Multifile Patent Sequence Searching on STN - STN International
Multifile Patent Sequence Searching on STN - STN International
Multifile Patent Sequence Searching on STN - STN International
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> <str<strong>on</strong>g>Patent</str<strong>on</strong>g> <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> <str<strong>on</strong>g>Searching</str<strong>on</strong>g> <strong>on</strong> <strong>STN</strong> ®<br />
Robert Austin – FIZ Karlsruhe
Agenda<br />
• <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> searchable databases <strong>on</strong> <strong>STN</strong> ®<br />
• Step-by-step through a multifile BLAST search<br />
• <str<strong>on</strong>g>Multifile</str<strong>on</strong>g> post-processing using <strong>STN</strong> Express<br />
• Overview of the search results<br />
• Summary and resources<br />
See also: <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> Basics e-Seminar (June 2010):<br />
http://www.stn-internati<strong>on</strong>al.com/<str<strong>on</strong>g>Sequence</str<strong>on</strong>g>_Basics_Seminar.html<br />
2
<strong>STN</strong> sequence searchable databases<br />
• DGENE<br />
– Thoms<strong>on</strong> Reuters GENESEQ TM<br />
– Value-added patent sequence data from around the globe<br />
• USGENE<br />
– The USPTO Genetic <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> Database<br />
– All available sequence data from the USPTO<br />
• PCTGEN<br />
– WIPO/PCT <str<strong>on</strong>g>Patent</str<strong>on</strong>g> Applicati<strong>on</strong> Biosequences<br />
– All available e-published sequence data from WIPO<br />
• CAS REGISTRY<br />
– Chemical Abstracts Service (CAS) REGISTRY<br />
– Worldwide value-added patent and n<strong>on</strong>-patent sequences<br />
3
DGENE, USGENE and PCTGEN offer three<br />
sequence search modes<br />
• <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> Code Match (motif) searching<br />
– Using the RUN GETSEQ command<br />
• BLAST similarity<br />
– Using the RUN BLAST command<br />
• FASTA similarity<br />
– Using the RUN GETSIM command<br />
Note: this e-Seminar covers BLAST.<br />
4
CAS REGISTRY/CAplus offers two<br />
sequence search modes<br />
• <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> Code Match (motif) searching<br />
– Using the Search (=> S) command<br />
• BLAST similarity<br />
– Using a separate Graphic User Interface<br />
Note: this e-Seminar covers BLAST.<br />
5
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> patent sequence searching<br />
Search Questi<strong>on</strong>:<br />
Find all patents that disclose Homo sapiens Damino-acid<br />
oxidase (NCBI NP_001908), or<br />
similar sequences (≥ 80%):<br />
MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLS<br />
DPNNPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGF<br />
RKLTPRELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVA<br />
REGADVIVNCTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPY<br />
IIPGTQTVTLGGIFQLGNWSELNNIQDHNTIWEGCCRLEPTLKNARIIGERTGFRPV<br />
RPQIRLEREQLRTGPSNTEVIHNYGHGGYGLTIHWGCALEAAKLFGRILEEKKLSRM<br />
PPSHL<br />
(Search c<strong>on</strong>ducted <strong>on</strong> 7 th July 2010)<br />
6
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> search strategy<br />
1) RUN BLAST in DGENE, USGENE and PCTGEN<br />
using offline BATCH mode<br />
2) Merge, organize by patent family, and display<br />
DGENE, USGENE and PCTGEN results<br />
3) Repeat the search using CAS REGISTRY BLAST<br />
4) Retrieve, identify, and display unique CAS<br />
REGISTRY BLAST CAplus records<br />
5) Post-process DGENE, USGENE and PCTGEN<br />
results using the <strong>STN</strong> Express Table Tool<br />
6) Post-process unique REGISTRY BLAST results<br />
using the BLAST Report Tool<br />
7
SAVE, UPLOAD and VERIFY the query<br />
• Prepare and save the query as a plain text file in<br />
a suitable text editor, e.g. Windows Notepad<br />
8
SAVE, UPLOAD and VERIFY the query (c<strong>on</strong>t.)<br />
(a) Click Upload <str<strong>on</strong>g>Sequence</str<strong>on</strong>g><br />
