single PDF file - Madagascar
single PDF file - Madagascar
single PDF file - Madagascar
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
MADAGASCAR DOCUMENTATION<br />
Maurice Aye-Aye, Sergey Fomel, Gilles Hennenfent, and Paul Sava<br />
http://ahay.org/
Copyright c○ 2011-12<br />
by <strong>Madagascar</strong> Community
i<br />
RSF — TABLE OF CONTENTS<br />
Maurice the Aye-Aye, <strong>Madagascar</strong> tutorial: Field data processing . . . . . . . 1<br />
Paul Sava, Seismic Imaging Tutorial: “exploding reflector” modeling/migration<br />
. . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 15<br />
Maurice the Aye-Aye, <strong>Madagascar</strong> tutorial . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 23<br />
Sergey Fomel, Guide to <strong>Madagascar</strong> programs . . . . . . . . . . . . . . . . . . . . . . . . . . . 37<br />
Sergey Fomel, Guide to RSF format . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 87<br />
Sergey Fomel, Revisiting SEP tour with <strong>Madagascar</strong> and SCons . . . . . . . . . 103<br />
Sergey Fomel, Guide to RSF API . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 119<br />
Paul Sava, Guide to programming using RSF . . . . . . . . . . . . . . . . . . . . . . . . . . . 135<br />
Sergey Fomel and Gilles Hennenfent, Reproducible computational experiments<br />
using SCons. . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 143
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
<strong>Madagascar</strong> tutorial: Field data processing<br />
Maurice the Aye-Aye<br />
ABSTRACT<br />
In this tutorial, you will learn about multiple attenuation using parabolic Radon<br />
transform (Hampson, 1986). You will first go through an example that explains<br />
the process step by step. You will be asked to change some parameters and add<br />
missing few lines. In the next part of the tutorial, you will be asked to apply the<br />
same workflow to another CMP gather. The CMP gathers used in the tutorial<br />
are from the Canterbury data set (Lu et al., 2003). By the end of this tutorial,<br />
you should have learned to:<br />
1. apply NMO and inverse NMO for a CMP gather,<br />
2. apply forward and inverse parabloic Radon transform,<br />
3. design a mute function that preserves multiples in the Radon domain,<br />
4. subtract multiples from the data,<br />
5. create a semblance scan for a CMP gather.<br />
Completing this tutorial requires<br />
PREREQUISITES<br />
• <strong>Madagascar</strong> software environment available from<br />
http://www.ahay.org<br />
• L A TEX environment with SEGTeX available from<br />
http://www.ahay.org/wiki/SEGTeX<br />
To do the assignment on your personal computer, you need to install the required<br />
environments. An Internet connection is required for access to the data repository.<br />
The tutorial itself is available from the <strong>Madagascar</strong> repository by running<br />
svn co https://rsf.svn.sourceforge.net/svnroot/rsf/trunk/book/rsf/school2012<br />
1
2 Maurice <strong>Madagascar</strong> Documentation<br />
INTRODUCTION<br />
In this tutorial, you will be asked to run commands from the Unix shell (identified<br />
by bash$) and to edit <strong>file</strong>s in a text editor. Different editors are available in a typical<br />
Unix environment (vi, emacs, nedit, etc.)<br />
Your first assignment:<br />
1. Open a Unix shell.<br />
2. Change directory to the tutorial directory<br />
bash$ cd $RSFSRC/book/rsf/school2012<br />
3. Open the tutorial.tex <strong>file</strong> in your favorite editor, for example by running<br />
bash$ nedit tutorial.tex &<br />
4. Look at the first line in the <strong>file</strong> and change the author name from Maurice the<br />
Aye-Aye to your name (first things first).<br />
Part One<br />
DEMO<br />
1. Change directory to the demo directory<br />
bash$ cd demo<br />
2. Run<br />
bash$ scons cmp.view<br />
in the Unix shell. A number of commands will appear in the shell followed by<br />
Figure 3(a) appearing on your screen.<br />
3. To understand the commands, examine the script that generated them by opening<br />
the SConstruct <strong>file</strong> in a text editor. Notice that, instead of Shell commands,<br />
the script contains rules.<br />
• The first rule, Fetch, allows the script to download the input data <strong>file</strong><br />
cmp1.rsf from the data server.<br />
• Other rules have the form Flow(target,source,command) for generating<br />
data <strong>file</strong>s or Plot and Result for generating picture <strong>file</strong>s.
<strong>Madagascar</strong> Documentation Tutorial 3<br />
• Fetch, Flow, Plot, and Result are defined in <strong>Madagascar</strong>’s rsf.proj<br />
package, which extends the functionality of SCons .<br />
4. To better understand how rules translate into commands, run<br />
bash$ scons -c cmp.rsf<br />
The -c flag tells scons to remove the cmp.rsf <strong>file</strong> and all its dependencies.<br />
5. Next, run<br />
bash$ scons -n cmp.rsf<br />
The -n flag tells scons not to run the command but simply to display it on the<br />
screen. Identify the lines in the SConstruct <strong>file</strong> that generate the output you<br />
see on the screen.<br />
6. Run<br />
bash$ scons cmp.rsf<br />
Examine the <strong>file</strong> cmp.rsf both by opening it in a text editor and by running<br />
bash$ sfin cmp.rsf<br />
Part Two<br />
Figure 3(a) shows a CMP gather from Canterbury data set Line 12. The multiple<br />
energy appears at time around 2.25 s. Figure 6 shows the same gather after applying<br />
NMO correction with veloctiy equals to 1500 m/s. The multiple events starting at<br />
around 2.25 s and below are flatened while primary events , e.g at 2 s, are over<br />
corrected. The difference in move-out between the primaries and multiples , hence,<br />
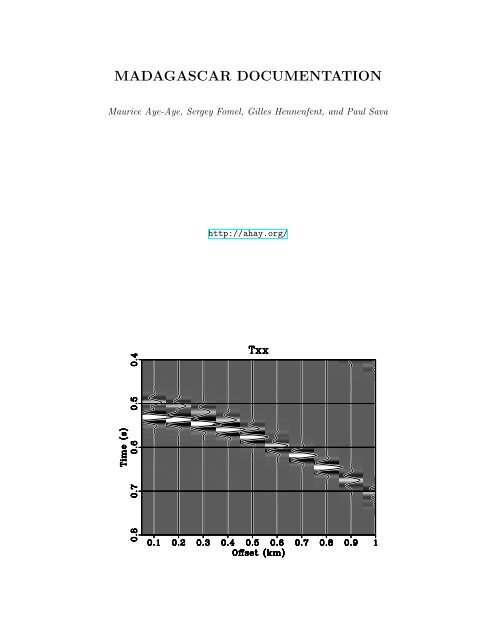
can be used in Radon domain to attenuate multiple energy. Figure 2(a) is generated by<br />
forward parabolic Radon transform while Figure 1(d) is generated by inverse parabloic<br />
Radon transform. The purpose was to make sure that forward and inverse transforms<br />
do not cause any data loss.<br />
Figure 2(a) shows the Radon transform of the CMP gather in Figure 3(a) while<br />
Figure 2(b) shows in the Radon domain the multiple energy only after mutting the<br />
primary energy. The protected multiples can be taken back to the time-offset domain<br />
and are subtracted from the data.<br />
CMP gather before multiple attenuation is shown in Figure 3(a) and the coresponding<br />
semblance scan is shown in Figure 3(c). The CMP gather after multiple<br />
attenuation is shown in Figure 3(b) and the coresponding semblance scan is shown in<br />
Figure 3(d). The semblance scans show how multiple energy is reduced for the CMP<br />
gather after multiple attenuation.
4 Maurice <strong>Madagascar</strong> Documentation<br />
Figure 1: CMP gather from Canterbury dataset before applying NMO (a), after applying<br />
NMO (b), after Forward parabolic Radon transfrom (c), after applying inverse<br />
parabolic Radon transform (d). The forward and inverse parabolic Radon transforms<br />
are applied in sequence to examine the parameters of the process and to ensure that<br />
no events are lost during the process school2012/demo cmp,nmo,taup,nmo2
<strong>Madagascar</strong> Documentation Tutorial 5<br />
Figure 2: Forward Radon transform of the gather (a). Mute is applied to preserve<br />
multiples (b); so that multiples can be transformed to time-offset domain for subtraction<br />
from the CMP gather. school2012/demo taup,taupmult<br />
1. To examine the forward and inverse Radon transform, Run<br />
bash$ scons taup_qc.view<br />
2. Edit the SConstruct <strong>file</strong> to modify the reference offset x0 for sfradon program.<br />
To get more details about sfradon parameters, run<br />
bash$ sfradon<br />
in a Unix shell. Check your result by running<br />
scons taup_qc.view<br />
3. Edit the SConstruct <strong>file</strong> to modify the starting time t0 for sfmutter. To get<br />
more details about sfmutter parameters, run sfmutter in a Unix shell. Check<br />
your result by running<br />
scons taup_mult.view<br />
4. Edit the SConstruct <strong>file</strong> to modify the starting time v0 for sfmutter. Check<br />
your result by running
6 Maurice <strong>Madagascar</strong> Documentation<br />
Figure 3: CMP gather before multiple attenuation (a). CMP gather after<br />
multiple attenuation (b). Gather in (a) is used to generated semblance<br />
scan in (c). Gather in (b) is used to generate semblance scan in (d).<br />
school2012/demo cmp,signal2,vscan-cmp,vscan-signal2
<strong>Madagascar</strong> Documentation Tutorial 7<br />
scons taup_mult.view<br />
5. Edit the SConstruct <strong>file</strong> and find the line that says ADD CODE to create<br />
signal2.vpl. To get more details about sfgrey parameters, run sfgrey in a<br />
Unix shell. Add your code and create the vpl <strong>file</strong> by running<br />
scons signal2.vpl<br />
6. Edit the SConstruct <strong>file</strong> and find the line that says ADD CODE to display<br />
cmp.vpl and signal2.vpl. Add your code and view the <strong>file</strong> by running<br />
scons cmp_signal2.view<br />
7. Edit the SConstruct <strong>file</strong> and find the line that says ADD CODE to display<br />
vscan-cmp.vpl and vscan-signal2.vpl. Add your code and view the <strong>file</strong> by<br />
running<br />
scons v_cmp_signal2.view<br />
1 from r s f . p r o j import ∗<br />
2<br />
3 # download cmp1 . r s f from the s e r v e r<br />
4 Fetch ( ’cmp1 . r s f ’ , ’ cant12 ’ )<br />
5<br />
6 # convert to n a t i v e format<br />
7 Flow ( ’cmp ’ , ’cmp1 ’ , ’ dd form=n a t i v e ’ )<br />
8<br />
9 # c r e a t e cmp . v p l f i l e<br />
10 Plot ( ’cmp ’ , ’ grey t i t l e=CMP ’ )<br />
11<br />
12 # water v e l o c i t y 1500 m/ s<br />
13 wvel=1500<br />
14<br />
15 # NMO with water v e l o c i t y<br />
16 Flow ( ’nmo ’ , ’cmp ’ , ’ nmostretch h a l f=n v0=%g ’%wvel )<br />
17<br />
18 # c r e a t e nmo. v p l<br />
19 Plot ( ’nmo ’ , ’ grey t i t l e=NMO’ )<br />
20<br />
21 # c r e a t e cmp nmo . v p l f i l e under Fig d i r e c t o r y<br />
22 # cmp . v p l and nmo. v p l c r e a t e d e a r l i e r using Plot<br />
23 # command w i l l be p l o t e d s i d e by s i d e<br />
24 Result ( ’cmp nmo ’ , ’cmp nmo ’ , ’ SideBySideAniso ’ )<br />
25<br />
26 ####################
8 Maurice <strong>Madagascar</strong> Documentation<br />
27 # radon parameters<br />
28 ####################<br />
29 ox =29.25<br />
30 nx=60<br />
31 dx=25<br />
32 #−−−−−−−−−−−−−−−−−−−−−<br />
33 x0=800 # CHANGE ME<br />
34 #−−−−−−−−−−−−−−−−−−−−−<br />
35 p0=−.05<br />
36 dp=.0005<br />
37 np=201<br />
38<br />
39 # forward Radon operator<br />
40 radono=’ ’ ’<br />
41 radon np=%d p0=%f dp=%f x0=%d parab=y<br />
42 ’ ’ ’ %(np , p0 , dp , x0 )<br />
43<br />
44 # i n v e r s e Radon operator<br />
45 radonoinv=’ ’ ’<br />
46 radon adj=n nx=%d ox=%g dx=%d x0=%d parab=y<br />
47 ’ ’ ’ %(nx , ox , dx , x0 )<br />
48<br />
49 # Test radon parameters , apply forward and<br />
50 # i n v e r s e Radon Transform , and QC r e s u l t s<br />
51 #########################################<br />
52 Flow ( ’ taup ’ , ’nmo ’ , radono )<br />
53<br />
54 # p l o t<br />
55 Plot ( ’ taup ’ , ’ grey t i t l e=forward RT ’ )<br />
56<br />
57 # I n v e r s e<br />
58 Flow ( ’nmo2 ’ , ’ taup ’ , radonoinv )<br />
59<br />
60 # p l o t<br />
61 Plot ( ’nmo2 ’ , ’ grey t i t l e=i n v e r s e RT ’ )<br />
62<br />
63 # Display t h r e e f i g u r e s to QC Radon parameters<br />
64 # Check t h a t forward and i n v e r s e Radon transforms<br />
65 # do not change the data i . e e v e n t s are p r e s e r v e d .<br />
66<br />
67 Result ( ’ taup qc ’ , ’nmo taup nmo2 ’ , ’ SideBySideAniso ’ )<br />
68<br />
69 ######################################<br />
70 # design a mute f u n c t i o n t h a t p r o t e c t s<br />
71 # m u l t i p l e s in the Radon domain
<strong>Madagascar</strong> Documentation Tutorial 9<br />
72 ######################################<br />
73 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
74 t0 =1.2 # CHANGE ME ; t r y 1.5<br />
75 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
76 # v e r t i c a l p o s i t i o n o f the t r i a n g l e v e r t i x<br />
77<br />
78 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
79 v0=.03 # CHANGE ME ; t r y .015<br />
80 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
81 # s l o p e o f the t r i a n g l e<br />
82<br />
83 Flow ( ’ taupmult ’ , ’ taup ’ , ’ mutter t0=%g v0=%g ’%(t0 , v0 ) )<br />
84 Plot ( ’ taupmult ’ , ’ grey t i t l e =”m u l t i p l e s in Radon domain” ’ )<br />
85<br />
86 # Display taup . v p l and taupmult . v p l<br />
87 # This d i s p l a y a l l o w s a f l i p between<br />
88 # the two f i g u r e s<br />
89 Result ( ’ taup mult ’ , ’ taup taupmult ’ ,<br />
90 ’ ’ ’<br />
91 cat a x i s=3 ${SOURCES[ 1 ] }<br />
92 | grey<br />
93 ’ ’ ’ )<br />
94<br />
95 # Transform m u l i t p l e s from Radon domain to time−o f f s e t domain<br />
96 Flow ( ’ m u l t i p l e ’ , ’ taupmult ’ , radonoinv )<br />
97<br />
98 # c r e a t e m u l t i p l e . v p l<br />
99 Plot ( ’ m u l t i p l e ’ , ’ grey t i t l e =”m u l t i p l e s ” ’ )<br />
100<br />
101 # p l o t CMP and m u l t i p l e s s i d e by s i d e<br />
102 Result ( ’ cmp mult ’ , ’nmo2 m u l t i p l e ’ , ’ SideBySideAniso ’ )<br />
103<br />
104 # S u b t r a c t m u l t i p l e s from the CMP<br />
105 Flow ( ’ s i g n a l ’ , ’ m u l t i p l e nmo2 ’ ,<br />
106 ’ ’ ’<br />
107 add s c a l e =−1,1 ${SOURCES[ 1 ] }<br />
108 ’ ’ ’ )<br />
109<br />
110 # i n v e r s e NMO<br />
111 Flow ( ’ s i g n a l 2 ’ , ’ s i g n a l ’ ,<br />
112 ’ ’ ’<br />
113 nmostretch inv=y h a l f=n v0=%g<br />
114 | mutter v0=1900 x0=200<br />
115 ’ ’ ’%wvel )<br />
116
10 Maurice <strong>Madagascar</strong> Documentation<br />
117 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
118 # ADD CODE to c r e a t e s i g n a l 2 . v p l<br />
119 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
120<br />
121<br />
122 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
123 # ADD CODE to d i s p l a y cmp . v p l and s i g n a l 2 . vpl ,<br />
124 # make the f i g u r e s f l i p back and f o r t h so you<br />
125 # can examine the the r e s u l t s of m u l t i p l e<br />
126 # a t t e n u a t i o n . Let us c a l l the output f i l e<br />
127 # cmp signal2<br />
128 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
129<br />
130<br />
131 ####################<br />
132 # Semblance Scan<br />
133 ####################<br />
134 dv=10<br />
135 nv=251<br />
136 v0=1400<br />
137 vscan=’ vscan v0=%d dv=%d nv=%d semblance=y h a l f=n ’%(v0 , dv , nv )<br />
138 pick=’ pick r e c t 1 =150 r e c t 2 =50 gate=20 ’<br />
139<br />
140 # semblance scan<br />
141 Flow ( ’ vscan−cmp ’ , ’cmp ’ , vscan )<br />
142<br />
143 # semblance scan<br />
144 Flow ( ’ vscan−s i g n a l 2 ’ , ’ s i g n a l 2 ’ , vscan )<br />
145<br />
146 Plot ( ’ vscan−cmp ’ ,<br />
147 ’ ’ ’<br />
148 grey c o l o r=j a l l p o s=y<br />
149 t i t l e =”V e l o c i t y Scan − CMP”<br />
150 ’ ’ ’ )<br />
151<br />
152 Plot ( ’ vscan−s i g n a l 2 ’ ,<br />
153 ’ ’ ’<br />
154 grey c o l o r=j a l l p o s=y<br />
155 t i t l e =”V e l o c i t y Scan − a f t e r demultiple ”<br />
156 ’ ’ ’ )<br />
157<br />
158 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
159 # ADD CODE to d i s p l a y the two f i g u r e s<br />
160 # vscan−cmp . v p l and vscan−s i g n a l 2 . v p l<br />
161 # s i d e by s i d e . Let us c a l l the output
<strong>Madagascar</strong> Documentation Tutorial 11<br />
162 # f i l e vcmp−s i g n a l 2<br />
163 #−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−−<br />
164<br />
165<br />
166<br />
167 ###################################################<br />
168 # This part i s to c r e a t e f i g u r e s f o r t u t o r i a l . pdf<br />
169 ###################################################<br />
170 # d e f i n e grey commands f o r f i g u r e s to be i n c l u d e d<br />
171 # in t u t o r i a l . pdf<br />
172 grey=’ ’ ’<br />
173 grey w a n t t i t l e=n l a b e l f a t =2 t i t l e f a t =2<br />
174 x l l =2 y l l =1.5 yur=9 xur=6<br />
175 ’ ’ ’<br />
176<br />
177 greyc=’ ’ ’<br />
178 grey w a n t t i t l e=n l a b e l f a t =2 t i t l e f a t =2<br />
179 x l l =2 y l l =1.5 yur=9 xur=6<br />
180 c o l o r=j a l l p o s=y<br />
181 ’ ’ ’<br />
182 # c r e a t e p l o t s<br />
183 Result ( ’cmp ’ , grey )<br />
184 Result ( ’nmo ’ , grey )<br />
185 Result ( ’ taup ’ , grey )<br />
186 Result ( ’nmo2 ’ , grey )<br />
187 Result ( ’ taupmult ’ , grey )<br />
188 Result ( ’ s i g n a l 2 ’ , grey )<br />
189 Result ( ’ vscan−cmp ’ , greyc )<br />
190 Result ( ’ vscan−s i g n a l 2 ’ , greyc )<br />
191<br />
192 End ( )<br />
EXERCISE<br />
In this part, your task is to apply the workflow explained above to a different CMP<br />
gather that requires different parameters. The same workflow should work here, but<br />
you need to observe that the CMP gather used for this exercise has shallow events.<br />
This means that, after applying NMO correction, amplitudes at far offstes of the<br />
shallow events get stretched. Therefore, an additional step is required for this CMP.<br />
We need to mute the distorted amplitudes. The mute is already applied in the<br />
SConstruct.<br />
1. Display the CMP gather after NMO with and without mute applied by running
12 Maurice <strong>Madagascar</strong> Documentation<br />
scons nmo1_nmo.view<br />
2. Your task is to add the necessary code to attenuate multiples for this CMP.<br />
The same work flow used in the SConstruct under demo directory should work<br />
here with only changes to<br />
• x0<br />
• t0<br />
• v0<br />
where it says CHANGE ME in the comments<br />
WRITING A REPORT<br />
1. Change directory to the parent directory<br />
bash$ cd ..<br />
This should be the directory that contains tutorial.tex.<br />
2. Run<br />
bash$ sftour scons lock<br />
The sftour command visits all subdirectories and runs scons lock, which<br />
copies result <strong>file</strong>s to a different location so that they do not get modified until<br />
further notice.<br />
3. You can also run<br />
bash$ sftour scons -c<br />
to clean intermediate results.<br />
4. Edit the <strong>file</strong> paper.tex to include your additional results. If you have not used<br />
L A TEX before, no worries. It is a descriptive language. Study the <strong>file</strong>, and it<br />
should become evident by example how to include figures.<br />
5. Run<br />
bash$ scons tutorial.pdf<br />
and open tutorial.pdf with a <strong>PDF</strong> viewing program such as Acrobat Reader.<br />
6. If you have L A TEX2HTML installed, you can also generate an HTML version of<br />
your paper by running<br />
bash$ scons tutorial.html<br />
and opening tutorial_html/index.html in a web browser.
