Cryopreservation of Quercus suber somatic embryos - Tree Physiology
Cryopreservation of Quercus suber somatic embryos - Tree Physiology
Cryopreservation of Quercus suber somatic embryos - Tree Physiology
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
ally associated with callus formation. <strong>Cryopreservation</strong> may<br />
therefore prove generally effective in reducing intermediate<br />
callus formation during clonal propagation (Takagi 2000).<br />
It is critical that embryogenic clones are maintained indefinitely,<br />
without change in genetic makeup or loss <strong>of</strong> viability<br />
during cryogenic storage (Park 2002). <strong>Cryopreservation</strong> may<br />
induce genetic instability (Sakai 2004); however, the number<br />
<strong>of</strong> studies on the occurrence <strong>of</strong> genetic variation in cryopreserved<br />
samples remains limited (e.g., Panis and Lombardi<br />
2005). For example, no genetic stability evaluations were performed<br />
in previously reported cryopreservation experiments<br />
with embryogenic cultures <strong>of</strong> Q. <strong>suber</strong> (Martinez et al. 2003,<br />
Valladares et al. 2004). However, Hao et al. (2002a) successfully<br />
cryopreserved cell suspensions <strong>of</strong> Citrus sp. without<br />
ploidy variation and randomly amplified polymorphic DNA<br />
(RAPD) variations. In addition, surviving cells <strong>of</strong> some genotypes<br />
regenerated <strong>somatic</strong> <strong>embryos</strong> with better performance<br />
than the controls. Cryopreserved strawberry shoot tips also<br />
showed ploidy stability, and AFLP analyses revealed only one<br />
band differing from non-cryopreserved samples (Hao et al.<br />
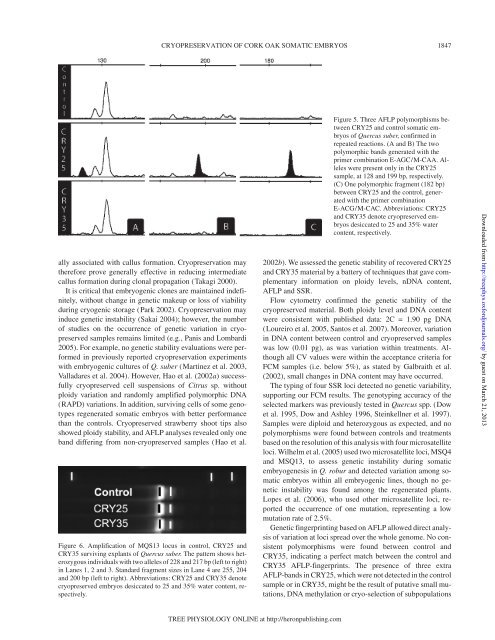
Figure 6. Amplification <strong>of</strong> MQS13 locus in control, CRY25 and<br />
CRY35 surviving explants <strong>of</strong> <strong>Quercus</strong> <strong>suber</strong>. The pattern shows heterozygous<br />
individuals with two alleles <strong>of</strong> 228 and 217 bp (left to right)<br />
in Lanes 1, 2 and 3. Standard fragment sizes in Lane 4 are 255, 204<br />
and 200 bp (left to right). Abbreviations: CRY25 and CRY35 denote<br />
cryopreserved <strong>embryos</strong> desiccated to 25 and 35% water content, respectively.<br />
CRYOPRESERVATION OF CORK OAK SOMATIC EMBRYOS 1847<br />
TREE PHYSIOLOGY ONLINE at http://heronpublishing.com<br />
Figure 5. Three AFLP polymorphisms between<br />
CRY25 and control <strong>somatic</strong> <strong>embryos</strong><br />
<strong>of</strong> <strong>Quercus</strong> <strong>suber</strong>, confirmed in<br />
repeated reactions. (A and B) The two<br />
polymorphic bands generated with the<br />
primer combination E-AGC/M-CAA. Alleles<br />
were present only in the CRY25<br />
sample, at 128 and 199 bp, respectively.<br />
(C) One polymorphic fragment (182 bp)<br />
between CRY25 and the control, generated<br />
with the primer combination<br />
E-ACG/M-CAC. Abbreviations: CRY25<br />
and CRY35 denote cryopreserved <strong>embryos</strong><br />
desiccated to 25 and 35% water<br />
content, respectively.<br />
2002b). We assessed the genetic stability <strong>of</strong> recovered CRY25<br />
and CRY35 material by a battery <strong>of</strong> techniques that gave complementary<br />
information on ploidy levels, nDNA content,<br />
AFLP and SSR.<br />
Flow cytometry confirmed the genetic stability <strong>of</strong> the<br />
cryopreserved material. Both ploidy level and DNA content<br />
were consistent with published data: 2C = 1.90 pg DNA<br />
(Loureiro et al. 2005, Santos et al. 2007). Moreover, variation<br />
in DNA content between control and cryopreserved samples<br />
was low (0.01 pg), as was variation within treatments. Although<br />
all CV values were within the acceptance criteria for<br />
FCM samples (i.e. below 5%), as stated by Galbraith et al.<br />
(2002), small changes in DNA content may have occurred.<br />
The typing <strong>of</strong> four SSR loci detected no genetic variability,<br />
supporting our FCM results. The genotyping accuracy <strong>of</strong> the<br />
selected markers was previously tested in <strong>Quercus</strong> spp. (Dow<br />
et al. 1995, Dow and Ashley 1996, Steinkellner et al. 1997).<br />
Samples were diploid and heterozygous as expected, and no<br />
polymorphisms were found between controls and treatments<br />
based on the resolution <strong>of</strong> this analysis with four microsatellite<br />
loci. Wilhelm et al. (2005) used two microsatellite loci, MSQ4<br />
and MSQ13, to assess genetic instability during <strong>somatic</strong><br />
embryogenesis in Q. robur and detected variation among <strong>somatic</strong><br />
<strong>embryos</strong> within all embryogenic lines, though no genetic<br />
instability was found among the regenerated plants.<br />
Lopes et al. (2006), who used other microsatellite loci, reported<br />
the occurrence <strong>of</strong> one mutation, representing a low<br />
mutation rate <strong>of</strong> 2.5%.<br />
Genetic fingerprinting based on AFLP allowed direct analysis<br />
<strong>of</strong> variation at loci spread over the whole genome. No consistent<br />
polymorphisms were found between control and<br />
CRY35, indicating a perfect match between the control and<br />
CRY35 AFLP-fingerprints. The presence <strong>of</strong> three extra<br />
AFLP-bands in CRY25, which were not detected in the control<br />
sample or in CRY35, might be the result <strong>of</strong> putative small mutations,<br />
DNA methylation or cryo-selection <strong>of</strong> subpopulations<br />
Downloaded from<br />
http://treephys.oxfordjournals.org/ by guest on March 21, 2013