(b) Choose the query file<br />
(c) Select the <strong>STN</strong> database<br />
(a)<br />
From the Discover! butt<strong>on</strong> menu.<br />
(b)<br />
(c)<br />
The sequence becomes a Query<br />
L-number in the database of<br />
choice for use with RUN BLAST.<br />
9
SAVE, UPLOAD and VERIFY the query (c<strong>on</strong>t.)<br />
=> FILE USGENE<br />
=> UPL R BLAST<br />
Commands in red are automatically run by the<br />
<strong>STN</strong> Express <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> Query Upload wizard.<br />
Uploading C:\. . . .\NP_001908 Homo sapiens DAO.txt<br />
UPLOAD SUCCESSFULLY COMPLETED<br />
L1 GENERATED<br />
Verify the sequence was uploaded<br />
=> D L1 LQUE successfully with D LQUE.<br />
L1 ANSWER 1 USGENE COPYRIGHT 2010 SEQUENCEBASE CORP <strong>on</strong> <strong>STN</strong><br />
LQUE MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSD<br />
PNNPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRK<br />
LTPRELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREG<br />
ADVIVNCTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPG<br />
TQTVTLGGIFQLGNWSELNNIQDHNTIWEGCCRLEPTLKNARIIGERTGFRPVRPQIR<br />
LEREQLRTGPSNTEVIHNYGHGGYGLTIHWGCALEAAKLFGRILEEKKLSRMPPSHL<br />
The sequence query is now ready for searching directly in<br />
DGENE, USGENE, or PCTGEN using the L-number (L1).<br />
10
RUN the DGENE, USGENE and PCTGEN<br />
BLAST searches in BATCH mode<br />
=> FILE DGENE<br />
FILE 'DGENE' ENTERED AT 17:05:31 ON 07 JUL 2010<br />
COPYRIGHT (C) 2010 THOMSON REUTERS<br />
=> RUN BLAST L1 /SQP -F F BATCH<br />
PLEASE ENTER BATCH IDENTIFIER (MAX. 8 CHARS):DAOP<br />
TO BE NOTIFIED WHEN THIS BATCH SEARCH IS COMPLETE,<br />
PLEASE ENTER YOUR EMAIL ADDRESS (MAX. 50 CHARS) OR "NONE"<br />
INPUT: OR (END):ROBERT.AUSTIN@FIZ-KARLSRUHE.DE<br />
BLAST Versi<strong>on</strong> 2.2<br />
The BLAST software is used herein with permissi<strong>on</strong> of the<br />
Nati<strong>on</strong>al Center for Biotechnology Informati<strong>on</strong> (NCBI) of<br />
the Nati<strong>on</strong>al Library of Medicine (NLM). . . .<br />
BATCH PROCESSING STARTED FOR DAOP<br />
Add BATCH to the end of<br />
a RUN BLAST command<br />
to search in offline batch<br />
search mode.<br />
New!<br />
Enter a valid email<br />
address to be notified<br />
when the BATCH<br />
search is completed.<br />
11
RUN the DGENE, USGENE and PCTGEN<br />
BLAST searches in BATCH mode (c<strong>on</strong>t.)<br />
=> FILE USGENE<br />
=> RUN BLAST L1 /SQP -F F BATCH<br />
. . . .<br />
PLEASE ENTER BATCH IDENTIFIER (MAX. 8 CHARS):DAOP<br />
. . . .<br />
=> FILE PCTGEN<br />
=> RUN BLAST L1 /SQP -F F BATCH<br />
. . . .<br />
PLEASE ENTER BATCH IDENTIFIER (MAX. 8 CHARS):DAOP<br />
. . . .<br />
=> LOG H<br />
Note: DGENE, USGENE and<br />
PCTGEN BLAST searches can be<br />
run in parallel using BATCH mode.<br />
Turn the Low Complexity Filter off<br />
with the syntax: /SQP –F F<br />
Tip: use LOGOFF HOLD (LOG H)<br />
to be able to return to the same<br />
<strong>STN</strong> sessi<strong>on</strong> within two hours.<br />
SESSION WILL BE HELD FOR 120 MINUTES<br />
<strong>STN</strong> INTERNATIONAL SESSION SUSPENDED AT 17:07:14 ON 07 JUL 2010<br />
12
Retrieve the BATCH search results<br />
=> FILE DGENE<br />
FILE 'DGENE' ENTERED AT 17:11:25 ON 07 JUL 2010<br />
COPYRIGHT (C) 2010 THOMSON REUTERS<br />
=> RUN GETBATCH DAOP<br />
Use RUN GETBATCH to retrieve<br />
Please enter your batch identifier completed BATCH search results.<br />
or enter # for batch id list<br />
or enter * for batch id at top of list<br />
or enter - before batch id to delete<br />
or enter . for (end)<br />
Database DGENE AA<br />
Posted date: Jun 25, 2010 11:33 PM<br />
. . . .<br />
ENTER EITHER THE NUMBER OF ANSWERS YOU WISH TO KEEP<br />
OR ENTER MINIMUM PERCENT OF SELF SCORE FOLLOWED BY %<br />
(BEST ANSWER PERCENTAGE OF SELF SCORE IS 100%)<br />
ENTER (ALL) OR ? :80%<br />
L2 RUN STATEMENT CREATED<br />
L2 19 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYAD. . . MPPSHL/SQP.-F F<br />
In this example, 80% of the Query<br />
Self Score is used to select out<br />