<strong>Madagascar</strong> Documentation Tutorial 13<br />
REFERENCES<br />
Hampson, D., 1986, Inverse velocity stacking for multiple elimination: J. Can. Soc.<br />
Expl. Geophys, 22, 44–55.<br />
Lu, H., C. S. Fulthorpe, and P. Mann, 2003, Three-dimensional architecture of shelfbuilding<br />
sediment drifts in the offshore canterbury basin, new zealand: Marine<br />
Geology, 193, 19 – 47.
14 Maurice <strong>Madagascar</strong> Documentation
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Seismic Imaging Tutorial:<br />
“exploding reflector” modeling/migration<br />
Paul Sava<br />
Center for Wave Phenomena<br />
Colorado School of Mines 1<br />
ABSTRACT<br />
This document demonstrates how reproducible numeric experiments constructed<br />
using the madagascar software package can be integrated into a document<br />
generated using the L A TEXtypesetting program. I use a simple modeling/migration<br />
exercise based on the exploding reflector model to illustrate reproducible<br />
document generation.<br />
INTRODUCTION<br />
Acoustic modeling and migration can be implemented using numeric solutions to the<br />
acoustic wave-equation (Clærbout, 1985):<br />
( )<br />
1<br />
1<br />
v Ẅ − ρ∇ · 2 ρ ∇W = f . (1)<br />
In Equation 1, W (x, t) represents the acoustic wavefield, v (x) and ρ (x) represent<br />
the velocity and density of the medium, respectively, and f (x, t) represents a source<br />
function.<br />
• In modeling, we use the distributed source f (x, t) to generate the wavefield<br />
W (x, t) at all positions and all times by wave propagation forward in time.<br />
The data represent a subset of the wavefield observed at receivers distributed<br />
in the medium: D (r, t) = W (x = r, t).<br />
• In migration, we use the observed data D (r, t) to generate the wavefield W (x, t)<br />
at all positions and all times by wave propagation backward in time. The image<br />
represents a subset of the wavefield at time zero: R (x) = W (x =, t = 0).<br />
In both cases, we solve Equation 1 with different initial conditions, but with the same<br />
model, v (x) and ρ (x) and with the same boundary conditions.<br />
1 e-mail: psava@mines.edu<br />
15
16 Sava <strong>Madagascar</strong> Documentation<br />
EXAMPLE<br />
I illustrate the method using the Sigsbee 2A synthetic model. This model is based on<br />
the Sigsbee structure in the Gulf of Mexico and the velocity model is illustrated in<br />
Figure 1. The model is characterized by a massive salt body close to the water bottom<br />
and surrounded by sediments. The salt velocity is 4.5 km/s and the surrounding<br />
sediment velocity ranges from approximately 1.5 to 3.25 km/s.<br />
Figure 1: Stratigraphic<br />
Sigsbee 2A velocity model<br />
school/sigsbee vstr<br />
In this experiment, I consider sources distributed uniformly in the subsalt region<br />
of the model. The data are acquired in a borehole array, located at x = 8.5 km and<br />
a horizontal array located at z = 1.5 km. In order to avoid multiple scattering in the<br />
subsurface, I simulate waves in a smooth version of the Sigsbee model, illustrated in<br />
Figure 2, and constant density.<br />
Figure 2: Smooth Sigsbee<br />
2A velocity model<br />
school/sigsbee vsmo<br />
Using the madagascar program sfawefd2d, we can simulate wavefields from the<br />
distributed sources. Figures 3(a)-3(h) show wavefield snapshots in order of increasing<br />
times. We can observe waves propagating from all subsalt sources, interacting with<br />
the variable velocity medium and arriving at the vertical and horizontal arrays.<br />
Figures 4(a) and 4(b) show the data observed at the horizontal array in variable<br />
density and wiggle plotting formats, respectively. Similarly, Figures 5(a) and 5(b)<br />
show the data observed in the vertical array using the same plotting formats. The<br />
data are just subsets of the same wavefields at the respective receiver positions and<br />
capture the complications observed in the wavefield, i.e. triplications due to lateral<br />
velocity variation.
<strong>Madagascar</strong> Documentation WSI tutorial 17<br />
Figure 3: Wavefield snapshots at increasing times.<br />
school/sigsbee wfld-01,wfld-03,wfld-05,wfld-07,wfld-09,wfld-11,wfld-13,wfld-15
18 Sava <strong>Madagascar</strong> Documentation<br />
Figure 4: Acoustic data observed in the horizontal array. school/sigsbee datH,wigH
<strong>Madagascar</strong> Documentation WSI tutorial 19<br />
Figure 5: Acoustic data observed in the vertical array. school/sigsbee datV,wigV
20 Sava <strong>Madagascar</strong> Documentation<br />
In zero-offset migration, we use the observed data to backprogate the wavefields<br />
using the data as boundary conditions. The image is the wavefield at time zero.<br />
Since we can acquire data at different locations in space, the reconstructed wavefields<br />
depend on the acquisition geometry, thus limiting the illumination in the subsurface.<br />
Therefore, the migrated images depend on the acquisition array, as illustrated in<br />
Figures 6(a) and 6(b) for the horizontal and vertical arrays, respectively. We can<br />
also obtain images by migrating the data observed in both the horizontal and vertical<br />
arrays, as illustrated in Figure 7, thus increasing the acquisition aperture and the<br />
subsurface illumination.<br />
CONCLUSIONS<br />
The combination of L A TEX and madagascar allows geoscientists to generate reproducible<br />
documents where the numeric examples can be verified by any user with the<br />
same computer setup. This allows for transparent peer-review, for recursive development<br />
and for transfer of technology between collaborative research groups.<br />
ACKNOWLEDGMENTS<br />
The reproducible numeric examples in this paper use the madagascar open-source<br />
software package freely available from http://www.reproducibility.org.<br />
REFERENCES<br />
Clærbout, J. F., 1985, Imaging the Earth’s interior: Blackwell Scientific Publications.
<strong>Madagascar</strong> Documentation WSI tutorial 21<br />
Figure 6: Migrated images for data acquired in (a) the horizontal array and (b) the<br />
vertical array. school/sigsbee imgH,imgV
22 Sava <strong>Madagascar</strong> Documentation<br />
Figure 7: Migrated image for data acquired in the horizontal and the vertical arrays.<br />
school/sigsbee imgA
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
<strong>Madagascar</strong> tutorial<br />
Maurice the Aye-Aye 1<br />
ABSTRACT<br />
In this tutorial, you will go through different steps required for writing a research<br />
paper with reproducible examples. In particular, you will<br />
1. identify a research problem,<br />
2. suggest a solution,<br />
3. test your solution using a synthetic example,<br />
4. apply your solution to field data,<br />
5. write a report about your work.<br />
Completing this tutorial requires<br />
PREREQUISITES<br />
• <strong>Madagascar</strong> software environment available from<br />
http://www.ahay.org<br />
• L A TEX environment with SEGTeX available from<br />
http://www.ahay.org/wiki/SEGTeX<br />
To do the assignment on your personal computer, you need to install the required<br />
environments. An Internet connection is required for access to the data repository.<br />
The tutorial itself is available from the <strong>Madagascar</strong> repository by running<br />
svn co https://rsf.svn.sourceforge.net/svnroot/rsf/trunk/book/rsf/school2009<br />
INTRODUCTION<br />
In this tutorial, you will be asked to run commands from the Unix shell (identified<br />
by bash$) and to edit <strong>file</strong>s in a text editor. Different editors are available in a typical<br />
Unix environment (vi, emacs, nedit, etc.)<br />
1 e-mail: psava@mines.edu<br />
23
24 Maurice <strong>Madagascar</strong> Documentation<br />
Your first assignment:<br />
1. Open a Unix shell.<br />
2. Change directory to the tutorial directory<br />
bash$ cd $RSFSRC/book/rsf/school2009<br />
3. Open the paper.tex <strong>file</strong> in your favorite editor, for example by running<br />
bash$ nedit paper.tex &<br />
4. Look at the first line in the <strong>file</strong> and change the author name from Maurice the<br />
Aye-Aye to your name (first things first).<br />
PROBLEM<br />
Figure 1: Depth slice from 3-D seismic (left) and output of edge detection (right).<br />
school2009/channel horizon<br />
The left plot in Figure 1 shows a depth slice from a 3-D seismic volume 2 . You<br />
notice a channel structure and decide to extract it using and edge detection algorithm<br />
from the image processing literature (Canny, 1986). In a nutshell, Canny’s edge<br />
detector picks areas of high gradient that seem to be aligned along an edge. The<br />
extracted edges are shown in the right plot of Figure 1. The initial result is not too<br />
clear, because it is affected by random fluctuations in seismic amplitudes. The goal<br />
of your research project is to achieve a better result in automatic channel extraction.<br />
1. Change directory to the project directory<br />
2 Courtesy of Matt Hall (ConocoPhillips Canada Ltd.)
<strong>Madagascar</strong> Documentation Tutorial 25<br />
bash$ cd channel<br />
2. Run<br />
bash$ scons horizon.view<br />
in the Unix shell. A number of commands will appear in the shell followed by<br />
Figure 1 appearing on your screen.<br />
3. To understand the commands, examine the script that generated them by opening<br />
the SConstruct <strong>file</strong> in a text editor. Notice that, instead of Shell commands,<br />
the script contains rules.<br />
• The first rule, Fetch, allows the script to download the input data <strong>file</strong><br />
horizon.asc from the data server.<br />
• Other rules have the form Flow(target,source,command) for generating<br />
data <strong>file</strong>s or Plot and Result for generating picture <strong>file</strong>s.<br />
• Fetch, Flow, Plot, and Result are defined in <strong>Madagascar</strong>’s rsf.proj<br />
package, which extends the functionality of SCons (Fomel and Hennenfent,<br />
2007).<br />
4. To better understand how rules translate into commands, run<br />
bash$ scons -c horizon.rsf<br />
The -c flag tells scons to remove the horizon.rsf <strong>file</strong> and all its dependencies.<br />
5. Next, run<br />
bash$ scons -n horizon.rsf<br />
The -n flag tells scons not to run the command but simply to display it on the<br />
screen. Identify the lines in the SConstruct <strong>file</strong> that generate the output you<br />
see on the screen.<br />
6. Run<br />
bash$ scons horizon.rsf<br />
Examine the <strong>file</strong> horizon.rsf both by opening it in a text editor and by running<br />
bash$ sfin horizon.rsf<br />
How many different <strong>Madagascar</strong> modules were used to create this <strong>file</strong>? What<br />
are the <strong>file</strong> dimensions? Where is the actual data stored?
