anaerobic dehalogenation of halogenated organic compounds
anaerobic dehalogenation of halogenated organic compounds
anaerobic dehalogenation of halogenated organic compounds
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
Mole %<br />
510 Max M. Häggblom et al.<br />
rdh61<br />
rdh63<br />
Dehalococcoides ethenogenes (Tce A)<br />
Dehalococcoides sp. VS<br />
Desulfitobacterium hafniense (CprA)<br />
Paleta creek2<br />
Paleta creek1<br />
85<br />
65<br />
Desulfitobacterium dehalogenans (CprA )<br />
63<br />
79<br />
35<br />
Sulfurospirillum<br />
multivorans (PceA)<br />
73<br />
33<br />
97<br />
SpongeE47<br />
Desulfitobacterium sp. Y51 (Pce A)<br />
100<br />
100<br />
0.1<br />
rdh81A<br />
E36<br />
E32<br />
E411<br />
Sponge group 2<br />
E42 E34<br />
E41 E310<br />
E29 E37 E45<br />
Sponge group 1<br />
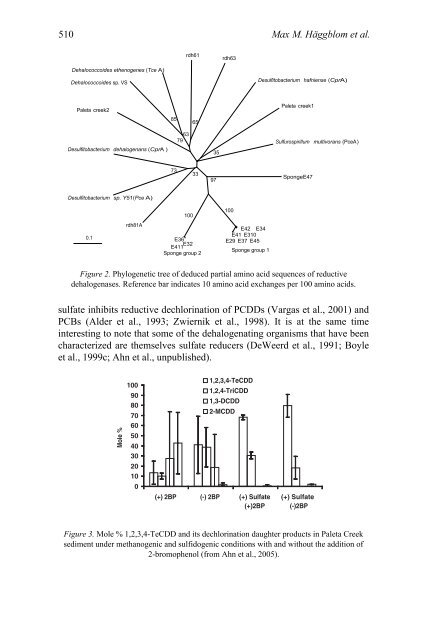
Figure 2. Phylogenetic tree <strong>of</strong> deduced partial amino acid sequences <strong>of</strong> reductive<br />
dehalogenases. Reference bar indicates 10 amino acid exchanges per 100 amino acids.<br />
sulfate inhibits reductive dechlorination <strong>of</strong> PCDDs (Vargas et al., 2001) and<br />
PCBs (Alder et al., 1993; Zwiernik et al., 1998). It is at the same time<br />
interesting to note that some <strong>of</strong> the dehalogenating organisms that have been<br />
characterized are themselves sulfate reducers (DeWeerd et al., 1991; Boyle<br />
et al., 1999c; Ahn et al., unpublished).<br />
100<br />
90<br />
80<br />
70<br />
60<br />
50<br />
40<br />
30<br />
20<br />
10<br />
0<br />
1,2,3,4-TeCDD<br />
1,2,4-TriCDD<br />
1,3-DCDD<br />
2-MCDD<br />
(+) 2BP (-) 2BP (+) Sulfate<br />
(+)2BP<br />
(+) Sulfate<br />
(-)2BP<br />
Figure 3. Mole % 1,2,3,4-TeCDD and its dechlorination daughter products in Paleta Creek<br />
sediment under methanogenic and sulfidogenic conditions with and without the addition <strong>of</strong><br />
2-bromophenol (from Ahn et al., 2005).