just the most relevant results (L2).<br />
Answer set arranged by accessi<strong>on</strong> number; to sort by descending<br />
similarity score, enter at an arrow prompt (=>) "sor score d".<br />
13
Retrieve the BATCH search results (c<strong>on</strong>t.)<br />
=> FILE USGENE<br />
=> RUN GETBATCH DAOP<br />
. . . .<br />
ENTER EITHER THE NUMBER OF ANSWERS YOU WISH TO KEEP<br />
OR ENTER MINIMUM PERCENT OF SELF SCORE FOLLOWED BY %<br />
(BEST ANSWER PERCENTAGE OF SELF SCORE IS 100%)<br />
ENTER (ALL) OR ? :80%<br />
L3 RUN STATEMENT CREATED<br />
L3 14 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYAD. . . MPPSHL/SQP.-F F<br />
=> FILE PCTGEN<br />
=> RUN GETBATCH DAOP<br />
. . . .<br />
Use RUN GETBATCH to retrieve<br />
completed BATCH search results.<br />
ENTER EITHER THE NUMBER OF ANSWERS YOU WISH TO KEEP<br />
OR ENTER MINIMUM PERCENT OF SELF SCORE FOLLOWED BY %<br />
(BEST ANSWER PERCENTAGE OF SELF SCORE IS 100%)<br />
ENTER (ALL) OR ? :80%<br />
L4 RUN STATEMENT CREATED<br />
L4 3 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYAD. . . MPPSHL/SQP.-F F<br />
14
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> search strategy<br />
1) RUN BLAST in DGENE, USGENE and PCTGEN<br />
using offline BATCH mode<br />
2) Merge, organize by patent family, and display<br />
DGENE, USGENE and PCTGEN results<br />
3) Repeat the search using CAS REGISTRY BLAST<br />
4) Retrieve, identify, and display unique CAS<br />
REGISTRY BLAST CAplus records<br />
5) Post-process DGENE, USGENE and PCTGEN<br />
results using the <strong>STN</strong> Express Table Tool<br />
6) Post-process unique REGISTRY BLAST results<br />
using the BLAST Report Tool<br />
15
Merge the results into a single L-number<br />
=> SET DUPORDER FILE<br />
SET COMMAND COMPLETED<br />
=> DUP IDE L2 L3 L4<br />
FILE 'DGENE' ENTERED AT 17:16:56 ON 07 JUL 2010<br />
COPYRIGHT (C) 2010 THOMSON REUTERS<br />
FILE 'USGENE' ENTERED AT 17:16:56 ON 07 JUL 2010<br />
COPYRIGHT (C) 2010 SEQUENCEBASE CORP<br />
FILE 'PCTGEN' ENTERED AT 17:16:56 ON 07 JUL 2010<br />
COPYRIGHT (C) 2010 WIPO<br />
PROCESSING COMPLETED FOR L2<br />
PROCESSING COMPLETED FOR L3<br />
PROCESSING COMPLETED FOR L4<br />
L5 36 DUP IDE L2 L3 L4 (INCLUDES 0 SETS OF DUPLICATES)<br />
ANSWERS '1-19' FROM FILE DGENE<br />
=> SOR IDENT D<br />
PROCESSING COMPLETED FOR L5<br />
L6 36 SOR L5 IDENT D<br />
ANSWERS '20-33' FROM FILE USGENE<br />
ANSWERS '34-36' FROM FILE PCTGEN<br />
New!<br />
SET DUPORER FILE ensures that<br />
multifile records merged using DUP<br />
IDE are organized by database (file).<br />
DUPLICATE IDENTIFY<br />
(DUP IDE) is used<br />
here to create a single<br />
multifile L-number (L5).<br />
The multifile L-number<br />
(L5) can be sorted by<br />
BLAST SCORE, or<br />
Percent Identity (IDENT).<br />
16
Review multifile answers with a free-of-charge<br />
format including alignment<br />
=> D L6 TRIAL SCORE ALIGN 1-36; FILE <strong>STN</strong>GUIDE<br />
L6 ANSWER 1 OF 36 DGENE COPYRIGHT 2010 THOMSON REUTERS <strong>on</strong> <strong>STN</strong><br />
AN AAO23074 Protein DGENE<br />
TI Determining a genotype of an individual for preparing a compositi<strong>on</strong><br />
for treating schizophrenia by determining the identity of a<br />
nucleotide at a biallelic marker of the D-amino acid oxidase gene of<br />
the polynucleotide in a sample -<br />
DESC Human D-amino acid oxidase wild-type protein.<br />
KW Biallelic marker; D-amino acid oxidase; DAO; neuroleptic; CNS<br />
disorder; movement; Parkins<strong>on</strong>'s disease; Huntingt<strong>on</strong>'s; motor<br />
neur<strong>on</strong>e; Alzheimer's; mood; unipolar depressi<strong>on</strong>; bipolar; . . . .<br />
SQL 347<br />
Query Self Score<br />
and percentage.<br />
SCORE 731 100% of query self score 731<br />
BLASTALIGN<br />
Query = 347 letters<br />
Length = 347<br />
Score = 731 bits (1886), Expect = 0.0<br />
Identities = 347/347 (100%), Positives = 347/347 (100%)<br />
Query: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQP . . .<br />
MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQP<br />