26 Maurice <strong>Madagascar</strong> Documentation<br />
7. Run<br />
bash$ scons smoothed.rsf<br />
Notice that the horizon.rsf <strong>file</strong> is not being rebuilt.<br />
8. What does the sfsmooth module do? Find it out by running<br />
bash$ sfsmooth<br />
without arguments. Has sfsmooth been used in any other <strong>Madagascar</strong> examples?<br />
9. What other <strong>Madagascar</strong> modules perform smoothing? To find out, run<br />
bash$ sfdoc -k smooth<br />
10. Notice that Figure 1 does not make a very good use of the color scale. To<br />
improve the scale, find the mean value of the data by running<br />
bash$ sfattr < horizon.rsf<br />
and insert it as a new value for the bias= parameter in the SConstruct <strong>file</strong>.<br />
Does smoothing by sfsmooth change the mean value?<br />
11. Save the SConstruct <strong>file</strong> and run<br />
bash$ scons view<br />
to view improved images. Notice that horizon.rsf and smoothed.rsf <strong>file</strong>s are<br />
not being rebuilt. SCons is smart enough to know that only the part affected<br />
by your changes needs to be updated.<br />
As shown in Figure 2, smoothing removes random amplitude fluctuations but at<br />
the same broadens the channel and thus makes the channel edge detection unreliable.<br />
In the next part of this tutorial, you will try to find a better solution by examining<br />
a simple one-dimensional synthetic example.<br />
1 from r s f . p r o j import ∗<br />
2<br />
3 # Download data<br />
4 Fetch ( ’ horizon . asc ’ , ’ h a l l ’ )<br />
5<br />
6 # Convert format<br />
7 Flow ( ’ horizon ’ , ’ horizon . asc ’ ,<br />
8 ’ ’ ’
<strong>Madagascar</strong> Documentation Tutorial 27<br />
Figure 2: Depth slice from Figure 1 after smoothing (left) and output of edge detection<br />
(right). school2009/channel smoothed<br />
9 echo in=$SOURCE data format=a s c i i f l o a t n1=3 n2=57036 |<br />
10 dd form=n a t i v e | window n1=1 f 1=−1 |<br />
11 put<br />
12 n1=196 o1 =33.139 d1=0.01 l a b e l 1=y unit1=km<br />
13 n2=291 o2 =35.031 d2=0.01 l a b e l 2=x unit2=km<br />
14 ’ ’ ’ )<br />
15<br />
16 # Triangle smoothing<br />
17 Flow ( ’ smoothed ’ , ’ horizon ’ , ’ smooth r e c t 1 =20 r e c t 2 =20 ’ )<br />
18<br />
19 # Display r e s u l t s<br />
20 for horizon in ( ’ horizon ’ , ’ smoothed ’ ) :<br />
21 # −−− CHANGE BELOW −−−<br />
22 Plot ( horizon , ’ grey c o l o r=j b i a s=0 y r e v e r s e=n w a n t t i t l e=n ’ )<br />
23 edge = ’ edge−’+horizon<br />
24 Flow ( edge , horizon , ’ canny max=98 | dd type=f l o a t ’ )<br />
25 Plot ( edge , ’ grey a l l p o s=y y r e v e r s e=n w a n t t i t l e=n ’ )<br />
26 Result ( horizon , [ horizon , edge ] , ’ SideBySideIso ’ )<br />
27<br />
28 End ( )<br />
1-D SYNTHETIC<br />
To better understand the effect of smoothing, you decide to create a one-dimensional<br />
synthetic example shown in Figure 3(a). The synthetic contains both sharp edges and<br />
random noise. The output of conventional triangle smoothing is shown in Figure 3(b).<br />
We see an effect similar to the one in the real data example: random noise gets
28 Maurice <strong>Madagascar</strong> Documentation<br />
Figure 3: (a) 1-D synthetic to test edge-preserving smoothing. (b) Output of conventional<br />
triangle smoothing. school2009/local step,smooth<br />
removed by smoothing at the expense of blurring the edges. Can you do better?<br />
Figure 4: (a) Input synthetic trace duplicated multiple times. (b) Duplicated traces<br />
shifted so that each data sample gets surrounded by its neighbors. The original trace<br />
is in the middle. school2009/local spray,local<br />
To better understand what is happening in the process of smoothing, let us convert<br />
1-D signal into a 2-D signal by first replicating the trace several times and then shifting<br />
the replicated traces with respect to the original trace (Figure 4). This creates a 2-<br />
D dataset, where each sample on the original trace is surrounded by samples from<br />
neighboring traces.<br />
Every local filtering operation can be understood as stacking traces from Figure<br />
4(b) multiplied by weights that correspond to the filter coefficients.<br />
1. Change directory to the project directory<br />
bash$ cd ../local
<strong>Madagascar</strong> Documentation Tutorial 29<br />
2. Verify the claim above by running<br />
bash$ scons smooth.view smooth2.view<br />
Open the SConstruct <strong>file</strong> in a text editor to verify that the first image is<br />
computed by sfsmooth and the second image is computed by applying triangle<br />
weights and stacking. To compare the two images by flipping between them,<br />
run<br />
bash$ sfpen Fig/smooth.vpl Fig/smooth2.vpl<br />
3. Edit SConstruct to change the weight from triangle<br />
to Gaussian<br />
W T (x) = 1 − |x|<br />
x 0<br />
(1)<br />
)<br />
W G (x) = exp<br />
(−α |x|2<br />
Repeat the previous computation. Does the result change? What is a good<br />
value for α?<br />
4. Thinking about this problem, you invent an idea 3 . Why not apply non-linear<br />
filter weights that would discriminate between points not only based on their<br />
distance from the center point but also on the difference in function values<br />
between the points. That is, instead of filtering by<br />
∫<br />
g(x) = f(y) W (x − y) dy , (3)<br />
where f(x) is input, g(y) is output, and W (x) is a linear weight, you decide to<br />
filter by<br />
∫<br />
g(x) = f(y) Ŵ (x − y, f(x) − f(y)) dy , (4)<br />
where and Ŵ (x, z) is a non-linear weight. Compare the two weights by running<br />
bash$ scons triangle.view similarity.view<br />
The results should look similar to Figure 5.<br />
5. The final output is Figure 6. By examining SConstruct, find how to reproduce<br />
this figure.<br />
6. EXTRA CREDIT If you are familiar with programming in C, add 1-D nonlocal<br />
filtering as a new <strong>Madagascar</strong> module sfnonloc. Ask the instructor<br />
for further instructions.<br />
x 2 0<br />
(2)
30 Maurice <strong>Madagascar</strong> Documentation<br />
Figure 5: (a) Linear and stationary triangle weights. (b) Non-linear and nonstationary<br />
weights reflecting both distance between data points and similarity in<br />
data values. school2009/local triangle,similarity<br />
Figure 6: Output of<br />
non-local<br />
smoothing<br />
school2009/local nlsmooth
<strong>Madagascar</strong> Documentation Tutorial 31<br />
Figure 6 shows that non-linear filtering can eliminate random noise while preserving<br />
the edges. The problem is solved! Now let us apply the result to our original<br />
problem.<br />
1 /∗ Non−l o c a l smoothing . ∗/<br />
2 #include <br />
3<br />
4 int main ( int argc , char ∗ argv [ ] )<br />
5 {<br />
6 int n1 , n2 , i1 , i2 , i s , ns ;<br />
7 float ∗ trace , ∗ trace2 , ax , ay , t ;<br />
8 s f f i l e inp , out ;<br />
9<br />
10 /∗ i n i t i a l i z e ∗/<br />
11 s f i n i t ( argc , argv ) ;<br />
12<br />
13 /∗ s e t input and output f i l e s ∗/<br />
14 inp = s f i n p u t ( ” in ” ) ;<br />
15 out = s f o u t p u t ( ” out ” ) ;<br />
16<br />
17 /∗ g e t input dimensions ∗/<br />
18 i f ( ! s f h i s t i n t ( inp , ”n1”,&n1 ) )<br />
19 s f e r r o r ( ”No n1= in input ” ) ;<br />
20 n2 = s f l e f t s i z e ( inp , 1 ) ;<br />
21<br />
22 /∗ g e t command−l i n e parameters ∗/<br />
23 i f ( ! s f g e t i n t ( ” ns ”,&ns ) ) s f e r r o r ( ”Need ns=” ) ;<br />
24 /∗ spray r a d i u s ∗/<br />
25<br />
26 i f ( ! s f g e t f l o a t ( ”ax”,&ax ) ) s f e r r o r ( ”Need ax=” ) ;<br />
27 /∗ e x p o n e n t i a l weight f o r the c o o r d i n a t e d i s t a n c e ∗/<br />
28<br />
29 t r a c e = s f f l o a t a l l o c ( n1 ) ;<br />
30 t r a c e 2 = s f f l o a t a l l o c ( n1 ) ;<br />
31<br />
32 /∗ loop over t r a c e s ∗/<br />
33 for ( i 2 =0; i 2 < n2 ; i 2++) {<br />
34 /∗ read input ∗/<br />
35 s f f l o a t r e a d ( trace , n1 , inp ) ;<br />
36<br />
37 /∗ loop over samples ∗/<br />
38 for ( i 1 =0; i 1 < n1 ; i 1++) {<br />
39 t = 0 . ;<br />
3 Actually, you reinvent the idea of bilateral or non-local filters (Tomasi and Manduchi, 1998;<br />
Gilboa and Osher, 2008).
32 Maurice <strong>Madagascar</strong> Documentation<br />
40<br />
41 /∗ accumulate s h i f t s ∗/<br />
42 for ( i s=−ns ; i s = 0 && i 1+i s < n1 ) {<br />
44<br />
45 /∗ ! ! ! MODIFY THE NEXT LINE ! ! ! ∗/<br />
46 t += t r a c e [ i 1+i s ] ∗ expf(−ax∗ i s ∗ i s ) ;<br />
47 }<br />
48 }<br />
49<br />
50 t r a c e 2 [ i 1 ] = t ;<br />
51 }<br />
52<br />
53 /∗ w r i t e output ∗/<br />
54 s f f l o a t w r i t e ( trace2 , n1 , out ) ;<br />
55 }<br />
56<br />
57 /∗ clean up ∗/<br />
58 s f f i l e c l o s e ( inp ) ;<br />
59 e x i t ( 0 ) ;<br />
60 }<br />
SOLUTION<br />
1. Change directory to the project directory<br />
bash$ cd ../channel2<br />
2. By now, you should know what to do next.<br />
3. Two-dimensional shifts generate a four-dimensional volume. Verify it by running<br />
bash$ scons local.rsf<br />
and<br />
bash$ sfin local.rsf<br />
View a movie of different shifts by running<br />
bash$ scons local.vpl<br />
4. Modify the filter weights by editing SConstruct in a text editor. Observe your<br />
final result by running
<strong>Madagascar</strong> Documentation Tutorial 33<br />
bash$ scons smoothed2.view<br />
5. The <strong>file</strong> norm.rsf contains the non-linear weights stacked over different shifts.<br />
Add a Result statement to SConstruct that would display the contents of<br />
norm.rsf in a figure. Do you notice anything interesting?<br />
6. Apply the Canny edge detection to your final result and display it in a figure.<br />
7. EXTRA CREDIT Change directory to ../mona and apply your method to<br />
the image of Mona Lisa. Can you extract her smile?<br />
1 from r s f . p r o j import ∗<br />
2<br />
3 # Download data<br />
4 Fetch ( ’ horizon . asc ’ , ’ h a l l ’ )<br />
5<br />
6 # Convert format<br />
7 Flow ( ’ horizon2 ’ , ’ horizon . asc ’ ,<br />
8 ’ ’ ’<br />
9 echo in=$SOURCE data format=a s c i i f l o a t n1=3 n2=57036 |<br />
10 dd form=n a t i v e | window n1=1 f 1=−1 |<br />
11 add add=−65 | put<br />
12 n1=196 o1 =33.139 d1=0.01 l a b e l 1=y unit1=km<br />
13 n2=291 o2 =35.031 d2=0.01 l a b e l 2=x unit2=km<br />
14 ’ ’ ’ , s t d i n =0)<br />
15 Result ( ’ horizon2 ’ , ’ grey y r e v e r s e=n c o l o r=j t i t l e=Input ’ )<br />
16<br />
17 # Spray<br />
18 Flow ( ’ spray ’ , ’ horizon2 ’ ,<br />
19 ’ ’ ’<br />
20 spray a x i s=3 n=21 o=−0.1 d=0.01 |<br />
21 spray a x i s=4 n=21 o=−0.1 d=0.01<br />
22 ’ ’ ’ )<br />
23<br />
24 # S h i f t<br />
25 Flow ( ’ s h i f t 1 ’ , ’ spray ’ , ’ window n1=1 | math output=x2 ’ )<br />
26 Flow ( ’ s h i f t 2 ’ , ’ spray ’ , ’ window n2=1 | math output=x3 ’ )<br />
27<br />
28 Flow ( ’ l o c a l ’ , ’ spray s h i f t 1 s h i f t 2 ’ ,<br />
29 ’ ’ ’<br />
30 d a t s t r e t c h datum=${SOURCES[ 1 ] } | transp |<br />
31 d a t s t r e t c h datum=${SOURCES[ 2 ] } | transp<br />
32 ’ ’ ’ )<br />
33 Plot ( ’ l o c a l ’ , ’ window j 3=4 j 4=4 | grey c o l o r=j ’ , view=1)<br />
34
34 Maurice <strong>Madagascar</strong> Documentation<br />
35 # −−− CHANGE BELOW −−−<br />
36 # t r y ” exp ( −0.1∗( input−l o c )ˆ2 −200∗( x3ˆ2+x4 ˆ2))”<br />
37 Flow ( ’ s i m i l ’ , ’ spray l o c a l ’ ,<br />
38 ’ ’ ’<br />
39 math l o c=${SOURCES[ 1 ] } output=1<br />
40 ’ ’ ’ )<br />
41<br />
42 Flow ( ’ norm ’ , ’ s i m i l ’ ,<br />
43 ’ stack a x i s=4 | stack a x i s=3 ’ )<br />
44<br />
45 Flow ( ’ smoothed2 ’ , ’ l o c a l s i m i l norm ’ ,<br />
46 ’ ’ ’<br />
47 add mode=p ${SOURCES[ 1 ] } |<br />
48 stack a x i s=4 | stack a x i s=3 |<br />
49 add mode=d ${SOURCES[ 2 ] }<br />
50 ’ ’ ’ )<br />
51 Result ( ’ smoothed2 ’ , ’ grey y r e v e r s e=n c o l o r=j t i t l e=Output ’ )<br />
52<br />
53 End ( )<br />
Figure 7: Your final result.<br />
school2009/channel2 smoothed2<br />
1 from r s f . p r o j import ∗<br />
2<br />
3 # Download data<br />
4 Fetch ( ’mona . img ’ , ’ imgs ’ )<br />
5<br />
6 # Convert to standard format<br />
7 Flow ( ’mona ’ , ’mona . img ’ ,<br />
8 ’ ’ ’<br />
9 echo n1=512 n2=513 in=$SOURCE data format=n a t i v e u c h a r |<br />
10 dd type=f l o a t<br />
11 ’ ’ ’ , s t d i n =0)<br />
12
<strong>Madagascar</strong> Documentation Tutorial 35<br />
Figure 8: Can you apply<br />
your algorithm to Mona Lisa?<br />
school2009/mona mona<br />
13 Result ( ’mona ’ ,<br />
14 ’ ’ ’<br />
15 grey transp=n a l l p o s=y t i t l e =”Mona Lisa ”<br />
16 c o l o r=b s c r e e n r a t i o =1 wantaxis=n<br />
17 ’ ’ ’ )<br />
18<br />
19 End ( )<br />
WRITING A REPORT<br />
1. Change directory to the parent directory<br />
bash$ cd ..<br />
This should be the directory that contains paper.tex.<br />
2. Run<br />
bash$ sftour scons lock<br />
The sftour command visits all subdirectories and runs scons lock, which<br />
copies result <strong>file</strong>s to a different location so that they do not get modified until<br />
further notice.<br />
3. You can also run<br />
bash$ sftour scons -c
36 Maurice <strong>Madagascar</strong> Documentation<br />
to clean intermediate results.<br />
4. Edit the <strong>file</strong> paper.tex to include your additional results. If you have not used<br />
L A TEX before, no worries. It is a descriptive language. Study the <strong>file</strong>, and it<br />
should become evident by example how to include figures.<br />
5. Run<br />
bash$ scons paper.pdf<br />
and open paper.pdf with a <strong>PDF</strong> viewing program such as Acrobat Reader.<br />
6. Want to submit your paper to Geophysics? Edit SConstruct in the paper<br />
directory to add option=manuscript to the End statement. Then run<br />
bash$ scons paper.pdf<br />
again and display the result.<br />
7. If you have L A TEX2HTML installed, you can also generate an HTML version of<br />
your paper by running<br />
bash$ scons html<br />
and opening paper_html/index.html in a web browser.<br />
REFERENCES<br />
Canny, J., 1986, A computational approach to edge detection: IEEE Trans. Pattern<br />
Analysis and Machine Intelligence, 8, 679–714.<br />
Fomel, S., and G. Hennenfent, 2007, Reproducible computational experiments using<br />
SCons: 32nd International Conference on Acoustics, Speech, and Signal Processing<br />
(ICASSP), IEEE, 1257–1260.<br />
Gilboa, G., and S. Osher, 2008, Nonlocal operators with applications to image processing:<br />
Multiscale Model & Simulation, 7, 1005–1028.<br />
Tomasi, C., and R. Manduchi, 1998, Bilateral filtering for gray and color images:<br />
Proceedings of IEEE International Conference on Computer Vision, IEEE, 836–<br />
846.
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Guide to <strong>Madagascar</strong> programs<br />
Sergey Fomel 1<br />
ABSTRACT<br />
This guide introduces some of the most used madagascar programs and illustrates<br />
their usage with examples.<br />
MAIN PROGRAMS<br />
The source <strong>file</strong>s for these programs can be found under system/main in the <strong>Madagascar</strong><br />
distribution.<br />
1 e-mail: sergey.fomel@beg.utexas.edu<br />
37
38 Fomel <strong>Madagascar</strong> Documentation<br />
sfadd: Add, multiply, or divide RSF datasets.<br />
sfadd > out.rsf scale= add= sqrt= abs= log= exp= mode= [< <strong>file</strong>0.rsf] <strong>file</strong>1.rsf<br />
<strong>file</strong>2.rsf ...<br />
The various operations, if selected, occur in the following order:<br />
(1) Take absolute value, abs=<br />
(2) Add a scalar, add=<br />
(3) Take the natural logarithm, log=<br />
(4) Take the square root, sqrt=<br />
(5) Multiply by a scalar, scale=<br />
(6) Compute the base-e exponential, exp=<br />
(7) Add, multiply, or divide the data sets, mode=<br />
sfadd operates on integer, float, or complex data, but all the input<br />
and output <strong>file</strong>s must be of the same data type.<br />
An alternative to sfadd is sfmath, which is more versatile, but may be<br />
less efficient.<br />
bools abs= If true take absolute value [nin]<br />
floats add= Scalar values to add to each dataset [nin]<br />
bools exp= If true compute exponential [nin]<br />
bools log= If true take logarithm [nin]<br />
string mode= ’a’ means add (default), ’p’ or ’m’ means<br />
multiply, ’d’ means divide<br />
floats scale= Scalar values to multiply each dataset<br />
with [nin]<br />
bools sqrt= If true take square root [nin]<br />
sfadd is useful for combining (adding, dividing, or multiplying) several datasets.<br />
What if you want to subtract two datasets? Easy. Use the scale parameter as<br />
follows:<br />
bash$ sfadd data1.rsf data2.rsf scale=1,-1 > diff.rsf<br />
or<br />
bash$ sfadd < data1.rsf data2.rsf scale=1,-1 > diff.rsf<br />
The same task can be accomplished with the more general sfmath program:<br />
bash$ sfmath one=data1.rsf two=data2.rsf output=’one-two’ > diff.rsf<br />
or
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 39<br />
bash$ sfmath < data1.rsf two=data2.rsf output=’input-two’ > diff.rsf<br />
In both cases, the size and shape of data1.rsf and data2.rsf hypercubes should be<br />
the same, and a warning message is printed out if the the axis sampling parameters<br />
(such as o1 or d1) in these <strong>file</strong>s are different.<br />
Implementation: system/main/add.c<br />
The first input <strong>file</strong> is either in the list or in the standard input.<br />
system/main/add.c<br />
103 /∗ f i n d number o f input f i l e s ∗/<br />
104 i f ( i s a t t y ( f i l e n o ( s t d i n ) ) ) {<br />
105 /∗ no input f i l e in s t d i n ∗/<br />
106 nin =0;<br />
107 } else {<br />
108 in [ 0 ] = s f i n p u t ( ” in ” ) ;<br />
109 nin =1;<br />
110 }<br />
Collect input <strong>file</strong>s in the in array from all command-line parameters that don’t<br />
contain an “=” sign. The total number of input <strong>file</strong>s is nin.<br />
system/main/add.c<br />
112 for ( i =1; i< argc ; i++) { /∗ c o l l e c t inputs ∗/<br />
113 i f (NULL != s t r c h r ( argv [ i ] , ’=’ ) ) continue ;<br />
114 in [ nin ] = s f i n p u t ( argv [ i ] ) ;<br />
115 nin++;<br />
116 }<br />
117 i f (0==nin ) s f e r r o r ( ”no input ” ) ;<br />
118 /∗ nin = no o f input f i l e s ∗/<br />
A helper function check compat checks the compatibility of input <strong>file</strong>s.<br />
Finally, we enter the main loop, where the input data are getting read buffer by<br />
buffer and combined in the total product depending on the data type.<br />
The data combination program for floating point numbers is add float.
