anaerobic dehalogenation of halogenated organic compounds
anaerobic dehalogenation of halogenated organic compounds
anaerobic dehalogenation of halogenated organic compounds
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
508 Max M. Häggblom et al.<br />
dehalogenating populations are distinct under different redox conditions,<br />
e.g., sulfidogenic, iron-reducing or methanogenic (Knight et al., 1999;<br />
Häggblom et al., 2000; Rhee et al., 2003; Fennell et al., 2004a). Differences<br />
in substrate specificity, differences in apparent rate <strong>of</strong> <strong>dehalogenation</strong>, and<br />
unique community pr<strong>of</strong>iles indicate that distinct microbial populations are<br />
responsible for reductive <strong>dehalogenation</strong> under different dominant terminal<br />
electron accepting conditions (Knight et al., 1999; Häggblom et al., 2000,<br />
2003).<br />
A.<br />
181 bp<br />
215 bp<br />
DNA: 2<br />
-BP + Sulfate master culture<br />
B.<br />
215 bp<br />
224 bp<br />
RNA: Phenol + Sulfate<br />
254 bp 298 bp<br />
C.<br />
181 bp<br />
212 bp<br />
RNA: 2<br />
-BP + Hydrogen<br />
D.<br />
181 bp<br />
DNA: Cloned SSU gene<br />
2-BP 48<br />
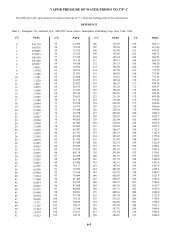
Figure 1. Comparison <strong>of</strong> T-RFLP analysis <strong>of</strong> genomic DNA from master culture 16S rRNA<br />
gene (A), reverse transcribed 16S rRNA from knock-out cultures phenol + sulfate (B), 2-<br />
bromophenol + hydrogen (C) and cloned 16S rRNA gene <strong>of</strong> 2BP-48 (from Fennell et al.,<br />
2004a).<br />
We have pursued the characterization <strong>of</strong> the structure and diversity<br />
<strong>of</strong> microbial food webs that are active during <strong>dehalogenation</strong> in <strong>anaerobic</strong><br />
estuarine and marine sediments in order to identify the dehalogenating<br />
organisms. We have applied an assay using a starved consortium (low<br />
ribosome content culture) fed a suite <strong>of</strong> selective substrates (Fennell et al.,<br />
2004a). The newly synthesized 16S rRNA was then analyzed using terminalrestriction<br />
fragment length polymorphism (T-RFLP) analysis <strong>of</strong> reverse<br />
transcribed rRNA as a means for assigning a metabolic role to the different<br />
members <strong>of</strong> the sulfidogenic 2-bromo-phenol-dehalogenating consortium.<br />
Those organisms that became active after substrate feeding were recognized<br />
as distinct peaks in the electropherogram and could be identified based on<br />
data from clonal libraries. Direct terminal restriction fragment length<br />
polymorphism fingerprinting <strong>of</strong> ribosomes in the various subcultures (fed