Sbjct: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQP . . .<br />
17
Review answers with a free-of-charge format<br />
including alignment (c<strong>on</strong>t.)<br />
L6 ANSWER 4 OF 36 USGENE COPYRIGHT 2010 SEQUENCEBASE CORP <strong>on</strong> <strong>STN</strong><br />
TI Collecti<strong>on</strong>s of matched biological reagents and methods for<br />
identifying matched reagents (PublishedApplicati<strong>on</strong>)<br />
MTY Protein<br />
SQL 347<br />
SCORE 731 100% of query self score 731<br />
BLASTALIGN<br />
Query = 347 letters<br />
Length = 347<br />
Score = 731 bits (1886), Expect = 0.0<br />
Identities = 347/347 (100%), Positives = 347/347 (100%)<br />
BLAST Percent<br />
Identity (IDENT).<br />
Query: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Sbjct: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Query: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Sbjct: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Query: 121 ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
Sbjct: 121 ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
Query: 181 CTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPGTQ . . .<br />
CTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPGTQ<br />
Sbjct: 181 CTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPGTQ . . .<br />
18
Review answers with a free-of-charge format<br />
including alignment (c<strong>on</strong>t.)<br />
L6 ANSWER 28 OF 36 PCTGEN COPYRIGHT 2010 WIPO <strong>on</strong> <strong>STN</strong><br />
TI ORGAN-SPECIFIC PROTEINS AND METHODS OFTHEIR USE<br />
MTY PRT<br />
SQL 347<br />
SCORE 728 99% of query self score 731<br />
BLASTALIGN<br />
Query = 347 letters<br />
Length = 347<br />
Score = 728 bits (1879), Expect = 0.0<br />
Identities = 346/347 (99%), Positives = 346/347 (99%)<br />
Query: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
MRVVVIGAGVIGLSTALCIHERYHSVLQPL IKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Sbjct: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLHIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Query: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Sbjct: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Query: 121 ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
Sbjct: 121 ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
Query: 181 CTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPGTQ . . .<br />
CTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPGTQ<br />
Sbjct: 181 CTGVWAGALQRDPLLQPGRGQIMKVDAPWMKHFILTHDPERGIYNSPYIIPGTQ . . .<br />
19
Ensure Capture Sessi<strong>on</strong> is <strong>on</strong> to record a<br />
transcript for use in post-processing<br />
Note: Check the Capture<br />
Retrospectively box to capture<br />
the sessi<strong>on</strong> so far, as well as the<br />
sessi<strong>on</strong> from this point forwards.<br />
20
Use the <strong>STN</strong> Express 8.4 <str<strong>on</strong>g>Patent</str<strong>on</strong>g> Family<br />
Manager wizard display the results<br />
Access the patent family<br />
manager wizard from the<br />
Discover! Menu.<br />
Choose a bibliographic display format with<br />
alignment for the first (best) hit, and a free-ofcharge<br />
format with alignment for the rest of<br />
the sequences in each patent family group.<br />
21
The patent family manager begins by<br />
organising the results using FSORT...<br />
=> FSORT L6<br />
. . . .<br />
L7 36 FSO L6<br />
11 Multi-record Families Answers 1-33<br />
Family 1 Answers 1-5<br />
Family 2 Answers 6-8<br />
Family 3 Answers 9-10<br />
Family 4 Answers 11-12<br />
Family 5 Answers 13-14<br />
Family 6 Answers 15-16<br />
Family 7 Answers 17-18<br />
Family 8 Answers 19-25<br />
Family 9 Answers 26-27<br />
Family 10 Answers 28-31<br />
Family 11 Answers 32-33<br />
3 Individual Records Answers 34-36<br />
0 N<strong>on</strong>-patent Records<br />
Commands in RED are those<br />
issued automatically by the <strong>STN</strong><br />
Express <str<strong>on</strong>g>Patent</str<strong>on</strong>g> Family Manager.<br />
FSORT organizes<br />
the patent<br />
sequence records<br />