40 Fomel <strong>Madagascar</strong> Documentation<br />
system/main/add.c<br />
424 check compat ( s f d a t a t y p e type /∗ data type ∗/ ,<br />
425 s i z e t nin /∗ number o f f i l e s ∗/ ,<br />
426 s f f i l e ∗ in /∗ input f i l e s [ nin ] ∗/ ,<br />
427 i n t dim /∗ f i l e d i m e n s i o n a l i t y ∗/ ,<br />
428 const o f f t ∗ n /∗ dimensions [ dim ] ∗/)<br />
429 /∗ Check that the input f i l e s are compatible .<br />
430 I s s u e e r r o r for type mismatch or s i z e mismatch .<br />
431 I s s u e warning for g r i d parameters mismatch . ∗/<br />
432 {<br />
433 i n t ni , id ;<br />
434 s i z e t i ;<br />
435 f l o a t d , di , o , o i ;<br />
436 char key [ 3 ] ;<br />
437 const f l o a t t o l =1.e −5; /∗ t o l e r a n c e for comparison ∗/<br />
438<br />
439 for ( i =1; i < nin ; i++) {<br />
440 i f ( s f g e t t y p e ( in [ i ] ) != type )<br />
441 s f e r r o r ( ” type mismatch : need %d” , type ) ;<br />
442 for ( id =1; id t o l ∗ f a b s f (d ) ) )<br />
456 s f w a r n i n g ( ”%s mismatch : need %g” , key , d ) ;<br />
457 } else {<br />
458 d = 1 . ;<br />
459 }<br />
460 ( void ) s n p r i n t f ( key , 3 , ”o%d” , id ) ;<br />
461 i f ( s f h i s t f l o a t ( in [ 0 ] , key ,&o ) &&<br />
462 ( ! s f h i s t f l o a t ( in [ i ] , key ,& o i ) | |<br />
463 ( f a b s f ( oi−o ) > t o l ∗ f a b s f ( d ) ) ) )<br />
464 s f w a r n i n g ( ”%s mismatch : need %g” , key , o ) ;<br />
465 }<br />
466 }<br />
467 }
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 41<br />
system/main/add.c<br />
183 for ( nbuf /= s f e s i z e ( in [ 0 ] ) ; n s i z > 0 ; n s i z −= nbuf ) {<br />
184 i f ( nbuf > n s i z ) nbuf=n s i z ;<br />
185<br />
186 for ( j =0; j < nin ; j++) {<br />
187 c o l l e c t = ( bool ) ( j != 0 ) ;<br />
188 switch ( type ) {<br />
189 case SF FLOAT:<br />
190 s f f l o a t r e a d ( ( f l o a t ∗) bufi ,<br />
191 nbuf ,<br />
192 in [ j ] ) ;<br />
193 a d d f l o a t ( c o l l e c t ,<br />
194 nbuf ,<br />
195 ( f l o a t ∗) buf ,<br />
196 ( const f l o a t ∗) bufi ,<br />
197 cmode ,<br />
198 s c a l e [ j ] ,<br />
199 add [ j ] ,<br />
200 a b s f l a g [ j ] ,<br />
201 l o g f l a g [ j ] ,<br />
202 s q r t f l a g [ j ] ,<br />
203 e x p f l a g [ j ] ) ;
42 Fomel <strong>Madagascar</strong> Documentation<br />
system/main/add.c<br />
264 s t a t i c void a d d f l o a t ( bool c o l l e c t , /∗ i f c o l l e c t ∗/<br />
265 s i z e t nbuf , /∗ b u f f e r s i z e ∗/<br />
266 f l o a t ∗ buf , /∗ output [ nbuf ] ∗/<br />
267 const f l o a t ∗ bufi , /∗ input [ nbuf ] ∗/<br />
268 char cmode , /∗ o p e r a t i o n ∗/<br />
269 f l o a t s c a l e , /∗ s c a l e f a c t o r ∗/<br />
270 f l o a t add , /∗ add f a c t o r ∗/<br />
271 bool a b s f l a g , /∗ i f abs ∗/<br />
272 bool l o g f l a g , /∗ i f l o g ∗/<br />
273 bool s q r t f l a g , /∗ i f s q r t ∗/<br />
274 bool e x p f l a g /∗ i f exp ∗/)<br />
275 /∗ Add f l o a t i n g point numbers ∗/<br />
276 {<br />
277 s i z e t j ;<br />
278 f l o a t f ;<br />
279<br />
280 for ( j =0; j < nbuf ; j++) {<br />
281 f = b u f i [ j ] ;<br />
282 i f ( a b s f l a g ) f = f a b s f ( f ) ;<br />
283 f += add ;<br />
284 i f ( l o g f l a g ) f = l o g f ( f ) ;<br />
285 i f ( s q r t f l a g ) f = s q r t f ( f ) ;<br />
286 i f ( 1 . != s c a l e ) f ∗= s c a l e ;<br />
287 i f ( e x p f l a g ) f = expf ( f ) ;<br />
288 i f ( c o l l e c t ) {<br />
289 switch ( cmode ) {<br />
290 case ’ p ’ : /∗ product ∗/<br />
291 case ’m’ : /∗ multiply ∗/<br />
292 buf [ j ] ∗= f ;<br />
293 break ;<br />
294 case ’ d ’ : /∗ d e l e t e ∗/<br />
295 i f ( f != 0 . ) buf [ j ] /= f ;<br />
296 break ;<br />
297 d e f a u l t : /∗ add ∗/<br />
298 buf [ j ] += f ;<br />
299 break ;<br />
300 }<br />
301 } else {<br />
302 buf [ j ] = f ;<br />
303 }<br />
304 }<br />
305 }
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 43<br />
sfattr: Display dataset attributes.<br />
sfattr < in.rsf lval=2 want=<br />
Sample output from "sfspike n1=100 | sfbandpass fhi=60 | sfattr"<br />
*******************************************<br />
rms = 0.992354<br />
mean = 0.987576<br />
2-norm = 9.92354<br />
variance = 0.00955481<br />
std dev = 0.0977487<br />
max = 1.12735 at 97<br />
min = 0.151392 at 100<br />
nonzero samples = 100<br />
total samples = 100<br />
*******************************************<br />
rms = sqrt[ sum(data^2) / n ]<br />
mean<br />
= sum(data) / n<br />
norm<br />
= sum(abs(data)^lval)^(1/lval)<br />
variance = [ sum(data^2) - n*mean^2 ] / [ n-1 ]<br />
standard deviation = sqrt [ variance ]<br />
int lval=2 norm option, lval is a non-negative integer,<br />
computes the vector lval-norm<br />
string want= ’all’(default), ’rms’, ’mean’, ’norm’, ’var’,<br />
’std’, ’max’, ’min’, ’nonzero’, ’samples’,<br />
’short’ want= ’rms’ displays the<br />
root mean square want= ’norm’ displays<br />
the square norm, otherwise specified by<br />
lval. want= ’var’ displays the variance<br />
want= ’std’ displays the standard deviation<br />
want= ’nonzero’ displays number of<br />
nonzero samples want= ’samples’ displays<br />
total number of samples want= ’short’<br />
displays a short one-line version<br />
sfattr is a useful diagnostic program. It reports certain statistical values for an<br />
RSF dataset: RMS (root-mean-square) amplitude, mean value, norm value, variance,<br />
standard deviation, maximum and minimum values, number of nonzero samples, and<br />
the total number of samples.<br />
√ If we denote data values as d i for i = 0, 1, 2, . . . , n, then √ the RMS value is<br />
n∑<br />
1<br />
d 2 n i , the mean value is n∑<br />
∑ n 1 d<br />
n i , the L 2 -norm value is d 2 i , the variance<br />
i=0[<br />
i=0<br />
i=0<br />
(<br />
n∑<br />
n<br />
)<br />
∑ 2<br />
1<br />
is d 2 n−1 i − 1 d<br />
n i<br />
], and the standard deviation is the square root of the<br />
i=0<br />
i=0<br />
variance. Using sfattr is a quick way to see the distribution of data values and<br />
check it for anomalies.
44 Fomel <strong>Madagascar</strong> Documentation<br />
Implementation: system/main/attr.c<br />
Computations start by finding the input data (in) size (nsiz) and dimensions (dim).<br />
system/main/attr.c<br />
81 dim = ( s i z e t ) s f l a r g e f i l e d i m s ( in , n ) ;<br />
82 for ( n s i z =1, i =0; i < dim ; i++) {<br />
83 n s i z ∗= n [ i ] ;<br />
84 }<br />
In the main loop, we read the input data buffer by buffer.<br />
system/main/attr.c<br />
100 for ( n l e f t=n s i z ; n l e f t > 0 ; n l e f t −= nbuf ) {<br />
101 nbuf = ( b u f s i z < n l e f t )? b u f s i z : n l e f t ;<br />
102 switch ( type ) {<br />
103 case SF FLOAT :<br />
104 s f f l o a t r e a d ( ( f l o a t ∗) buf , nbuf , in ) ;<br />
105 break ;<br />
106 case SF INT :<br />
107 s f i n t r e a d ( ( i n t ∗) buf , nbuf , in ) ;<br />
108 break ;<br />
109 case SF SHORT:<br />
110 s f s h o r t r e a d ( ( s h o r t ∗) buf , nbuf , in ) ;<br />
111 break ;<br />
112 case SF COMPLEX:<br />
113 sf complexread ( ( sf complex ∗) buf , nbuf , in ) ;<br />
114 break ;<br />
115 case SF UCHAR:<br />
116 s f u c h a r r e a d ( ( unsigned char ∗) buf , nbuf , in ) ;<br />
117 break ;<br />
118 case SF CHAR:<br />
119 d e f a u l t :<br />
120 s f c h a r r e a d ( buf , nbuf , in ) ;<br />
121 break ;<br />
122 }<br />
The data attributes are accumulated in corresponding double-precision variables.<br />
Finally, the attributes are reduced and printed out.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 45<br />
system/main/attr.c<br />
146 fsum += f ;<br />
147 f s q r += ( double ) f ∗ f ;<br />
180 fmean = fsum/ n s i z ;<br />
system/main/attr.c<br />
181 i f ( l v a l ==2) fnorm = s q r t ( f s q r ) ;<br />
182 else i f ( l v a l ==0) fnorm = nsiz −nzero ;<br />
183 else fnorm = pow( f l v a l , 1 . / l v a l ) ;<br />
184 frms = s q r t ( f s q r / n s i z ) ;<br />
185 i f ( n s i z > 1) f v a r = f a b s ( f s q r −n s i z ∗fmean∗fmean ) / ( nsiz −1);<br />
186 else f v a r = 0 . 0 ;<br />
187 f s t d = s q r t ( f v a r ) ;<br />
system/main/attr.c<br />
194 i f (NULL==want | | 0==strcmp ( want , ”rms” ) )<br />
195 p r i n t f ( ” rms = %13.6g \n” , ( f l o a t ) frms ) ;<br />
196 i f (NULL==want | | 0==strcmp ( want , ”mean” ) )<br />
197 p r i n t f ( ” mean = %13.6g \n” , ( f l o a t ) fmean ) ;<br />
198 i f (NULL==want | | 0==strcmp ( want , ”norm” ) )<br />
199 p r i n t f ( ” %d−norm = %13.6g \n” , l v a l , ( f l o a t ) fnorm ) ;<br />
200 i f (NULL==want | | 0==strcmp ( want , ” var ” ) )<br />
201 p r i n t f ( ” v a r i a n c e = %13.6g \n” , ( f l o a t ) f v a r ) ;<br />
202 i f (NULL==want | | 0==strcmp ( want , ” std ” ) )<br />
203 p r i n t f ( ” std dev = %13.6g \n” , ( f l o a t ) f s t d ) ;
46 Fomel <strong>Madagascar</strong> Documentation<br />
sfcat: Concatenate datasets.<br />
sfcat > out.rsf order= space= axis=3 nspace=(int) (ni/(20*nin) + 1) o= d=<br />
[ one.rsf<br />
bash$ sfin one.rsf<br />
one.rsf:<br />
in="/tmp/one.rsf@"<br />
esize=4 type=float form=native<br />
n1=2 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
6 elements 24 bytes<br />
bash$ sfcat one.rsf one.rsf axis=1 > two.rsf<br />
bash$ sfin two.rsf<br />
two.rsf:<br />
in="/tmp/two.rsf@"<br />
esize=4 type=float form=native<br />
n1=4 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
12 elements 48 bytes<br />
Example of sfmerge:<br />
bash$ sfmerge one.rsf one.rsf axis=2 > two.rsf<br />
bash$ sfin two.rsf<br />
two.rsf:<br />
in="/tmp/two.rsf@"
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 47<br />
esize=4 type=float form=native<br />
n1=2 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=7 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
14 elements 56 bytes<br />
In this case, an extra empty trace is inserted between the two merged <strong>file</strong>s.<br />
The axes that are not being merged are checked for consistency:<br />
bash$ sfcat one.rsf two.rsf > three.rsf<br />
sfcat: n2 mismatch: need 3<br />
Implementation: system/main/cat.c<br />
The first input <strong>file</strong> is either in the list or in the standard input.<br />
system/main/cat.c<br />
64 i f ( ! s f s t d i n ( ) ) { /∗ no input f i l e in s t d i n ∗/<br />
65 nin =0;<br />
66 } else {<br />
67 f i l e n a m e [ 0 ] = ” in ” ;<br />
68 nin =1;<br />
69 }<br />
Everything on the command line that does not contain a “=” sign is treated as a<br />
<strong>file</strong> name, and the corresponding <strong>file</strong> object is added to the list.<br />
system/main/cat.c<br />
71 for ( i =1; i< argc ; i++) { /∗ c o l l e c t inputs ∗/<br />
72 i f (NULL != s t r c h r ( argv [ i ] , ’=’ ) )<br />
73 continue ; /∗ not a f i l e ∗/<br />
74 f i l e n a m e [ nin ] = argv [ i ] ;<br />
75 nin++;<br />
76 }<br />
77 i f (0==nin ) s f e r r o r ( ”no input ” ) ;<br />
As explained above, if the space= parameter is not set, it is inferred from the<br />
program name: sfmerge corresponds to space=y and sfcat corresponds to space=n.<br />
Find the axis for the merging (from the command line axis= argument) and figure<br />
out two sizes: n1 for everything after the axis and n2 for everything before the axis.
