in the flowering Chinese<br />
cabbage was inhibited by hpRNA-mediated RNA<br />
interference in the transgenic plant.<br />
ARTICLE IN PRESS<br />
Q. Zhang et al.<br />
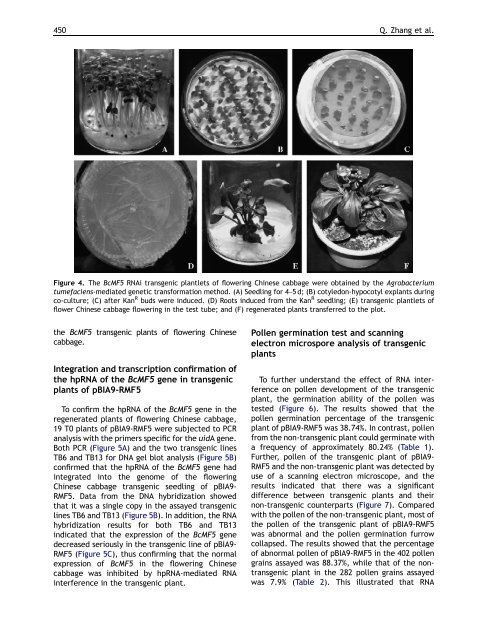
Figure 4. The BcMF5 RNAi transgenic plantlets of flowering Chinese cabbage were obtained by the Agrobacterium<br />
tumefaciens-mediated genetic transformation method. (A) Seedling for 4–5 d; (B) cotyledon-hypocotyl explants during<br />
co-culture; (C) after Kan R buds were induced. (D) Roots induced from the Kan R seedling; (E) transgenic plantlets of<br />
flower Chinese cabbage flowering in the test tube; and (F) regenerated plants transferred to the plot.<br />
Pollen germination test and scanning<br />
electron microspore analysis of transgenic<br />
plants<br />
To further understand the effect of RNA interference<br />
on pollen development of the transgenic<br />
plant, the germination ability of the pollen was<br />
tested (Figure 6). The results showed that the<br />
pollen germination percentage of the transgenic<br />
plant of pBIA9-RMF5 was 38.74%. In contrast, pollen<br />
from the non-transgenic plant could germinate with<br />
a frequency of approximately 80.24% (Table 1).<br />
Further, pollen of the transgenic plant of pBIA9-<br />
RMF5 and the non-transgenic plant was detected by<br />
use of a scanning electron microscope, and the<br />
results indicated that there was a significant<br />
difference between transgenic plants and their<br />
non-transgenic counterparts (Figure 7). Compared<br />
with the pollen of the non-transgenic plant, most of<br />
the pollen of the transgenic plant of pBIA9-RMF5<br />
was abnormal and the pollen germination furrow<br />
collapsed. The results showed that the percentage<br />
of abnormal pollen of pBIA9-RMF5 in the 402 pollen<br />
grains assayed was 88.37%, while that of the nontransgenic<br />
plant in the 282 pollen grains assayed<br />
was 7.9% (Table 2). This illustrated that RNA