by Publicati<strong>on</strong>,<br />
Applicati<strong>on</strong>,<br />
Related, and<br />
Priority numbers.<br />
In this example, 14 patent family<br />
groups (i.e. 11 + 3) are retrieved.<br />
22
...and then c<strong>on</strong>tinues by displaying the<br />
family groups in the specified formats<br />
=> DIS L7 PFAM=7 1 BIB,SQL,SCORE,IDENT,ALIGN<br />
L7 ANSWER 17 OF 36 DGENE COPYRIGHT 2010 THOMSON REUTERS <strong>on</strong> <strong>STN</strong> FAMILY7<br />
AN AEL25470 protein DGENE<br />
TI Identifying compound that reduce/inhibit internal ribosome . . . .<br />
IN Fear M<br />
PA (TELE-N) TELETHON INST CHILD HEALTH RES.<br />
PI WO 2006102720 A1 20061005 197<br />
AI WO 2006-AU435 20060331<br />
PRAI AU 2005-901574 20050331<br />
PSL Disclosure; SEQ ID NO 18<br />
LA English<br />
OS 2006-747347 [76]<br />
CR N-PSDB: AEL25469<br />
PC-NCBI: gi30446<br />
PC-SWISSPROT: P14920<br />
DESC Reporter protein SEQ ID NO:18.<br />
SQL 347<br />
SCORE 726 99% of query self score 731<br />
IDENT 99%<br />
BLASTALIGN<br />
Query = 347 letters<br />
Length = 347<br />
Score = 726 bits (1873), Expect = 0.0<br />
Identities = 345/347 (99%), Positives = 345/347 (99%)<br />
. . . .<br />
. . . .<br />
Commands in RED are those<br />
issued automatically by the <strong>STN</strong><br />
Express <str<strong>on</strong>g>Patent</str<strong>on</strong>g> Family Manager.<br />
23
...and then c<strong>on</strong>tinues by displaying the<br />
family groups in the specified formats (c<strong>on</strong>t.)<br />
=> DIS L7 PFAM=7 2-TOT TRIAL,SCORE,IDENT,ALIGN<br />
L7 ANSWER 18 OF 36 USGENE COPYRIGHT 2010 SEQUENCEBASE CORP <strong>on</strong> <strong>STN</strong>FAMILY7<br />
TI Isolati<strong>on</strong> of Inhibitors of IRES-Mediated Translati<strong>on</strong><br />
(PublishedApplicati<strong>on</strong>)<br />
DESC Homo Sapiens Protein; sequence 18 of 148<br />
MTY Protein<br />
SQL 347<br />
SCORE 726 99% of query self score 731<br />
IDENT 99%<br />
BLASTALIGN<br />
Query = 347 letters<br />
Length = 347<br />
Score = 726 bits (1873), Expect = 0.0<br />
Identities = 345/347 (99%), Positives = 345/347 (99%)<br />
This USGENE hit is in the same<br />
family as the DGENE record <strong>on</strong><br />
the previous slide (FAMILY 7).<br />
Query: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
MRVVVIGAGVIGLSTALCIHERYHSVLQPL IKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Sbjct: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLHIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Query: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Sbjct: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Query: 121 ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
Sbjct: 121 ELDMFPDYGYGWFHTSLILEGKNYLQWLTERLTERGVKFFQRKVESFEEVAREGADVIVN<br />
. . . .<br />
24
...and then c<strong>on</strong>tinues by displaying the<br />
family groups in the specified formats (c<strong>on</strong>t.)<br />
=> DIS L7 34-36 BIB,SQL,SCORE,IDENT,ALIGN<br />
L7 ANSWER 34 OF 36 USGENE COPYRIGHT 2010 SEQUENCEBASE CORP <strong>on</strong> <strong>STN</strong><br />
AN 20060275794.63099 Protein USGENE<br />
TI Collecti<strong>on</strong>s of matched biological reagents and methods for<br />
identifying matched reagents (PublishedApplicati<strong>on</strong>)<br />
IN Carrino John (San Diego, CA); Liang Feng (San Diego, CA)<br />
PA Invitrogen Corporati<strong>on</strong> (Carlsbad CA)<br />
PI US 20060275794 A1 20061207<br />
AI US 2006-371354 20060307<br />
DT <str<strong>on</strong>g>Patent</str<strong>on</strong>g><br />
SQL 347<br />
SCORE 731 100% of query self score 731<br />
IDENT 100%<br />
BLASTALIGN<br />
Query = 347 letters<br />
Length = 347<br />
Score = 731 bits (1886), Expect = 0.0<br />
Identities = 347/347 (100%), Positives = 347/347 (100%)<br />
This USGENE record is the first<br />
of the 3 “individual records” in<br />
the FSORT answer set (L7).<br />
Query: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Sbjct: 1 MRVVVIGAGVIGLSTALCIHERYHSVLQPLDIKVYADRFTPLTTTDVAAGLWQPYLSDPN<br />
Query: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