48 Fomel <strong>Madagascar</strong> Documentation<br />
system/main/cat.c<br />
99 i f ( ! s f g e t b o o l ( ” space ”,& space ) ) {<br />
100 /∗ I n s e r t a d d i t i o n a l space .<br />
101 y i s d e f a u l t for sfmerge , n i s d e f a u l t for s f c a t ∗/<br />
102 prog = s f g e t p r o g ( ) ;<br />
103 i f (NULL != s t r s t r ( prog , ”merge” ) ) {<br />
104 space = true ;<br />
105 } else i f (NULL != s t r s t r ( prog , ” cat ” ) ) {<br />
106 space = f a l s e ;<br />
107 } else {<br />
108 s f w a r n i n g ( ”%s i s n e i t h e r merge nor cat , ”<br />
109 ” assume merge” , prog ) ;<br />
110 space = true ;<br />
111 }<br />
112 }<br />
132 n1=1;<br />
133 n2=1;<br />
system/main/cat.c<br />
134 for ( i =1; i a x i s ) n2 ∗= n [ i −1];<br />
137 }
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 49<br />
In the output, the selected axis will get extended.<br />
system/main/cat.c<br />
149 /∗ f i g u r e out the length o f extended a x i s ∗/<br />
150 ni = 0 ;<br />
151 for ( j =0; j < nin ; j++) {<br />
152 ni += naxis [ j ] ;<br />
153 }<br />
154<br />
155 i f ( space ) {<br />
156 i f ( ! s f g e t i n t ( ” nspace ”,& nspace ) )<br />
157 nspace = ( i n t ) ( ni /(20∗ nin ) + 1 ) ;<br />
158 /∗ i f space=y , number o f t r a c e s to i n s e r t ∗/<br />
159 ni += nspace ∗( nin −1);<br />
160 }<br />
161<br />
162 ( void ) s n p r i n t f ( key , 3 , ”n%d” , a x i s ) ;<br />
163 s f p u t i n t ( out , key , ( i n t ) ni ) ;<br />
The rest is simple: loop through the datasets reading and writing the data in<br />
buffer-size chunks and adding extra empty chunks if space=y.<br />
sfcmplx: Create a complex dataset from its real and imaginary<br />
parts.<br />
sfcmplx < real.rsf > cmplx.rsf real.rsf imag.rsf<br />
There has to be only two input <strong>file</strong>s specified and no additional parameters.<br />
sfcmplx simply creates a complex dataset from its real and imaginary parts. The<br />
reverse operation can be accomplished with sfreal and sfimag.<br />
Example of sfcmplx:<br />
bash$ sfspike n1=2 n2=3 > one.rsf<br />
bash$ sfin one.rsf<br />
one.rsf:<br />
in="/tmp/one.rsf@"<br />
esize=4 type=float form=native<br />
n1=2 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
6 elements 24 bytes
50 Fomel <strong>Madagascar</strong> Documentation<br />
system/main/cat.c<br />
184 for ( i 2 =0; i 2 < n2 ; i 2++) {<br />
185 for ( j =0; j < nin ; j++) {<br />
186 k = order [ j ] ;<br />
187 for ( ni = n1∗ naxis [ k ] ∗ e s i z e ; ni > 0 ; ni −= nbuf ) {<br />
188 nbuf = (BUFSIZ < ni )? BUFSIZ : ni ;<br />
189 s f c h a r r e a d ( buf , nbuf , in [ k ] ) ;<br />
190 s f c h a r w r i t e ( buf , nbuf , out ) ;<br />
191 }<br />
192 i f ( ! space | | j == nin −1) continue ;<br />
193 /∗ Add spaces ∗/<br />
194 memset ( buf , 0 , BUFSIZ ) ;<br />
195 for ( ni = n1∗ nspace ∗ e s i z e ; ni > 0 ; ni −= nbuf ) {<br />
196 nbuf = (BUFSIZ < ni )? BUFSIZ : ni ;<br />
197 s f c h a r w r i t e ( buf , nbuf , out ) ;<br />
198 }<br />
199 }<br />
200 }<br />
bash$ sfcmplx one.rsf one.rsf > cmplx.rsf<br />
bash$ sfin cmplx.rsf<br />
cmplx.rsf:<br />
in="/tmp/cmplx.rsf@"<br />
esize=8 type=complex form=native<br />
n1=2 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
6 elements 48 bytes<br />
Implementation: system/main/cmplx.c<br />
The program flow is simple. First, get the names of the input <strong>file</strong>s.<br />
The main part of the program reads the real and imaginary parts buffer by buffer<br />
and assembles and writes out the complex input.<br />
sfconjgrad: Generic conjugate-gradient solver for linear inversion<br />
sfconjgrad < dat.rsf mod=mod.rsf > to.rsf < from.rsf > out.rsf niter=1<br />
<strong>file</strong> mod= auxiliary input <strong>file</strong> name<br />
int niter=1 number of iterations
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 51<br />
system/main/cmplx.c<br />
41 /∗ the f i r s t two non−parameters are r e a l and imag f i l e s ∗/<br />
42 for ( i =1; i< argc ; i++) {<br />
43 i f (NULL == s t r c h r ( argv [ i ] , ’=’ ) ) {<br />
44 i f (NULL == r e a l ) {<br />
45 r e a l = s f i n p u t ( argv [ i ] ) ;<br />
46 } else {<br />
47 imag = s f i n p u t ( argv [ i ] ) ;<br />
48 break ;<br />
49 }<br />
50 }<br />
51 }<br />
52 i f (NULL == imag ) {<br />
53 i f (NULL == r e a l ) s f e r r o r ( ” not enough input ” ) ;<br />
54 /∗ i f only one input , r e a l i s in s t d i n ∗/<br />
55 imag = r e a l ;<br />
56 r e a l = s f i n p u t ( ” in ” ) ;<br />
57 }<br />
system/main/cmplx.c<br />
81 for ( n l e f t= ( s i z e t ) ( r s i z e ∗ r e s i z e ) ;<br />
82 n l e f t > 0 ; n l e f t −= nbuf ) {<br />
83 nbuf = (BUFSIZ < n l e f t )? BUFSIZ : n l e f t ;<br />
84 s f c h a r r e a d ( rbuf , nbuf , r e a l ) ;<br />
85 s f c h a r r e a d ( ibuf , nbuf , imag ) ;<br />
86 for ( i =0; i < nbuf ; i += r e s i z e ) {<br />
87 memcpy( cbuf+2∗i , rbuf+i , ( s i z e t ) r e s i z e ) ;<br />
88 memcpy( cbuf+2∗ i+r e s i z e , i b u f+i , ( s i z e t ) r e s i z e ) ;<br />
89 }<br />
90 s f c h a r w r i t e ( cbuf ,2∗ nbuf , cmplx ) ;<br />
91 }
52 Fomel <strong>Madagascar</strong> Documentation<br />
sfconjgrad is a generic program for least-squares linear inversion with the conjugategradient<br />
method. Suppose you have an executable program that takes an RSF<br />
<strong>file</strong> from the standard input and produces an RSF <strong>file</strong> in the standard output. It may<br />
take any number of additional parameters but one of them must be adj= that sets the<br />
forward (adj=0) or adjoint (adj=1) operations. The program is typically an<br />
RSF program but it could be anything (a script, a multiprocessor MPI program, etc.)<br />
as long as it implements a linear operator L and its adjoint. There are no restrictions<br />
on the data size or shape. You can easily test the adjointness with sfdottest. The<br />
sfconjgrad program searches for a vector m that minimizes the least-square misfit<br />
‖d − L m‖ 2 for the given input data vector d.<br />
Here is an example. The sfhelicon program implements Claerbout’s multidimensional<br />
helical filtering (Claerbout, 1998). It requires a filter to be specified in<br />
addition to the input and output vectors. We create a helical 2-D filter using the<br />
Unix echo command.<br />
bash$ echo 1 19 20 n1=3 n=20,20 data_format=ascii_int in=lag.rsf > lag.rsf<br />
bash$ echo 1 1 1 a0=-3 n1=3 data_format=ascii_float in=flt.rsf > flt.rsf<br />
Next, we create an example 2-D model and data vector with sfspike.<br />
bash$ sfspike n1=50 n2=50 > vec.rsf<br />
The sfdottest program can perform the dot product test to check that the adjoint<br />
mode works correctly.<br />
bash$ sfdottest sfhelicon filt=flt.rsf lag=lag.rsf \<br />
mod=vec.rsf dat=vec.rsf<br />
sfdottest: L[m]*d=5.28394<br />
sfdottest: L’[d]*m=5.28394<br />
Your numbers may be different because sfdottest generates new random input on<br />
each run. Next, let us make some random data with sfnoise.<br />
bash$ sfnoise seed=2005 rep=y < vec.rsf > dat.rsf<br />
and try to invert the filtering operation using sfconjgrad:<br />
bash$ sfconjgrad sfhelicon filt=flt.rsf lag=lag.rsf \<br />
mod=vec.rsf < dat.rsf > mod.rsf niter=10<br />
sfconjgrad: iter 1 of 10<br />
sfconjgrad: grad=3253.65<br />
sfconjgrad: iter 2 of 10<br />
sfconjgrad: grad=289.421
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 53<br />
sfconjgrad: iter 3 of 10<br />
sfconjgrad: grad=92.3481<br />
sfconjgrad: iter 4 of 10<br />
sfconjgrad: grad=36.9417<br />
sfconjgrad: iter 5 of 10<br />
sfconjgrad: grad=18.7228<br />
sfconjgrad: iter 6 of 10<br />
sfconjgrad: grad=11.1794<br />
sfconjgrad: iter 7 of 10<br />
sfconjgrad: grad=7.26941<br />
sfconjgrad: iter 8 of 10<br />
sfconjgrad: grad=5.15945<br />
sfconjgrad: iter 9 of 10<br />
sfconjgrad: grad=4.23055<br />
sfconjgrad: iter 10 of 10<br />
sfconjgrad: grad=3.57495<br />
The output shows that, in 10 iterations, the norm of the gradient vector decreases by<br />
almost 1000. We can check the residual misfit before<br />
bash$ < dat.rsf sfattr want=norm<br />
norm value = 49.7801<br />
and after<br />
bash$ sfhelicon filt=flt.rsf lag=lag.rsf < mod.rsf | \<br />
sfadd scale=1,-1 dat.rsf | sfattr want=norm<br />
norm value = 5.73563<br />
In 10 iterations, the misfit decreased by an order of magnitude. The result can be<br />
improved by running the program for more iterations.<br />
Implementation: system/main/conjgrad.c<br />
sfcp: Copy or move a dataset.<br />
sfcp < in.rsf > out.rsf in.rsf out.rsf<br />
sfcp - copy, sfmv - move.<br />
Mimics standard Unix commands.<br />
The sfcp and sfmv command imitate the Unix cp and mv commands and serve<br />
for copying and moving RSF <strong>file</strong>s. Example:
54 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ sfspike n1=2 n2=3 > one.rsf<br />
bash$ sfin one.rsf<br />
one.rsf:<br />
in="/tmp/one.rsf@"<br />
esize=4 type=float form=native<br />
n1=2 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
6 elements 24 bytes<br />
bash$ sfcp one.rsf two.rsf<br />
bash$ sfin two.rsf<br />
two.rsf:<br />
in="/tmp/two.rsf@"<br />
esize=4 type=float form=native<br />
n1=2 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
6 elements 24 bytes<br />
Implementation: system/main/cp.c<br />
First, we look for the two first command-line arguments that don’t have the “=”<br />
character in them and consider them as the names of the input and the output <strong>file</strong>s.<br />
system/main/cp.c<br />
47 /∗ the f i r s t two non−parameters are in and out f i l e s ∗/<br />
48 for ( i =1; i< argc ; i++) {<br />
49 i f (NULL == s t r c h r ( argv [ i ] , ’=’ ) ) {<br />
50 i f (NULL == in ) {<br />
51 i n f i l e = argv [ i ] ;<br />
52 in = s f i n p u t ( i n f i l e ) ;<br />
53 } else {<br />
54 out = s f o u t p u t ( argv [ i ] ) ;<br />
55 break ;<br />
56 }<br />
57 }<br />
58 }<br />
Next, we use library functions sf_cp and sf_mv to do the actual work.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 55<br />
66 s f c p ( in , out ) ;<br />
system/main/cp.c<br />
67 i f (NULL != s t r s t r ( prog , ”mv” ) )<br />
68 sf rm ( i n f i l e , f a l s e , f a l s e , f a l s e ) ;<br />
sfcut: Zero a portion of the dataset.<br />
sfcut < in.rsf > out.rsf verb=n j#=(1,...) d#=(d1,d2,...) f#=(0,...)<br />
min#=(o1,o2,,...) n#=(0,...) max#=(o1+(n1-1)*d1,o2+(n1-1)*d2,,...)<br />
Reverse of window.<br />
float d#=(d1,d2,...) sampling in #-th dimension<br />
largeint f#=(0,...) window start in #-th dimension<br />
int j#=(1,...) jump in #-th dimension<br />
float<br />
max#=(o1+(n1-<br />
maximum in #-th dimension<br />
1)*d1,o2+(n1-<br />
1)*d2,,...)<br />
float min#=(o1,o2,,...) minimum in #-th dimension<br />
int n#=(0,...) window size in #-th dimension<br />
bool verb=n [y/n] Verbosity flag<br />
The sfcut command is related to sfwindow and has the same set of arguments<br />
only instead of extracting the selected window, it fills it with zeroes. The size of the<br />
input data is preserved.<br />
Examples:<br />
bash$ sfspike n1=5 n2=5 > in.rsf<br />
bash$ < in.rsf sfdisfil<br />
0: 1 1 1 1 1<br />
5: 1 1 1 1 1<br />
10: 1 1 1 1 1<br />
15: 1 1 1 1 1<br />
20: 1 1 1 1 1<br />
bash$ < in.rsf sfcut n1=2 f1=1 n2=3 f2=2 | sfdisfil<br />
0: 1 1 1 1 1<br />
5: 1 1 1 1 1<br />
10: 1 0 0 1 1<br />
15: 1 0 0 1 1<br />
20: 1 0 0 1 1<br />
bash$ < in.rsf sfcut j1=2 | sfdisfil<br />
0: 0 1 0 1 0<br />
5: 0 1 0 1 0
56 Fomel <strong>Madagascar</strong> Documentation<br />
10: 0 1 0 1 0<br />
15: 0 1 0 1 0<br />
20: 0 1 0 1 0<br />
sfdd: Convert between different formats.<br />
sfdd < in.rsf > out.rsf trunc=n line=8 ibm=n form= type= format=<br />
string form= ascii, native, xdr<br />
string format= Element format (for conversion to ASCII)<br />
bool ibm=n [y/n] Special case - assume integers actually<br />
represent IBM floats<br />
int line=8 Number of numbers per line (for conversion<br />
to ASCII)<br />
bool trunc=n [y/n] Truncate or round to nearest when converting<br />
from float to int/short<br />
string type= int, float, complex, short<br />
The sfdd program is used to change either the form (ascii, xdr, native) or the<br />
type (complex, float, int, char) of the input dataset.<br />
In the example below, we create a plain text (ASCII) <strong>file</strong> with numbers and then<br />
use sfdd to generate an RSF <strong>file</strong> in xdr form with complex numbers.<br />
bash$ cat test.txt<br />
1 2 3 4 5 6<br />
bash$ echo n1=6 data_format=ascii_int in=test.txt > test.rsf<br />
bash$ sfin test.rsf<br />
test.rsf:<br />
in="test.txt"<br />
esize=0 type=int form=ascii<br />
n1=6 d1=? o1=?<br />
6 elements<br />
bash$ sfdd < test.rsf form=xdr type=complex > test2.rsf<br />
bash$ sfin test2.rsf<br />
test2.rsf:<br />
in="/tmp/test2.rsf@"<br />
esize=8 type=complex form=xdr<br />
n1=3 d1=? o1=?<br />
3 elements 24 bytes<br />
bash$ sfdisfil < test2.rsf<br />
0: 1, 2i 3, 4i 5, 6i<br />
To learn more about the RSF data format, consult the guide to RSF format.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 57<br />
sfdisfil: Print out data values.<br />
sfdisfil < in.rsf number=y col=0 format= header= trailer=<br />
Alternatively, use sfdd and convert to ASCII form.<br />
int col=0 Number of columns. The default depends<br />
on the data type: 10 for int and char, 5<br />
for float, 3 for complex<br />
string format= Format for numbers (printf-style). The<br />
default depends on the data type: ”””<br />
string header= Optional header string to output before<br />
data<br />
bool number=y [y/n] If number the elements<br />
string trailer= Optional trailer string to output after<br />
data<br />
The sfdisfil program simply dumps the data contents to the standard output<br />
in a text form. It is used mostly for debugging purposes to quickly examine RSF <strong>file</strong>s.<br />
Here is an example:<br />
bash$ sfmath o1=0 d1=2 n1=12 output=x1 > test.rsf<br />
bash$ < test.rsf sfdisfil<br />
0: 0 2 4 6 8<br />
5: 10 12 14 16 18<br />
10: 20 22<br />
The output format is easily configurable.<br />
bash$ < test.rsf sfdisfil col=6 number=n format="%5.1f"<br />
0.0 2.0 4.0 6.0 8.0 10.0<br />
12.0 14.0 16.0 18.0 20.0 22.0<br />
Along with sfdd, sfdisfil provides a simple way to convert RSF data to an ASCII<br />
form.<br />
sfdottest: Generic dot-product test for linear operators with<br />
adjoints<br />
sfdottest mod=mod.rsf dat=dat.rsf > pip.rsf<br />
<strong>file</strong> dat= auxiliary input <strong>file</strong> name<br />
<strong>file</strong> mod= auxiliary input <strong>file</strong> name<br />
sfdottest is a generic dot-product test program for testing linear operators. Suppose<br />
there is an executable program that takes an RSF <strong>file</strong> from the standard
58 Fomel <strong>Madagascar</strong> Documentation<br />
input and produces an RSF <strong>file</strong> in the standard output. It may take any number of<br />
additional parameters but one of them must be adj= that sets the forward (adj=0)<br />
or adjoint (adj=1) operations. The program is typically an RSF program<br />
but it could be anything (a script, a multiprocessor MPI program, etc.) as long as<br />
it implements a linear operator L and its adjoint L T . The sfdottest program is<br />
testing the equality<br />
d T L m = m T L T d (1)<br />
by using random vectors m and d. You can invoke it with<br />
bash$ sfdottest [optional aruments] mod=mod.rsf dat=dat.rsf<br />
where mod.rsf and dat.rsf are RSF <strong>file</strong>s that represent vectors from the model and<br />
data spaces. sfdottest does not create any temporary <strong>file</strong>s and does not have any<br />
restrictive limitations on the size of the vectors.<br />
Here is an example. We first setup a vector with 100 elements using sfspike and<br />
then run sfdottest to test the sfcausint program. sfcausint implements a linear<br />
operator of causal integration and its adjoint, the anti-causal integration.<br />
bash$ sfspike n1=100 > vec.rsf<br />
bash$ sfdottest sfcausint mod=vec.rsf dat=vec.rsf<br />
sfdottest: L[m]*d=1410.2<br />
sfdottest: L’[d]*m=1410.2<br />
bash$ sfdottest sfcausint mod=vec.rsf dat=vec.rsf<br />
sfdottest: L[m]*d=1165.87<br />
sfdottest: L’[d]*m=1165.87<br />
The numbers are different on subsequent runs because of changing seed in the random<br />
number generator.<br />
Here is a somewhat more complicated example. The sfhelicon program implements<br />
Claerbout’s multidimensional helical filtering (Claerbout, 1998). It requires a<br />
filter to be specified in addition to the input and output vectors. We create a helical<br />
2-D filter using the Unix echo command.<br />
bash$ echo 1 19 20 n1=3 n=20,20 data_format=ascii_int in=lag.rsf > lag.rsf<br />
bash$ echo 1 1 1 a0=-3 n1=3 data_format=ascii_float in=flt.rsf > flt.rsf<br />
Next, we create an example 2-D model and data vector with sfspike.<br />
bash$ sfspike n1=50 n2=50 > vec.rsf<br />
Now the sfdottest program can perform the dot product test.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 59<br />
bash$ sfdottest sfhelicon filt=flt.rsf lag=lag.rsf \<br />
> mod=vec.rsf dat=vec.rsf<br />
sfdottest: L[m]*d=8.97375<br />
sfdottest: L’[d]*m=8.97375<br />
Here is the same program tested in the inverse filtering mode:<br />
bash$ sfdottest sfhelicon filt=flt.rsf lag=lag.rsf \<br />
> mod=vec.rsf dat=vec.rsf div=y<br />
sfdottest: L[m]*d=15.0222<br />
sfdottest: L’[d]*m=15.0222<br />
sfget: Output parameters from the header.<br />
sfget parform=y all=n par1 par2 ...<br />
bool all=n [y/n] If output all values.<br />
bool parform=y [y/n] If y, print out parameter=value. If n,<br />
print out value.<br />
The sfget program extracts a parameter value from an RSF <strong>file</strong>. It is useful<br />
mostly for scripting. Here is, for example, a quick calculation of the maximum value<br />
on the first axis in an RSF dataset (the output of sfspike) using the standard Unix<br />
bc calculator.<br />
bash$ ( sfspike n1=100 | sfget n1 d1 o1; echo "o1+(n1-1)*d1" ) | bc<br />
.396<br />
See also sfput.<br />
Implementation: system/main/get.c<br />
The implementation is trivial. Loop through all command-line parameters that contain<br />
the “=” character.<br />
system/main/get.c<br />
41 i f ( ! s f g e t b o o l ( ” a l l ”,& a l l ) ) a l l=f a l s e ;<br />
42 /∗ I f output a l l v a l u e s . ∗/<br />
Get the parameter value (as string) and output it as either key=value or value,<br />
depending on the parform parameter.
60 Fomel <strong>Madagascar</strong> Documentation<br />
system/main/get.c<br />
44 t a b l e = s f s i m t a b i n i t ( t a b s i z e ) ;<br />
45 s f s i m t a b i n p u t ( table , stdin , NULL) ;<br />
46<br />
47 i f ( a l l ) {<br />
48 s f s i m t a b o u t p u t ( table , stdout ) ;<br />
49 } else {<br />
50 for ( i = 1 ; i < argc ; i++) {<br />
sfheadercut: Zero a portion of a dataset based on a header<br />
mask.<br />
sfheadercut mask=head.rsf < in.rsf > out.rsf<br />
The input data is a collection of traces n1xn2,<br />
mask is an integer array of size n2.<br />
<strong>file</strong> mask= auxiliary input <strong>file</strong> name<br />
sfheadercut is close to sfheaderwindow but instead of windowing the dataset,<br />
it fills the traces specified by the header mask with zeroes. The size of the input data<br />
is preserved.<br />
Here is an example of using sfheaderwindow for zeroing every other trace in the<br />
input <strong>file</strong>. First, let us create an input <strong>file</strong> with ten traces:<br />
bash$ sfmath n1=5 n2=10 output=x2+1 > input.rsf<br />
bash$ < input.rsf sfdisfil<br />
0: 1 1 1 1 1<br />
5: 2 2 2 2 2<br />
10: 3 3 3 3 3<br />
15: 4 4 4 4 4<br />
20: 5 5 5 5 5<br />
25: 6 6 6 6 6<br />
30: 7 7 7 7 7<br />
35: 8 8 8 8 8<br />
40: 9 9 9 9 9<br />
45: 10 10 10 10 10<br />
Next, we can create a mask with alternating ones and zeros using sfinterleave.<br />
bash$ sfspike n1=5 mag=1 | sfdd type=int > ones.rsf
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 61<br />
bash$ sfspike n1=5 mag=0 | sfdd type=int > zeros.rsf<br />
bash$ sfinterleave axis=1 ones.rsf zeros.rsf > mask.rsf<br />
bash$ sfdisfil < mask.rsf<br />
0: 1 0 1 0 1 0 1 0 1 0<br />
Finally, sfheadercut zeros the input traces.<br />
bash$ sfheadercut < input.rsf mask=mask.rsf > output.rsf<br />
bash$ sfdisfil < output.rsf<br />
0: 1 1 1 1 1<br />
5: 0 0 0 0 0<br />
10: 3 3 3 3 3<br />
15: 0 0 0 0 0<br />
20: 5 5 5 5 5<br />
25: 0 0 0 0 0<br />
30: 7 7 7 7 7<br />
35: 0 0 0 0 0<br />
40: 9 9 9 9 9<br />
45: 0 0 0 0 0<br />
sfheadersort: Sort a dataset according to a header key.<br />
sfheadersort < in.rsf > out.rsf head=<br />
string head= header <strong>file</strong><br />
sfheadersort is used to sort traces in the input <strong>file</strong> according to trace header<br />
information.<br />
Here is an example of using sfheadersort for randomly shuffling traces in the<br />
input <strong>file</strong>. First, let us create an input <strong>file</strong> with seven traces:<br />
bash$ sfmath n1=5 n2=7 output=x2+1 > input.rsf<br />
bash$ < input.rsf sfdisfil<br />
0: 1 1 1 1 1<br />
5: 2 2 2 2 2<br />
10: 3 3 3 3 3<br />
15: 4 4 4 4 4<br />
20: 5 5 5 5 5<br />
25: 6 6 6 6 6<br />
30: 7 7 7 7 7<br />
Next, we can create a random <strong>file</strong> with seven header values using sfnoise.
