Compiler Usage Guidelines for 64-Bit Operating Systems on AMD64 ...
Compiler Usage Guidelines for 64-Bit Operating Systems on AMD64 ...
Compiler Usage Guidelines for 64-Bit Operating Systems on AMD64 ...
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
<str<strong>on</strong>g>Compiler</str<strong>on</strong>g> <str<strong>on</strong>g>Usage</str<strong>on</strong>g> <str<strong>on</strong>g>Guidelines</str<strong>on</strong>g> <str<strong>on</strong>g>for</str<strong>on</strong>g> AMD<str<strong>on</strong>g>64</str<strong>on</strong>g> Plat<str<strong>on</strong>g>for</str<strong>on</strong>g>ms<br />
32035 Rev. 3.22 November 2007<br />
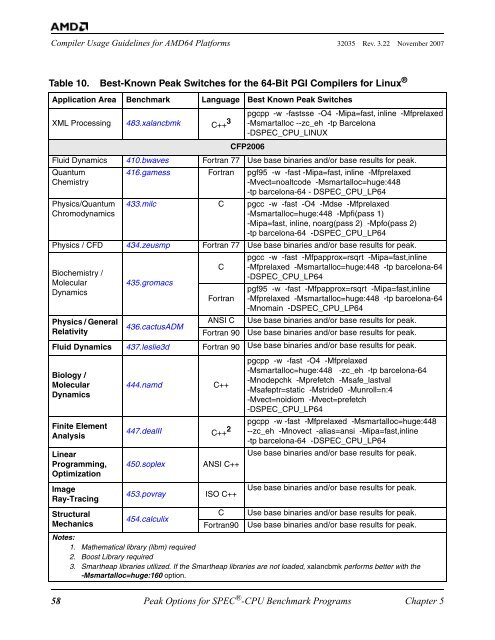
Table 10. Best-Known Peak Switches <str<strong>on</strong>g>for</str<strong>on</strong>g> the <str<strong>on</strong>g>64</str<strong>on</strong>g>-<str<strong>on</strong>g>Bit</str<strong>on</strong>g> PGI <str<strong>on</strong>g>Compiler</str<strong>on</strong>g>s <str<strong>on</strong>g>for</str<strong>on</strong>g> Linux ®<br />
XML Processing 483.xalancbmk C++ 3<br />
Applicati<strong>on</strong> Area Benchmark Language Best Known Peak Switches<br />
pgcpp -w -fastsse -O4 -Mipa=fast, inline -Mfprelaxed<br />
-Msmartalloc --zc_eh -tp Barcel<strong>on</strong>a<br />
-DSPEC_CPU_LINUX<br />
CFP2006<br />
Fluid Dynamics 410.bwaves Fortran 77 Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Quantum<br />
416.gamess Fortran pgf95 -w -fast -Mipa=fast, inline -Mfprelaxed<br />
Chemistry<br />
-Mvect=noaltcode -Msmartalloc=huge:448<br />
-tp barcel<strong>on</strong>a-<str<strong>on</strong>g>64</str<strong>on</strong>g> - DSPEC_CPU_LP<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
Physics/Quantum 433.milc C pgcc -w -fast -O4 -Mdse -Mfprelaxed<br />
Chromodynamics<br />
-Msmartalloc=huge:448 -Mpfi(pass 1)<br />
-Mipa=fast, inline, noarg(pass 2) -Mpfo(pass 2)<br />
-tp barcel<strong>on</strong>a-<str<strong>on</strong>g>64</str<strong>on</strong>g> -DSPEC_CPU_LP<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
Physics / CFD 434.zeusmp Fortran 77 Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Biochemistry /<br />
Molecular<br />
Dynamics<br />
435.gromacs<br />
C<br />
Fortran<br />
pgcc -w -fast -Mfpapprox=rsqrt -Mipa=fast,inline<br />
-Mfprelaxed -Msmartalloc=huge:448 -tp barcel<strong>on</strong>a-<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
-DSPEC_CPU_LP<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
pgf95 -w -fast -Mfpapprox=rsqrt -Mipa=fast,inline<br />
-Mfprelaxed -Msmartalloc=huge:448 -tp barcel<strong>on</strong>a-<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
-Mnomain -DSPEC_CPU_LP<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
Physics / General<br />
Relativity<br />
436.cactusADM<br />
ANSI C<br />
Fortran 90<br />
Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Fluid Dynamics 437.leslie3d Fortran 90 Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Biology /<br />
Molecular<br />
Dynamics<br />
Finite Element<br />
Analysis<br />
Linear<br />
Programming,<br />
Optimizati<strong>on</strong><br />
Image<br />
Ray-Tracing<br />
Structural<br />
Mechanics<br />
444.namd C++<br />
447.dealII C++ 2<br />
450.soplex ANSI C++<br />
453.povray ISO C++<br />
454.calculix<br />
pgcpp -w -fast -O4 -Mfprelaxed<br />
-Msmartalloc=huge:448 -zc_eh -tp barcel<strong>on</strong>a-<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
-Mnodepchk -Mprefetch -Msafe_lastval<br />
-Msafeptr=static -Mstride0 -Munroll=n:4<br />
-Mvect=noidiom -Mvect=prefetch<br />
-DSPEC_CPU_LP<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
pgcpp -w -fast -Mfprelaxed -Msmartalloc=huge:448<br />
--zc_eh -Mnovect -alias=ansi -Mipa=fast,inline<br />
-tp barcel<strong>on</strong>a-<str<strong>on</strong>g>64</str<strong>on</strong>g> -DSPEC_CPU_LP<str<strong>on</strong>g>64</str<strong>on</strong>g><br />
Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
C Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Fortran90 Use base binaries and/or base results <str<strong>on</strong>g>for</str<strong>on</strong>g> peak.<br />
Notes:<br />
1. Mathematical library (libm) required<br />
2. Boost Library required<br />
3. Smartheap libraries utilized. If the Smartheap libraries are not loaded, xalancbmk per<str<strong>on</strong>g>for</str<strong>on</strong>g>ms better with the<br />
-Msmartalloc=huge:160 opti<strong>on</strong>.<br />
58 Peak Opti<strong>on</strong>s <str<strong>on</strong>g>for</str<strong>on</strong>g> SPEC ® -CPU Benchmark Programs Chapter 5