Sbjct: 61 NPQEADWSQQTFDYLLSHVHSPNAENLGLFLISGYNLFHEAIPDPSWKDTVLGFRKLTPR<br />
. . . .<br />
25
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> search strategy<br />
1) RUN BLAST in DGENE, USGENE and PCTGEN<br />
using offline BATCH mode<br />
2) Merge, organize by patent family, and display<br />
DGENE, USGENE and PCTGEN results<br />
3) Repeat the search using CAS REGISTRY BLAST<br />
4) Retrieve, identify, and display unique CAS<br />
REGISTRY BLAST CAplus records<br />
5) Post-process DGENE, USGENE and PCTGEN<br />
results using the <strong>STN</strong> Express Table Tool<br />
6) Post-process unique REGISTRY BLAST results<br />
using the BLAST Report Tool<br />
26
Typical steps of CAS REGISTRY BLAST<br />
1. Launch BLAST<br />
2. Search the sequence<br />
3. Examine and evaluate alignment/relevance of<br />
sequence answers<br />
4. Display <strong>STN</strong> data <strong>on</strong> sequences – REGISTRY<br />
5. Display <strong>STN</strong> data <strong>on</strong> sequences – CAplus SM<br />
– Limit CAplus results, if necessary<br />
– Display CAplus data (references and HITRN)<br />
6. Post-process BLAST alignment data<br />
27
Launch CAS REGISTRY BLAST<br />
• The Result Set Manager is<br />
the starting point<br />
• To begin a new sequence<br />
search<br />
• To review results of previous<br />
sequence searches<br />
28
Input the search query<br />
• <str<strong>on</strong>g>Sequence</str<strong>on</strong>g>s can be input by Copy/paste<br />
• Read from a file<br />
• Recall a previously searched sequence<br />
within the same sessi<strong>on</strong><br />
• <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> line numbers do not<br />
interfere with the search.<br />
29
Select the BLAST program<br />
The following programs are<br />
most typically run:<br />
• BLASTn for nucleotides<br />
• BLASTp for proteins/peptides<br />
30
Verify BLAST settings<br />
Default values have been set to<br />
optimize sequence searches for<br />
researchers.<br />
Recommended settings for<br />
patent searches:<br />
• Low Complexity Filtering –<br />
unchecked<br />
• Max No. of Answers - 1000<br />
31
View results<br />
Highlight the result set<br />
to be viewed, and click<br />
<strong>on</strong> View Results.<br />
32
Evaluate the alignment report<br />
The negative sign represents<br />
that the alignment details are<br />
shown.<br />
Detail informati<strong>on</strong> such as the<br />
sequence length, score,<br />
percent identity are available.<br />
33
Select sequences of interest<br />
<str<strong>on</strong>g>Sequence</str<strong>on</strong>g>s can be selected:<br />
• In groups, using the color bar in the<br />
Alignment Scores<br />
• Individually, by selecting the check box<br />
• To transfer the sequence data to <strong>STN</strong>,<br />
click the Get <strong>STN</strong> Data butt<strong>on</strong>.<br />
34
Get <strong>STN</strong> Data and Save alignments (.xss)<br />
Alignment data needs<br />
to be transferred for<br />
post-processing.<br />
The alignment data is saved in <strong>STN</strong><br />
Express Saved <str<strong>on</strong>g>Sequence</str<strong>on</strong>g>s (.xss) format.<br />
35
Transfer sequences to <strong>STN</strong><br />
• Log<strong>on</strong> to <strong>STN</strong> and a REGISTRY search<br />
of the sequences is automatic.<br />
• Results display can be accomplished<br />
using either Discover! wizards or<br />
command line input.<br />
• Note: Type END or click Cancel to get<br />
out of the “Display Wizard”. You can turn<br />
off the “Display Wizard” in Preferences.<br />
Display sequences if desired.<br />
36
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> search strategy<br />
1) RUN BLAST in DGENE, USGENE and PCTGEN<br />
using offline BATCH mode<br />
2) Merge, organize by patent family, and display<br />
DGENE, USGENE and PCTGEN results<br />
3) Repeat the search using CAS REGISTRY BLAST<br />
4) Retrieve, identify, and display unique CAS<br />
REGISTRY BLAST CAplus records<br />
5) Post-process DGENE, USGENE and PCTGEN<br />
results using the <strong>STN</strong> Express Table Tool<br />