62 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ sfspike n1=7 | sfnoise rep=y type=n > random.rsf<br />
bash$ < random.rsf sfdisfil<br />
0: 0.05256 -0.2879 0.1487 0.4097 0.1548<br />
5: 0.4501 0.2836<br />
If you reproduce this example, your numbers will most likely be different, because,<br />
in the absence of seed= parameter, sfnoise uses a random seed value to generate<br />
pseudo-random numbers. Finally, we apply sfheadersort to shuffle the input traces.<br />
bash$ < input.rsf sfheadersort head=random.rsf > output.rsf<br />
bash$ < output.rsf sfdisfil<br />
0: 2 2 2 2 2<br />
5: 1 1 1 1 1<br />
10: 3 3 3 3 3<br />
15: 5 5 5 5 5<br />
20: 7 7 7 7 7<br />
25: 4 4 4 4 4<br />
30: 6 6 6 6 6<br />
As expected, the order of traces in the output <strong>file</strong> corresponds to the order of values<br />
in the header. Thanks to the separation between headers and data, the operation of<br />
sfheadersort is optimally efficient. It first sorts the headers and only then accesses<br />
the data, reading each data trace only once.<br />
sfheaderwindow: Window a dataset based on a header mask.<br />
sfheaderwindow mask=head.rsf < in.rsf > out.rsf<br />
The input data is a collection of traces n1xn2,<br />
mask is an integer array os size n2, windowed is n1xm2,<br />
where m2 is the number of nonzero elements in mask.<br />
<strong>file</strong> mask= auxiliary input <strong>file</strong> name<br />
sfheaderwindow is used to window traces in the input <strong>file</strong> according to trace<br />
header information.<br />
Here is an example of using sfheaderwindow for randomly selecting part of the<br />
traces in the input <strong>file</strong>. First, let us create an input <strong>file</strong> with ten traces:<br />
bash$ sfmath n1=5 n2=10 output=x2+1 > input.rsf<br />
bash$ < input.rsf sfdisfil<br />
0: 1 1 1 1 1<br />
5: 2 2 2 2 2<br />
10: 3 3 3 3 3
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 63<br />
15: 4 4 4 4 4<br />
20: 5 5 5 5 5<br />
25: 6 6 6 6 6<br />
30: 7 7 7 7 7<br />
35: 8 8 8 8 8<br />
40: 9 9 9 9 9<br />
45: 10 10 10 10 10<br />
Next, we can create a random <strong>file</strong> with ten header values using sfnoise.<br />
bash$ sfspike n1=10 | sfnoise rep=y type=n > random.rsf<br />
bash$ < random.rsf sfdisfil<br />
0: -0.005768 0.02258 -0.04331 -0.4129 -0.3909<br />
5: -0.03582 0.4595 -0.3326 0.498 -0.3517<br />
If you reproduce this example, your numbers will most likely be different, because,<br />
in the absence of seed= parameter, sfnoise uses a random seed value to generate<br />
pseudo-random numbers. Finally, we apply sfheaderwindow to window the input<br />
traces selecting only those for which the header is greater than zero.<br />
bash$ < random.rsf sfmask min=0 > mask.rsf<br />
bash$ < mask.rsf sfdisfil<br />
0: 0 1 0 0 0 0 1 0 1 0<br />
bash$ < input.rsf sfheaderwindow mask=mask.rsf > output.rsf<br />
bash$ < output.rsf sfdisfil<br />
0: 2 2 2 2 2<br />
5: 7 7 7 7 7<br />
10: 9 9 9 9 9<br />
In this case, only three traces are selected for the output. Thanks to the separation<br />
between headers and data, the operation of sfheaderwindow is optimally efficient.<br />
sfin: Display basic information about RSF <strong>file</strong>s.<br />
sfin info=y check=2. trail=y [
64 Fomel <strong>Madagascar</strong> Documentation<br />
sfin is one of the most useful programs for operating with RSF <strong>file</strong>s. It produces<br />
quick information on the <strong>file</strong> hypercube dimensions and checks the consistency of the<br />
associated data <strong>file</strong>.<br />
Here is an example. Let us create an RSF <strong>file</strong> and examine it with sfin.<br />
bash$ sfspike n1=100 n2=20 > spike.rsf<br />
bash$ sfin spike.rsf<br />
spike.rsf:<br />
in="/tmp/spike.rsf@"<br />
esize=4 type=float form=native<br />
n1=100 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=20 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
2000 elements 8000 bytes<br />
sfin reports the following information:<br />
• location of the data <strong>file</strong> (/tmp/spike.rsf)<br />
• element size (4 bytes)<br />
• element type (floating point)<br />
• element form (native)<br />
• hypercube dimensions (100 by 20)<br />
• axes scale (0.004 and 0.1)<br />
• axes origin (0 and 0)<br />
• axes labels<br />
• axes units<br />
• total number of elements<br />
• total number of bytes in the data <strong>file</strong><br />
Suppose that the <strong>file</strong> got corrupted by a buggy program and reports incorrect<br />
dimensions. The sfin program should be able to catch the discrepancy.<br />
bash$ echo n2=100 >> spike.rsf<br />
bash$ sfin spike.rsf > /dev/null<br />
sfin:<br />
Actually 8000 bytes, 20% of expected.<br />
sfin also checks the first records in the <strong>file</strong> for zeros.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 65<br />
bash$ sfspike n1=100 n2=100 k2=99 > spike2.rsf<br />
bash$ sfin spike2.rsf >/dev/null<br />
sfin: The first 32768 bytes are all zeros<br />
The number of bytes to check is adjustable<br />
bash$ sfin spike2.rsf check=0.01 >/dev/null<br />
sfin: The first 16384 bytes are all zeros<br />
You can also output only the location of the data <strong>file</strong>. This is sometimes handy<br />
in scripts.<br />
bash$ sfin spike.rsf spike2.rsf info=n<br />
/tmp/spike.rsf@ /tmp/spike2.rsf@<br />
An alternative is to use sfget, as follows:<br />
bash$ sfget parform=n in < spike.rsf<br />
/tmp/spike.rsf@<br />
sfinterleave: Combine several datasets by interleaving.<br />
sfinterleave > out.rsf axis=3 [< <strong>file</strong>0.rsf] <strong>file</strong>1.rsf <strong>file</strong>2.rsf ...<br />
int axis=3 Axis for interleaving<br />
sfinterleave combines two or more datasets by interleaving them on one of the<br />
axes. Here is a quick example:<br />
bash$ sfspike n1=5 n2=5 > one.rsf<br />
bash$ sfdisfil < one.rsf<br />
0: 1 1 1 1 1<br />
5: 1 1 1 1 1<br />
10: 1 1 1 1 1<br />
15: 1 1 1 1 1<br />
20: 1 1 1 1 1<br />
bash$ sfscale < one.rsf dscale=2 > two.rsf<br />
bash$ sfdisfil < two.rsf<br />
0: 2 2 2 2 2<br />
5: 2 2 2 2 2<br />
10: 2 2 2 2 2<br />
15: 2 2 2 2 2<br />
20: 2 2 2 2 2
66 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ sfinterleave one.rsf two.rsf axis=1 | sfdisfil<br />
0: 1 2 1 2 1<br />
5: 2 1 2 1 2<br />
10: 1 2 1 2 1<br />
15: 2 1 2 1 2<br />
20: 1 2 1 2 1<br />
25: 2 1 2 1 2<br />
30: 1 2 1 2 1<br />
35: 2 1 2 1 2<br />
40: 1 2 1 2 1<br />
45: 2 1 2 1 2<br />
bash$ sfinterleave < one.rsf two.rsf axis=2 | sfdisfil<br />
0: 1 1 1 1 1<br />
5: 2 2 2 2 2<br />
10: 1 1 1 1 1<br />
15: 2 2 2 2 2<br />
20: 1 1 1 1 1<br />
25: 2 2 2 2 2<br />
30: 1 1 1 1 1<br />
35: 2 2 2 2 2<br />
40: 1 1 1 1 1<br />
45: 2 2 2 2 2<br />
sfmask: Create a mask.<br />
sfmask < in.rsf > out.rsf min= max= min= max=<br />
Mask is an integer data with ones and zeros.<br />
Ones correspond to input values between min and max.<br />
The output can be used with sfheaderwindow.<br />
int max= maximum header value<br />
int min= minimum header value<br />
sfmask creates an integer output of ones and zeros comparing the values of the<br />
input data to specified min= and max= parameters. It is useful for sfheaderwindow<br />
and in many other applications. Here is a quick example:<br />
bash$ sfmath n1=10 output="sin(x1)" > sin.rsf<br />
bash$ < sin.rsf sfdisfil<br />
0: 0 0.8415 0.9093 0.1411 -0.7568<br />
5: -0.9589 -0.2794 0.657 0.9894 0.4121<br />
bash$ < sin.rsf sfmask min=-0.5 max=0.5 | sfdisfil<br />
0: 1 0 0 1 0 0 1 0 0 1
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 67<br />
sfmath: Mathematical operations on data <strong>file</strong>s.<br />
sfmath > out.rsf n#= d#=(1,1,...) o#=(0,0,...) label#= unit#= type= label= unit=<br />
output=<br />
Known functions:<br />
cos, sin, tan, acos, asin, atan,<br />
cosh, sinh, tanh, acosh, asinh, atanh,<br />
exp, log, sqrt, abs,<br />
erf, erfc (for float data),<br />
arg, conj, real, imag (for complex data).<br />
sfmath will work on float or complex data, but all the input and output<br />
<strong>file</strong>s must be of the same data type.<br />
An alternative to sfmath is sfadd, which may be more efficient, but is<br />
less versatile.<br />
Examples:<br />
sfmath x=<strong>file</strong>1.rsf y=<strong>file</strong>2.rsf power=<strong>file</strong>3.rsf output=’sin((x+2*y)^power)’ > out.rsf<br />
sfmath < <strong>file</strong>1.rsf tau=<strong>file</strong>2.rsf output=’exp(tau*input)’ > out.rsf<br />
sfmath n1=100 type=complex output="exp(I*x1)" > out.rsf<br />
Arguments which are not treated as variables in mathematical expressions:<br />
datapath=, type=, out=<br />
See also: sfheadermath.<br />
float d#=(1,1,...) sampling on #-th axis<br />
string label= data label<br />
string label#= label on #-th axis<br />
largeint n#= size of #-th axis<br />
float o#=(0,0,...) origin on #-th axis<br />
string output= Mathematical description of the output<br />
string type= output data type [float,complex]<br />
string unit= data unit<br />
string unit#= unit on #-th axis<br />
sfmath is a versatile program for mathematical operations with RSF <strong>file</strong>s. It can<br />
operate with several input <strong>file</strong>, all of the same dimensions and data type. The data<br />
type can be real (floating point) or complex. Here is an example that demonstrates<br />
several features of sfmath.<br />
bash$ sfmath n1=629 d1=0.01 o1=0 n2=40 d2=1 o2=5 \<br />
output="x2*(8+sin(6*x1+x2/10))" > rad.rsf<br />
bash$ < rad.rsf sfrtoc | sfmath output="input*exp(I*x1)" > rose.rsf<br />
bash$ < rose.rsf sfgraph title=Rose screenratio=1 wantaxis=n | sfpen<br />
The first line creates a 2-D dataset that consists of 40 traces 629 samples each. The
68 Fomel <strong>Madagascar</strong> Documentation<br />
values of the data are computed with the formula "x2*(8+sin(6*x1+x2/10))", where<br />
x1 refers to the coordinate on the first axis, and x2 is the coordinate of the second<br />
axis. In the second line, we convert the data from real to complex using sfrtoc and<br />
produce a complex dataset using formula "input*exp(I*x1)", where input refers to<br />
the input <strong>file</strong>. Finally, we plot the complex data as a collection of parametric curves<br />
using sfgraph and display the result using sfpen. The plot appearing on your screen<br />
should look similar to Figure 1.<br />
Figure 1: This figure was created with sfmath. rsf/sfmath rose<br />
One possible alternative to the second line above is<br />
bash$ < rad.rsf sfmath output=x1 > ang.rsf<br />
bash$ sfmath r=rad.rsf a=ang.rsf output="r*cos(a)" > cos.rsf<br />
bash$ sfmath r=rad.rsf a=ang.rsf output="r*sin(a)" > sin.rsf<br />
bash$ sfcmplx cos.rsf sin.rsf > rose.rsf<br />
Here we refer to input <strong>file</strong>s by names (r and a) and combine the names in a formula.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 69<br />
sfpad: Pad a dataset with zeros.<br />
sfpad < in.rsf > out.rsf beg#=0 end#=0<br />
n\#out is equivalent to n\#, both of them overwrite end\#.<br />
int beg#=0 the number of zeros to add before the beginning<br />
of #-th axis<br />
int end#=0 the number of zeros to add after the end<br />
of #-th axis<br />
pad increases the dimensions of the input dataset by padding the data with zeroes.<br />
Here are some simple examples.<br />
bash$ sfspike n1=5 n2=3 > one.rsf<br />
bash$ sfdisfil < one.rsf<br />
0: 1 1 1 1 1<br />
5: 1 1 1 1 1<br />
10: 1 1 1 1 1<br />
bash$ < one.rsf sfpad n2=5 | sfdisfil<br />
0: 1 1 1 1 1<br />
5: 1 1 1 1 1<br />
10: 1 1 1 1 1<br />
15: 0 0 0 0 0<br />
20: 0 0 0 0 0<br />
bash$ < one.rsf sfpad beg2=2 | sfdisfil<br />
0: 0 0 0 0 0<br />
5: 0 0 0 0 0<br />
10: 1 1 1 1 1<br />
15: 1 1 1 1 1<br />
20: 1 1 1 1 1<br />
bash$ < one.rsf sfpad beg2=1 end2=1 | sfdisfil<br />
0: 0 0 0 0 0<br />
5: 1 1 1 1 1<br />
10: 1 1 1 1 1<br />
15: 1 1 1 1 1<br />
20: 0 0 0 0 0<br />
bash$ < one.rsf sfwindow n1=3 | sfpad n1=5 n2=5 beg1=1 beg2=1 | sfdisfil<br />
0: 0 0 0 0 0<br />
5: 0 1 1 1 0<br />
10: 0 1 1 1 0<br />
15: 0 1 1 1 0<br />
20: 0 0 0 0 0<br />
You can use sfcat to pad data with values other than zeroes.
70 Fomel <strong>Madagascar</strong> Documentation<br />
sfput: Input parameters into a header.<br />
sfput < in.rsf > out.rsf<br />
sfput is a very simple program. It simply appends parameters from the command<br />
line to the output RSF <strong>file</strong>. One can achieve similar results with editing by hand or<br />
with standard Unix utilities like sed and echo. sfput is sometimes more convenient<br />
because it handles input/output operations similarly to other regular RSF programs.<br />
bash$ sfspike n1=10 > spike.rsf<br />
bash$ sfin spike.rsf<br />
spike.rsf:<br />
in="/tmp/spike.rsf@"<br />
esize=4 type=float form=native<br />
n1=10 d1=0.004 o1=0 label1="Time" unit1="s"<br />
10 elements 40 bytes<br />
bash$ sfput < spike.rsf d1=25 label1=Depth unit1=m > spike2.rsf<br />
bash$ sfin spike2.rsf<br />
spike2.rsf:<br />
in="/tmp/spike2.rsf@"<br />
esize=4 type=float form=native<br />
n1=10 d1=25 o1=0 label1="Depth" unit1="m"<br />
10 elements 40 bytes<br />
sfreal: Extract real (sfreal) or imaginary (sfimag) part of a<br />
complex dataset.<br />
sfreal < cmplx.rsf > real.rsf<br />
sfreal extracts the real part of a complex type dataset. The imaginary part can<br />
be extracted with sfimag, an the real and imaginary part can be combined together<br />
with sfcmplx.<br />
Here is a simple example. Let us first create a complex dataset with sfmath<br />
bash$ sfmath n1=10 type=complex output="(2+I)*x1" > cmplx.rsf<br />
bash$ fdisfil < cmplx.rsf<br />
0: 0, 0i 2, 1i 4, 2i<br />
3: 6, 3i 8, 4i 10, 5i<br />
6: 12, 6i 14, 7i 16, 8i<br />
9: 18, 9i<br />
Extracting the real part with sfreal:
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 71<br />
bash$ sfreal < cmplx.rsf | sfdisfil<br />
0: 0 2 4 6 8<br />
5: 10 12 14 16 18<br />
Extracting the imaginary part with sfimag:<br />
bash$ sfimag < cmplx.rsf | sfdisfil<br />
0: 0 1 2 3 4<br />
5: 5 6 7 8 9<br />
sfreverse: Reverse one or more axes in the data hypercube.<br />
sfreverse < in.rsf > out.rsf which=-1 verb=n memsize=sf memsize() opt=<br />
int memsize=sf memsize() Max amount of RAM (in Mb) to be used<br />
string opt= If y, change o and d parameters on the<br />
reversed axis; if i, don’t change o and d<br />
bool verb=n [y/n] Verbosity flag<br />
int which=-1 Which axis to reverse. To reverse a given<br />
axis, start with 0, add 1 to number to reverse<br />
n1 dimension, add 2 to number to<br />
reverse n2 dimension, add 4 to number to<br />
reverse n3 dimension, etc. Thus, which=7<br />
would reverse the first three dimensions,<br />
which=5 just n1 and n3, etc. which=0<br />
will just pass the input on through unchanged.<br />
Here is an example of using sfreverse. First, let us create a 2-D dataset.<br />
bash$ sfmath n1=5 d1=1 n2=3 d2=1 output=x1+x2 > test.rsf<br />
bash$ < test.rsf sfdisfil<br />
0: 0 1 2 3 4<br />
5: 1 2 3 4 5<br />
10: 2 3 4 5 6<br />
Reversing the first axis:<br />
bash$ < test.rsf sfreverse which=1 | sfdisfil<br />
0: 4 3 2 1 0<br />
5: 5 4 3 2 1<br />
10: 6 5 4 3 2<br />
Reversing the second axis:
72 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ < test.rsf sfreverse which=2 | sfdisfil<br />
0: 2 3 4 5 6<br />
5: 1 2 3 4 5<br />
10: 0 1 2 3 4<br />
Reversing both the first and the second axis:<br />
bash$ < test.rsf sfreverse which=3 | sfdisfil<br />
0: 2 3 4 5 6<br />
5: 1 2 3 4 5<br />
10: 0 1 2 3 4<br />
As you can see, the which= parameter controls the axes that are being reversed by<br />
encoding them into one number.<br />
When an axis is reversed, what happens with its axis origin and sampling parameters?<br />
This behavior is controlled by opt=. In our example,<br />
bash$ < test.rsf sfget n1 o1 d1<br />
n1=5<br />
o1=0<br />
d1=1<br />
bash$ < test.rsf sfreverse which=1 | sfget o1 d1<br />
o1=4<br />
d1=-1<br />
The default behavior (equivalent to opt=y) puts the origin o1 at the end of the axis<br />
and reverses the sampling parameter d1. Using opt=n preserves the sampling but<br />
reverses the origin.<br />
bash$ < test.rsf sfreverse which=1 opt=n | sfget o1 d1<br />
o1=-4<br />
d1=1<br />
Using opt=i preserves both the sampling and the origin while reversing the axis.<br />
bash$ < test.rsf sfreverse which=1 opt=i | sfget o1 d1<br />
o1=0<br />
d1=1<br />
One of the three possible behaviors may be desirable depending on the application.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 73<br />
sfrm: Remove RSF <strong>file</strong>s together with their data.<br />
sfrm <strong>file</strong>1.rsf [<strong>file</strong>2.rsf ...] [-i] [-v] [-f]<br />
Mimics the standard Unix rm command.<br />
See also: sfmv, sfcp.<br />
sfrm is a program for removing RSF <strong>file</strong>s. Its arguments mimic the arguments of<br />
the standard Unix rm utility: -v for verbosity, -i for interactive inquiry, -f for force<br />
removal of suspicious <strong>file</strong>s. Unlike the Unix rm, sfrm removes both the RSF header<br />
<strong>file</strong>s and the binary <strong>file</strong>s that the headers point to.<br />
Example:<br />
bash$ sfspike n1=10 > spike.rsf datapath=./<br />
bash$ sfget in < spike.rsf<br />
in=./spike.rsf@<br />
bash$ ls spike*<br />
spike.rsf spike.rsf@<br />
bash$ sfrm -v spike.rsf<br />
sfrm: sf_rm: Removing header spike.rsf<br />
sfrm: sf_rm: Removing data ./spike.rsf@<br />
bash$ ls spike*<br />
ls: No match.<br />
sfrotate: Rotate a portion of one or more axes in the data<br />
hypercube.<br />
sfrotate < in.rsf > out.rsf verb=n memsize=sf memsize() rot#=(0,0,...)<br />
int memsize=sf memsize() Max amount of RAM (in Mb) to be used<br />
int rot#=(0,0,...) length of #-th axis that is moved to the<br />
end<br />
bool verb=n [y/n] Verbosity flag<br />
sfrotate modifies the input dataset by splitting it into parts and putting the<br />
parts back in a different order. Here is a quick example.<br />
bash$ sfmath n1=5 d1=1 n2=3 d2=1 output=x1+x2 > test.rsf<br />
bash$ < test.rsf sfdisfil<br />
0: 0 1 2 3 4<br />
5: 1 2 3 4 5<br />
10: 2 3 4 5 6
74 Fomel <strong>Madagascar</strong> Documentation<br />
Rotating the first axis by putting the last two columns in front:<br />
bash$ < test.rsf sfrotate rot1=2 | sfdisfil<br />
0: 3 4 0 1 2<br />
5: 4 5 1 2 3<br />
10: 5 6 2 3 4<br />
Rotating the second axis by putting the last row in front:<br />
bash$ < test.rsf sfrotate rot2=1 | sfdisfil<br />
0: 2 3 4 5 6<br />
5: 0 1 2 3 4<br />
10: 1 2 3 4 5<br />
Rotating both the first and the second axis:<br />
bash$ < test.rsf sfrotate rot1=3 rot2=1 | sfdisfil<br />
0: 4 5 6 2 3<br />
5: 2 3 4 0 1<br />
10: 3 4 5 1 2<br />
The transformation is shown schematically in Figure 2.<br />
before<br />
after<br />
Figure 2: Schematic transformation of data with sfrotate. rsf/XFig rotate<br />
sfrtoc: Convert real data to complex (by adding zero imaginary<br />
part).<br />
sfrtoc < real.rsf > cmplx.rsf<br />
See also: sfcmplx<br />
The input to sfrtoc can be any type=float dataset:
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 75<br />
bash$ sfspike n1=10 n2=20 n3=30 >real.rsf<br />
bash$ sfin real.rsf<br />
real.rsf:<br />
in="/var/tmp/real.rsf@"<br />
esize=4 type=float form=native<br />
n1=10 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=20 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
n3=30 d3=0.1 o3=0 label3="Distance" unit3="km"<br />
6000 elements 24000 bytes<br />
The output dataset will have type=complex, and its binary will be twice the size of<br />
the input:<br />
bash$ complex.rsf<br />
bash$ sfin complex.rsf<br />
complex.rsf:<br />
in="/var/tmp/complex.rsf@"<br />
esize=8 type=complex form=native<br />
n1=10 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=20 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
n3=30 d3=0.1 o3=0 label3="Distance" unit3="km"<br />
6000 elements 48000 bytes<br />
sfscale: Scale data.<br />
sfscale < in.rsf > out.rsf axis=0 rscale=0.<br />
dscale=1.<br />
To scale by a constant factor, you can also use sfmath.<br />
int axis=0 Scale by maximum in the dimensions up<br />
to this axis.<br />
float dscale=1. Scale by this factor (works if rscale=0)<br />
float rscale=0. Scale by this factor.<br />
sfscale scales the input dataset by a factor. Here are some simple examples.<br />
First, let us create a test dataset.<br />
bash$ sfmath n1=5 n2=3 o1=1 o2=1 output="x1*x2" > test.rsf<br />
bash$ < test.rsf sfdisfil<br />
0: 1 2 3 4 5<br />
5: 2 4 6 8 10<br />
10: 3 6 9 12 15<br />
Scale every data point by 2:
76 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ < test.rsf sfscale dscale=2 | sfdisfil<br />
0: 2 4 6 8 10<br />
5: 4 8 12 16 20<br />
10: 6 12 18 24 30<br />
Divide every trace by its maximum value:<br />
bash$ < test.rsf sfscale axis=1 | sfdisfil<br />
0: 0.2 0.4 0.6 0.8 1<br />
5: 0.2 0.4 0.6 0.8 1<br />
10: 0.2 0.4 0.6 0.8 1<br />
Divide by the maximum value in the whole 2-D dataset:<br />
bash$ < test.rsf sfscale axis=2 | sfdisfil<br />
0: 0.06667 0.1333 0.2 0.2667 0.3333<br />
5: 0.1333 0.2667 0.4 0.5333 0.6667<br />
10: 0.2 0.4 0.6 0.8 1<br />
The rscale= parameter is synonymous to dscale= except when it is equal to zero.<br />
With sfscale dscale=0, the dataset gets multiplied by zero. If using rscale=0,<br />
the other parameters are used to define scaling. Thus, sfscale rscale=0 axis=1<br />
is equivalent to sfscale axis=1, and sfscale rscale=0 is equivalent to sfscale<br />
dscale=1.<br />
sfspike: Generate simple data: spikes, boxes, planes, constants.<br />
sfspike < in.rsf > spike.rsf mag= nsp=1 k#=[0,...] l#=[k1,k2,...] p#=[0,...]<br />
n#= o#=[0,0,...] d#=[0.004,0.1,0.1,...] label#=[Time,Distance,Distance,...]<br />
unit#=[s,km,km,...] title=<br />
Spike positioning is given in samples and starts with 1.<br />
float d#=[0.004,0.1,0.1,...] sampling on #-th axis<br />
ints k#=[0,...] spike starting position [nsp]<br />
ints l#=[k1,k2,...] spike ending position [nsp]<br />
string<br />
label#=[Time,Distance,Distance,...] label on #-th axis<br />
floats mag= spike magnitudes [nsp]<br />
int n#= size of #-th axis<br />
int nsp=1 Number of spikes<br />
float o#=[0,0,...] origin on #-th axis<br />
floats p#=[0,...] spike inclination (in samples) [nsp]<br />
string title= title for plots<br />
string unit#=[s,km,km,...] unit on #-th axis
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 77<br />
sfspike takes no input and generates an output with “spikes”. It is an easy way<br />
to create data. Here is an example:<br />
bash$ sfspike n1=5 n2=3 k1=4 k2=1 | sfdisfil<br />
0: 0 0 0 1 0<br />
5: 0 0 0 0 0<br />
10: 0 0 0 0 0<br />
The spike location is specified by parameters k1=4 and k2=1. Note that the locations<br />
are numbered starting from 1. If one of the parameters is omitted or given the value<br />
of zero, the spike in the corresponding direction becomes a plane:<br />
bash$ sfspike n1=5 n2=3 k1=4 | sfdisfil<br />
0: 0 0 0 1 0<br />
5: 0 0 0 1 0<br />
10: 0 0 0 1 0<br />
If no spike parameters are given, the whole dataset is filled with ones:<br />
bash$ sfspike n1=5 n2=3 | sfdisfil<br />
0: 1 1 1 1 1<br />
5: 1 1 1 1 1<br />
10: 1 1 1 1 1<br />
To create several spikes, use the nsp= parameter and give a comma-separated list<br />
of values to k#= arguments:<br />
bash$ sfspike n1=5 n2=3 nsp=3 k1=1,3,4 k2=1,2,3 | sfdisfil<br />
0: 1 0 0 0 0<br />
5: 0 0 1 0 0<br />
10: 0 0 0 1 0<br />
If the number of values in the list is smaller than nsp, the last value gets repeated,<br />
and the spikes add on top of each other, creating larger amplitudes:<br />
bash$ sfspike n1=5 n2=3 nsp=3 k1=1,3 k2=1,2 | sfdisfil<br />
0: 1 0 0 0 0<br />
5: 0 0 2 0 0<br />
10: 0 0 0 0 0<br />
The magnitude of the spikes can be controlled explicitly with the mag= parameter:
78 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ sfspike n1=5 n2=3 nsp=3 k1=1,3,4 k2=1,2,3 mag=1,4,2 | sfdisfil<br />
0: 1 0 0 0 0<br />
5: 0 0 4 0 0<br />
10: 0 0 0 2 0<br />
You can create boxes instead of spikes by using l#= parameters:<br />
bash$ sfspike n1=5 n2=3 k1=2 l1=4 k2=2 mag=8 | sfdisfil<br />
0: 0 0 0 0 0<br />
5: 0 8 8 8 0<br />
10: 0 0 0 0 0<br />
In this case, k1=2 specifies the box start, and l1=4 specifies the box end.<br />
Finally, multi-dimensional planes can be given an inclination by using p#= parameters:<br />
bash$ sfspike n1=5 n2=3 k1=2 p2=1 | sfdisfil<br />
0: 0 1 0 0 0<br />
5: 0 0 1 0 0<br />
10: 0 0 0 1 0<br />
When the inclination value is not integer, simple linear interpolation is used:<br />
bash$ sfspike n1=5 n2=3 k1=2 p2=0.7 | sfdisfil<br />
0: 0 1 0 0 0<br />
5: 0 0.3 0.7 0 0<br />
10: 0 0 0.6 0.4 0<br />
sfspike supplies default dimensions and labels to all axis:<br />
bash$ sfspike n1=5 n2=3 n3=4 > spike.rsf<br />
bash$ sfin spike.rsf<br />
spike.rsf:<br />
in="/var/tmp/spike.rsf@"<br />
esize=4 type=float form=native<br />
n1=5 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
n3=4 d3=0.1 o3=0 label3="Distance" unit3="km"<br />
60 elements 240 bytes<br />
As you can see, the first axis is assumed to be time, with sampling of 0.004 seconds.<br />
All other axes are assumed to be distance, with sampling of 0.1 kilometers. All these<br />
parameters can be changed on the command line.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 79<br />
bash$ sfspike n1=5 n2=3 n3=4 label3=Offset unit3=ft d3=20 > spike.rsf<br />
bash$ sfin spike.rsf<br />
spike.rsf:<br />
in="/var/tmp/spike.rsf@"<br />
esize=4 type=float form=native<br />
n1=5 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=3 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
n3=4 d3=20 o3=0 label3="Offset" unit3="ft"<br />
60 elements 240 bytes<br />
sfspray: Extend a dataset by duplicating in the specified axis<br />
dimension.<br />
sfspray < in.rsf > out.rsf axis=2 n= d= o= label= unit=<br />
This operation is adjoint to sfstack.<br />
int axis=2 which axis to spray<br />
float d= Sampling of the newly created dimension<br />
string label= Label of the newly created dimension<br />
int n= Size of the newly created dimension<br />
float o= Origin of the newly created dimension<br />
string unit= Units of the newly created dimension<br />
sfspray extends the input hypercube by replicating the data in one of the dimensions.<br />
The output dataset acquires one additional dimension. Here is an example:<br />
Start with a 2-D dataset<br />
bash$ sfmath n1=5 n2=2 output=x1+x2 > test.rsf<br />
bash$ sfin test.rsf<br />
test.rsf:<br />
in="/var/tmp/test.rsf@"<br />
esize=4 type=float form=native<br />
n1=5 d1=1 o1=0<br />
n2=2 d2=1 o2=0<br />
10 elements 40 bytes<br />
bash$ < test.rsf sfdisfil<br />
0: 0 1 2 3 4<br />
5: 1 2 3 4 5<br />
Extend the data in the second dimension<br />
bash$ < test.rsf sfspray axis=2 n=3 > test2.rsf<br />
bash$ sfin test2.rsf
80 Fomel <strong>Madagascar</strong> Documentation<br />
test2.rsf:<br />
in="/var/tmp/test2.rsf@"<br />
esize=4 type=float form=native<br />
n1=5 d1=1 o1=0<br />
n2=3 d2=1 o2=0<br />
n3=2 d3=1 o3=0<br />
30 elements 120 bytes<br />
bash$ < test2.rsf sfdisfil<br />
0: 0 1 2 3 4<br />
5: 0 1 2 3 4<br />
10: 0 1 2 3 4<br />
15: 1 2 3 4 5<br />
20: 1 2 3 4 5<br />
25: 1 2 3 4 5<br />
The output is three-dimensional, with traces from the original data duplicated along<br />
the second axis.<br />
Extend the data in the third dimension<br />
bash$ < test.rsf sfspray axis=3 n=2 > test3.rsf<br />
bash$ sfin test3.rsf<br />
test3.rsf:<br />
in="/var/tmp/test3.rsf@"<br />
esize=4 type=float form=native<br />
n1=5 d1=1 o1=0<br />
n2=2 d2=1 o2=0<br />
n3=2 d3=? o3=?<br />
20 elements 80 bytes<br />
bash$ < test3.rsf sfdisfil<br />
0: 0 1 2 3 4<br />
5: 1 2 3 4 5<br />
10: 0 1 2 3 4<br />
15: 1 2 3 4 5<br />
The output is also three-dimensional, with the original data replicated along the third<br />
axis.