6) Post-process unique REGISTRY BLAST results<br />
using the BLAST Report Tool<br />
37
Display additi<strong>on</strong>al CAplus answers including<br />
the HITRN for alignment post-processing<br />
=> FILE HCAPLUS<br />
FILE 'HCAPLUS' ENTERED AT 17:25:10 ON 07 JUL 2010<br />
COPYRIGHT (C) 2010 AMERICAN CHEMICAL SOCIETY (ACS)<br />
=> S L12 AND PATENT/DT<br />
The 44 REGISTRY records (L12)<br />
L13 12 L12 AND PATENT/DT<br />
corresp<strong>on</strong>d to 12 HCAplus patent<br />
records (L13).<br />
=> TRANSFER L6 PN 1-<br />
L14 TRANSFER L6 1- PN : Transfer 20 TERMS Publicati<strong>on</strong> Numbers (PN)<br />
L15 29 L14<br />
from DGENE/USGENE/PCTGEN<br />
ALL TERMS IN L14 RETRIEVED.<br />
(L6) to find corresp<strong>on</strong>ding HCAplus<br />
records (L15).<br />
=> S L13 NOT L15<br />
L16 2 L13 NOT L15<br />
=> D BIB HITRN 1-2<br />
In this example, 2 additi<strong>on</strong>al, highly<br />
relevant references have been<br />
found by including the<br />
REGISTRY/HCAplus search (L16).<br />
38
Example: Unique REGISTRY/CAplus result<br />
L16 ANSWER 1 OF 2 HCAPLUS COPYRIGHT 2010 ACS <strong>on</strong> <strong>STN</strong><br />
AN 2002:391912 HCAPLUS<br />
DN 137:1836<br />
TI Measurement of DNA methylati<strong>on</strong> for analysis of the toxicology . . . .<br />
IN Olek, Alexander; Piepenbrock, Christian; Berlin, Kurt<br />
PA Epigenomics Ag, Germany<br />
SO PCT Int. Appl., 113 pp.<br />
CODEN: PIXXD2<br />
LA German<br />
FAN.CNT 1<br />
PATENT NO. KIND DATE APPLICATION NO. DATE<br />
--------------- ---- -------- -------------------- --------<br />
PI WO 2002040710 A2 20020523 WO 2001-EP12951 20011108<br />
. . . .<br />
PRAI DE 2000-10056802 A 20001114<br />
WO 2001-EP12951 W 20011108<br />
Note: HITRN must be included,<br />
IT 391975-30-7, Protein (human 347-amino acid)<br />
RL: BSU (Biological study, unclassified); so that PRP the (Properties); CAS REGISTRY BIOL<br />
(Biological study)<br />
BLAST alignments can be<br />
(amino acid sequence; measurement of DNA methylati<strong>on</strong> for anal. of<br />
the toxicol. of substances)<br />
merged into the BLAST Report.<br />
39
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> search strategy<br />
1) RUN BLAST in DGENE, USGENE and PCTGEN<br />
using offline BATCH mode<br />
2) Merge, organize by patent family, and display<br />
DGENE, USGENE and PCTGEN results<br />
3) Repeat the search using CAS REGISTRY BLAST<br />
4) Retrieve, identify, and display unique CAS<br />
REGISTRY BLAST CAplus records<br />
5) Post-process DGENE, USGENE and PCTGEN<br />
results using the <strong>STN</strong> Express Table Tool<br />
6) Post-process unique REGISTRY BLAST results<br />
using the BLAST Report Tool<br />
40
Access the Table Tool and select the<br />
multifile search Transcript file<br />
The most recent <strong>STN</strong> sessi<strong>on</strong><br />
Transcript is usually listed here.<br />
41
Choose a template and select c<strong>on</strong>tent<br />
Opti<strong>on</strong>: choose a predefined<br />
custom template<br />
from a previous project.<br />
L7 is the DGENE,<br />
USGENE and PCTGEN<br />
FSORTed answer set.<br />
42
Select fields, column order, headings, f<strong>on</strong>ts<br />
and spacing for the table<br />
The pre-defined custom<br />
template included a list<br />
of fields. These can be<br />
further customized and<br />
the template re-saved.<br />
43
Review, adjust, and export the table<br />
44
Explore the results further in Microsoft Excel<br />
Some tips for Microsoft Excel:<br />
• Resize columns and rows as desired –<br />
especially the BLAST alignment<br />
column to approx 77<br />
• View, Freeze panes – holds the top row<br />
fixed when scrolling down<br />
• Add Filters – provides a great way to<br />
navigate results – for example by<br />
BLAST percent identity (above)<br />
45
<str<strong>on</strong>g>Multifile</str<strong>on</strong>g> search strategy<br />
1) RUN BLAST in DGENE, USGENE and PCTGEN<br />
using offline BATCH mode<br />
2) Merge, organize by patent family, and display<br />
DGENE, USGENE and PCTGEN results<br />
3) Repeat the search using CAS REGISTRY BLAST<br />
4) Retrieve, identify, and display unique CAS<br />
REGISTRY BLAST CAplus records<br />
5) Post-process DGENE, USGENE and PCTGEN<br />
results using the <strong>STN</strong> Express Table Tool<br />
6) Post-process unique REGISTRY BLAST results<br />
using the BLAST Report Tool<br />
46
Post-process REGISTRY BLAST alignments<br />
Download the post-processing template (.PRF) files used in this seminar:<br />
http://www.stn-internati<strong>on</strong>al.com/stn_biosequence_searching_mfs.html<br />
47
Select BLAST alignment report<br />
• The first step is to select the XSS<br />
file to include in the BLAST report.<br />
• Important: If your BLAST query is<br />
fairly l<strong>on</strong>g, or a nucleic acid, or the<br />
answers may exceed 1000<br />
characters, make sure you change<br />
the value in the Do not include<br />
alignments l<strong>on</strong>ger than box.<br />
Post-processing then c<strong>on</strong>tinues<br />
via standard <strong>STN</strong> Express<br />
Custom Report Tool steps.<br />
48
Select the sessi<strong>on</strong> Transcript and template<br />
Opti<strong>on</strong>: choose a predefined<br />
custom template<br />
from a previous project.<br />
The most recent <strong>STN</strong><br />
sessi<strong>on</strong> Transcript is<br />
usually listed here.<br />
49
Select the records to be processed<br />
L16 is REGISTRY/CAplus<br />
additi<strong>on</strong>al unique answers.<br />
50
Select fields, f<strong>on</strong>ts and spacing for the report<br />
The pre-defined custom<br />
template included a list<br />
of fields. These can be<br />
further customized and<br />
the template re-saved.<br />
51
Review, adjust, and export the report<br />
52
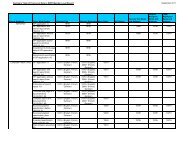
Overview of search results for Homo sapiens Damino-acid<br />
oxidase – unique in (red)<br />
SEQs<br />
≥ 80%<br />
PNs<br />
<str<strong>on</strong>g>Patent</str<strong>on</strong>g><br />
Families*<br />
DGENE 19 10 8 (1)<br />
USGENE 14 10 7 (2)<br />
PCTGEN 3 3 3 (1)<br />
REGISTRY 18 12 9 (2)<br />
NCBI 6 4 4 (0)<br />
Total Unique - - 14<br />
(* <str<strong>on</strong>g>Patent</str<strong>on</strong>g> families = INPADOC <str<strong>on</strong>g>Patent</str<strong>on</strong>g> Families. Specifically, family records in INPAFAMDB.)
Summary<br />
• RUN BLAST is available for searching DGENE,<br />
USGENE and PCTGEN directly <strong>on</strong> <strong>STN</strong><br />
• CAS REGISTRY BLAST provides BLAST searching<br />
opti<strong>on</strong>s for the REGISTRY database<br />
• DGENE, USGENE and PCTGEN multifile search<br />
results can be post-processed into tables, and<br />
exported to Microsoft Excel, using <strong>STN</strong> Express<br />
• CAS REGISTRY BLAST alignment data can be<br />
merged with CAplus records, and exported in to RTF<br />
format, to form single unified report<br />
• All four <strong>STN</strong> sequence databases are required for a<br />
comprehensive patent sequence search<br />
54
Resources for sequence searching <strong>on</strong> <strong>STN</strong><br />
• <str<strong>on</strong>g>Sequence</str<strong>on</strong>g> <str<strong>on</strong>g>Searching</str<strong>on</strong>g> <strong>on</strong> <strong>STN</strong> modular workshop<br />
http://www.stn-internati<strong>on</strong>al.com/sequence_searching.html<br />
• CAS REGISTRY sequence searching resources<br />
http://www.cas.org/support/stngen/stndoc/sequences.html<br />
• DGENE Workshop Manual<br />
http://www.stn-internati<strong>on</strong>al.com/dgene_wm.html<br />
• USGENE Workshop Manual<br />
http://www.stn-internati<strong>on</strong>al.com/usgene_wm.html<br />
• USGENE Workshop Manual <str<strong>on</strong>g>Multifile</str<strong>on</strong>g> Supplement:<br />
http://www.stn-internati<strong>on</strong>al.com/usgene_wm_mfs.html<br />
55
CAS<br />
E-mail: help@cas.org<br />
Support and Training:<br />
www.cas.org<br />
For more informati<strong>on</strong> …<br />
FIZ Karlsruhe<br />
helpdesk@fiz-karlsruhe.de<br />
Support and Training:<br />
www.stn-internati<strong>on</strong>al.de