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 81<br />
sfstack: Stack a dataset over one of the dimensions.<br />
sfstack < in.rsf > out.rsf scale= axis=2 rms=n norm=y min=n max=n prod=n<br />
This operation is adjoint to sfspray.<br />
int axis=2 which axis to stack<br />
bool max=n [y/n] If y, find maximum instead of stack. Ignores<br />
rms and norm.<br />
bool min=n [y/n] If y, find minimum instead of stack. Ignores<br />
rms and norm.<br />
bool norm=y [y/n] If y, normalize by fold.<br />
bool prod=n [y/n] If y, find product instead of stack. Ignores<br />
rms and norm.<br />
bool rms=n [y/n] If y, compute the root-mean-square instead<br />
of stack.<br />
floats scale= optionally scale before stacking [n2]<br />
While sfspray adds a dimension to a hypercube, sfstack effectively removes one<br />
of the dimensions by stacking over it. Here are some examples:<br />
bash$ sfmath n1=5 n2=3 output=x1+x2 > test.rsf<br />
bash$ < test.rsf sfdisfil<br />
0: 0 1 2 3 4<br />
5: 1 2 3 4 5<br />
10: 2 3 4 5 6<br />
bash$ < test.rsf sfstack axis=2 | sfdisfil<br />
0: 1.5 2 3 4 5<br />
bash$ < test.rsf sfstack axis=1 | sfdisfil<br />
0: 2.5 3 4<br />
Why is the first value not 1 (in the first case) or 2 (in the second case)? By default,<br />
sfstack normalizes the stack by the fold (the number of non-zero entries). To avoid<br />
normalization, use norm=n, as follows:<br />
bash$ < test.rsf sfstack norm=n | sfdisfil<br />
0: 3 6 9 12 15<br />
sfstack can also compute root-mean-square values as well as minimum and maximum<br />
values.<br />
bash$ < test.rsf sfstack rms=y | sfdisfil<br />
0: 1.581 2.16 3.109 4.082 5.066<br />
bash$ < test.rsf sfstack min=y | sfdisfil<br />
0: 0 1 2 3 4<br />
bash$ < test.rsf sfstack axis=1 max=y | sfdisfil<br />
0: 4 5 6
82 Fomel <strong>Madagascar</strong> Documentation<br />
sftransp: Transpose two axes in a dataset.<br />
sftransp < in.rsf > out.rsf memsize=sf memsize() plane=<br />
If you get a "Cannot allocate memory" error, give the program a<br />
memsize=1 command-line parameter to force out-of-core operation.<br />
int memsize=sf memsize() Max amount of RAM (in Mb) to be used<br />
int plane= Two-digit number with axes to transpose.<br />
The default is 12<br />
The sftransp program transposes the input hypercube exchanging the two axes<br />
specified by the plane= parameter.<br />
bash$ sfspike n1=10 n2=20 n3=30 > orig123.rsf<br />
bash$ sfin orig123.rsf<br />
orig123.rsf:<br />
in="/var/tmp/orig123.rsf@"<br />
esize=4 type=float form=native<br />
n1=10 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=20 d2=0.1 o2=0 label2="Distance2" unit2="km"<br />
n3=30 d3=0.1 o3=0 label3="Distance3" unit3="km"<br />
6000 elements 24000 bytes<br />
bash$ out132.rsf<br />
bash$ sfin out132.rsf<br />
out132.rsf:<br />
in="/var/tmp/out132.rsf@"<br />
esize=4 type=float form=native<br />
n1=10 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=30 d2=0.1 o2=0 label2="Distance3" unit2="km"<br />
n3=20 d3=0.1 o3=0 label3="Distance2" unit3="km"<br />
6000 elements 24000 bytes<br />
bash$ out321.rsf<br />
bash$ sfin out321.rsf<br />
out321.rsf:<br />
in="/var/tmp/out132.rsf@"<br />
esize=4 type=float form=native<br />
n1=30 d1=0.1 o1=0 label1="Distance" unit1="km"<br />
n2=20 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
n3=10 d3=0.004 o3=0 label3="Time" unit3="s"<br />
6000 elements 24000 bytes<br />
sftransp tries to fit the dataset in memory to transpose it there but, if not enough<br />
memory is available, it performs a slower transpose out of core using disk operations.<br />
You can control the amount of available memory using the memsize= parameter or
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 83<br />
the RSFMEMSIZE environmental variable.<br />
sfwindow: Window a portion of a dataset.<br />
sfwindow < in.rsf > out.rsf verb=n squeeze=y j#=(1,...) d#=(d1,d2,...)<br />
f#=(0,...) min#=(o1,o2,,...) n#=(0,...) max#=(o1+(n1-1)*d1,o2+(n1-1)*d2,,...)<br />
float d#=(d1,d2,...) sampling in #-th dimension<br />
largeint f#=(0,...) window start in #-th dimension<br />
int j#=(1,...) jump in #-th dimension<br />
float<br />
max#=(o1+(n1-<br />
maximum in #-th dimension<br />
1)*d1,o2+(n1-<br />
1)*d2,,...)<br />
float min#=(o1,o2,,...) minimum in #-th dimension<br />
largeint n#=(0,...) window size in #-th dimension<br />
bool squeeze=y [y/n] if y, squeeze dimensions equal to 1 to the<br />
end<br />
bool verb=n [y/n] Verbosity flag<br />
sfwindow is used to window a portion of the dataset. Here is a quick example:<br />
Start by creating some data.<br />
bash$ sfmath n1=5 n2=3 o1=1 o2=1 output="x1*x2" > test.rsf<br />
bash$ < test.rsf sfdisfil<br />
0: 1 2 3 4 5<br />
5: 2 4 6 8 10<br />
10: 3 6 9 12 15<br />
Now window the first two rows:<br />
bash$ < test.rsf sfwindow n2=2 | sfdisfil<br />
0: 1 2 3 4 5<br />
5: 2 4 6 8 10<br />
Window the first three columns:<br />
bash$ < test.rsf sfwindow n1=3 | sfdisfil<br />
0: 1 2 3 2 4<br />
5: 6 3 6 9<br />
Window the middle row:<br />
bash$ < test.rsf sfwindow f2=1 n2=1 | sfdisfil<br />
0: 2 4 6 8 10<br />
You can interpret the f# and n# parameters as meaning ”skip that many rows/-<br />
columns” and ”select that many rows/columns” correspondingly. Window the middle<br />
point in the dataset:
84 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ < test.rsf sfwindow f1=2 n1=1 f2=1 n2=1 | sfdisfil<br />
0: 6<br />
Window every other column:<br />
bash$ < test.rsf sfwindow j1=2 | sfdisfil<br />
0: 1 3 5 2 6<br />
5: 10 3 9 15<br />
Window every third column:<br />
bash$ < test.rsf sfwindow j1=3 | sfdisfil<br />
0: 1 4 2 8 3<br />
5: 12<br />
Alternatively, sfwindow can use the minimum and maximum parameters to select<br />
a window. In the following example, we are creating a dataset with sfspike and<br />
then windowing a portion of it between 1 and 2 seconds in time and sampled at 8<br />
miliseconds.<br />
bash$ sfspike n1=1000 n2=10 > spike.rsf<br />
bash$ sfin spike.rsf<br />
spike.rsf:<br />
in="/var/tmp/spike.rsf@"<br />
esize=4 type=float form=native<br />
n1=1000 d1=0.004 o1=0 label1="Time" unit1="s"<br />
n2=10 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
10000 elements 40000 bytes<br />
bash$ < spike.rsf sfwindow min1=1 max1=2 d1=0.008 > window.rsf<br />
bash$ sfin window.rsfwindow.rsf:<br />
in="/var/tmp/window.rsf@"<br />
esize=4 type=float form=native<br />
n1=126 d1=0.008 o1=1 label1="Time" unit1="s"<br />
n2=10 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
1260 elements 5040 bytes<br />
By default, sfwindow “squeezes” the hypercube dimensions that are equal to one<br />
toward the end of the dataset. Here is an example of taking a time slice:<br />
bash$ < spike.rsf sfwindow n1=1 min1=1 > slice.rsf<br />
bash$ sfin slice.rsf<br />
slice.rsf:<br />
in="/var/tmp/slice.rsf@"
<strong>Madagascar</strong> Documentation <strong>Madagascar</strong> programs 85<br />
esize=4 type=float form=native<br />
n1=10 d1=0.1 o1=0 label1="Distance" unit1="km"<br />
n2=1 d2=0.004 o2=1 label2="Time" unit2="s"<br />
10 elements 40 bytes<br />
You can change this behavior by specifying squeeze=n.<br />
bash$ < spike.rsf sfwindow n1=1 min1=1 squeeze=n > slice.rsf<br />
bash$ sfin slice.rsf slice.rsf:<br />
in="/var/tmp/slice.rsf@"<br />
esize=4 type=float form=native<br />
n1=1 d1=0.004 o1=1 label1="Time" unit1="s"<br />
n2=10 d2=0.1 o2=0 label2="Distance" unit2="km"<br />
10 elements 40 bytes<br />
REFERENCES<br />
Claerbout, J., 1998, Multidimensional recursive filters via a helix: Geophysics, 63,<br />
1532–1541.
86 Fomel <strong>Madagascar</strong> Documentation
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Guide to RSF format<br />
Sergey Fomel 1<br />
ABSTRACT<br />
This guide explains the RSF <strong>file</strong> format.<br />
PRINCIPLES<br />
The main design principle behind the RSF <strong>file</strong> format is KISS (“Keep It Simple,<br />
Stupid!”). The RSF format is borrowed from the SEPlib data format originally<br />
designed at the Stanford Exploration Project (Claerbout, 1991). The format is made<br />
as simple as possible for maximum convenience, transparency and flexibility.<br />
According to the Unix tradition, common <strong>file</strong> formats should be in a readable<br />
textual form so that they can be easily examined and processed with universal tools.<br />
Raymond (2004) writes:<br />
To design a perfect anti-Unix, make all <strong>file</strong> formats binary and opaque,<br />
and require heavyweight tools to read and edit them.<br />
If you feel an urge to design a complex binary <strong>file</strong> format, or a complex<br />
binary application protocol, it is generally wise to lie down until the feeling<br />
passes.<br />
Storing large-scale datasets in a text format may not be economical. RSF chooses<br />
the next best thing: it allows data values to be stored in a binary format but puts all<br />
data attributes in text <strong>file</strong>s that can be read by humans and processed with universal<br />
text-processing utilities.<br />
Example<br />
Let us first create some synthetic RSF data.<br />
bash$ sfmath n1=1000 output=’sin(0.5*x1)’ > sin.rsf<br />
Open and read the <strong>file</strong> sin.rsf.<br />
1 e-mail: sergey.fomel@beg.utexas.edu<br />
87
88 Fomel <strong>Madagascar</strong> Documentation<br />
bash$ cat sin.rsf<br />
sfmath rsf/rsf/rsftour: fomels@egl Sun Jul 31 07:18:48 2005<br />
o1=0<br />
data_format="native_float"<br />
esize=4<br />
in="/tmp/sin.rsf@"<br />
x1=0<br />
d1=1<br />
n1=1000<br />
The <strong>file</strong> contains nine lines with simple readable text. The first line shows the name<br />
of the program, the working directory, the user and computer that created the <strong>file</strong> and<br />
the time it was created (that information is recorded for accounting purposes). Other<br />
lines contain parameter-value pairs separated by the “=” sign. The “in” parameter<br />
points to the location of the binary data. Before we discuss the meaning of parameters<br />
in more detail, let us plot the data.<br />
bash$ < sin.rsf<br />
sfwiggle title=’One Trace’ | sfpen<br />
On your screen, you should see a plot similar to Figure 1.<br />
Suppose you want to reformat the data so that instead of one trace of a thousand<br />
samples, it contains twenty traces with fifty samples each. Try running<br />
bash$ < sin.rsf sed ’s/n1=1000/n1=100 n2=10/’ > sin10.rsf<br />
bash$ < sin10.rsf sfwiggle title=Traces | sfpen<br />
or (using pipes)<br />
bash$ < sin.rsf sed ’s/n1=1000/n1=50 n2=20/’ | sfwiggle title=Traces | sfpen<br />
On your screen, you should see a plot similar to Figure 2.<br />
What happened? We used sed, a standard Unix line editing utility to change<br />
the parameters describing the data dimensions. Because of the simplicity of this<br />
operation, there is no need to create specialized data formatting tools or to make the<br />
sfwiggle program accept additional formatting parameters. Other general-purpose<br />
Unix tools that can be applied on RSF <strong>file</strong>s include cat, echo, grep, etc.<br />
An alternative way to obtain the previous result is to run<br />
bash$ ( cat sin.rsf; echo n1=50 n2=20 ) > sin10.rsf<br />
bash$ < sin10.rsf sfwiggle title=Traces | sfpen
<strong>Madagascar</strong> Documentation RSF format 89<br />
Figure 1: An example sinusoid plot. rsf/format sin1
90 Fomel <strong>Madagascar</strong> Documentation<br />
Figure 2: An example sinusoid plot, with data reformatted to twenty traces.<br />
rsf/format sin2
<strong>Madagascar</strong> Documentation RSF format 91<br />
In this case, the cat utility simply copies the contents of the previous <strong>file</strong>, and the<br />
echo utility appends new line “n1=50 n2=20”. A new value of the n1 parameter<br />
overwrites the old value of n1=1000, and we achieve the same result as before.<br />
Of course, one could also edit the <strong>file</strong> by hand with one of the general purpose<br />
text editors. For recording the history of data processing, it is usually preferable to<br />
be able to process <strong>file</strong>s with non-interactive tools.<br />
HEADER AND DATA FILES<br />
A simple way to check the layout of an RSF <strong>file</strong> is with the sfin program.<br />
bash$ sfin sin10.rsf<br />
sin10.rsf:<br />
in="/tmp/sin.rsf@"<br />
esize=4 type=float form=native<br />
n1=50 d1=1 o1=0<br />
n2=20 d2=? o2=?<br />
1000 elements 4000 bytes<br />
The program reports the following information: the location of the data <strong>file</strong> (/tmp/sin.rsf),<br />
the element size (4 bytes), the element type (floating point), the element form (native),<br />
the hypercube dimensions (50 × 20), axis scaling (1 and unspecified), and axis<br />
origin (0 and unspecified). It also checks the total number of elements and bytes in<br />
the data <strong>file</strong>.<br />
Let us examine this information in detail. First, we can verify that the data <strong>file</strong><br />
exists and contains the specified number of bytes:<br />
bash$ ls -l /tmp/sin.rsf@<br />
-rw-r--r-- 1 sergey users 4000 2004-10-04 00:35 /tmp/sin.rsf@<br />
4000 bytes in this <strong>file</strong> are required to store 50 × 20 floating-point 4-byte numbers in<br />
a binary form. Thus, the data <strong>file</strong> contains nothing but the raw data in a contiguous<br />
binary form.<br />
Datapath<br />
How did the RSF program (sfmath) decide where to put the data <strong>file</strong>? In the order<br />
of priority, the rules for selecting the data <strong>file</strong> name and the data <strong>file</strong> directory are as<br />
follows:<br />
1. Check out= parameter on the command line. The parameter specifies the output<br />
data <strong>file</strong> location explicitly.
92 Fomel <strong>Madagascar</strong> Documentation<br />
2. Specify the path and the <strong>file</strong> name separately.<br />
• The rules for the path selection are:<br />
(a) Check datapath= parameter on the command line. The parameter<br />
specifies a string to prepend to the <strong>file</strong> name. The string may contain<br />
the <strong>file</strong> directory.<br />
(b) Check DATAPATH environmental variable. It has the same meaning as<br />
the parameter specified with datapath=.<br />
(c) Check for .datapath <strong>file</strong> in the current directory. The <strong>file</strong> may contain<br />
a line<br />
datapath=/path/to_<strong>file</strong>/<br />
or<br />
machine_name datapath=/path/to_<strong>file</strong>/<br />
if you indent to use different paths on different platforms.<br />
(d) Check for .datapath <strong>file</strong> in the user home directory.<br />
(e) Put the data <strong>file</strong> in the current directory (similar to datapath=./).<br />
• The rules for the <strong>file</strong>name selection are:<br />
(a) If the output RSF <strong>file</strong> is in the current directory, the name of the data<br />
<strong>file</strong> is made by appending .<br />
(b) If the output <strong>file</strong> is not in the current directory or if it is created<br />
temporarily by a program, the name is made by appending random<br />
characters to the name of the program and selected to be unique.<br />
Examples:<br />
•<br />
bash$ sfspike n1=10 out=test1 > spike.rsf<br />
bash$ grep in spike.rsf<br />
in="test1"<br />
•<br />
bash$ sfspike n1=10 datapath=/tmp/ > spike.rsf<br />
bash$ grep in spike.rsf<br />
in="/tmp/spike.rsf@"<br />
•
<strong>Madagascar</strong> Documentation RSF format 93<br />
bash$ DATAPATH=/tmp/ sfspike n1=10 > spike.rsf<br />
bash$ grep in spike.rsf<br />
in="/tmp/spike.rsf@"<br />
•<br />
bash$ sfspike n1=10 datapath=/tmp/ > /tmp/spike.rsf<br />
bash$ grep in /tmp/spike.rsf<br />
in="/tmp/sfspikejcARVf"<br />
Packing header and data together<br />
While the header and data <strong>file</strong>s are separated by default, it is also possible to pack<br />
them together into one <strong>file</strong>. To do that, specify the program’s “out” parameter as<br />
out=stdout. Example:<br />
bash$ sfspike n1=10 out=stdout > spike.rsf<br />
bash$ grep in spike.rsf<br />
Binary <strong>file</strong> spike.rsf matches<br />
bash$ sfin spike.rsf<br />
spike.rsf:<br />
in="stdin"<br />
esize=4 type=float form=native<br />
n1=10 d1=0.004 o1=0 label1="Time" unit1="s"<br />
10 elements 40 bytes<br />
bash$ ls -l spike.rsf<br />
-rw-r--r-- 1 sergey users 196 2004-11-10 21:39 spike.rsf<br />
If you examine the contents of spike.rsf, you will find that it starts with the text<br />
header information, followed by special symbols, followed by binary data.<br />
Packing headers and data together may not be a good idea for data processing<br />
but it works well for storing data: it is easier to move the packed <strong>file</strong> around than to<br />
move two different <strong>file</strong>s (header and binary) together while remembering to preserve<br />
their connection. Packing header and data together is also the current mechanism<br />
used to push RSF <strong>file</strong>s through Unix pipes.<br />
Type<br />
The data stored with RSF can have different types: character, unsigned character,<br />
integer, floating point, or complex. By default, <strong>single</strong> precision is used for numbers<br />
(int and float data types in the C programming language). The number of bytes<br />
required for represent these numbers may depend on the platform.
94 Fomel <strong>Madagascar</strong> Documentation<br />
Form<br />
The data stored with RSF can also be in a different form: ASCII, native binary, and<br />
XDR binary. Native binary is often used by default. It is the binary format employed<br />
by the machine that is running the application. On Linux-running PC, the native<br />
binary format will typically correspond to the so-called little-endian byte ordering.<br />
On some other platform, it might be big-endian ordering. XDR is a binary format<br />
designed by Sun for exchanging <strong>file</strong>s over network. It typically corresponds to bigendian<br />
byte ordering. It is more efficient to process RSF <strong>file</strong>s in the native binary<br />
format but, if you intend to access data from different platforms, it might be a good<br />
idea to store the corresponding <strong>file</strong> in an XDR format. RSF also allows for an ASCII<br />
(plain text) form of data <strong>file</strong>s.<br />
Conversion between different types and forms is accomplished with sfdd program.<br />
Here are some examples. First, let us create synthetic data.<br />
bash$ sfmath n1=10 output=’10*sin(0.5*x1)’ > sin.rsf<br />
bash$ sfin sin.rsf<br />
sin.rsf:<br />
in="/tmp/sin.rsf@"<br />
esize=4 type=float form=native<br />
n1=10 d1=1 o1=0<br />
10 elements 40 bytes<br />
bash$ < sin.rsf sfdisfil<br />
0: 0 4.794 8.415 9.975 9.093<br />
5: 5.985 1.411 -3.508 -7.568 -9.775<br />
Converting the data to the integer type:<br />
bash$ < sin.rsf sfdd type=int > isin.rsf<br />
bash$ sfin isin.rsf<br />
isin.rsf:<br />
in="/tmp/isin.rsf@"<br />
esize=4 type=int form=native<br />
n1=10 d1=1 o1=0<br />
10 elements 40 bytes<br />
bash$ < isin.rsf sfdisfil<br />
0: 0 4 8 9 9 5 1 -3 -7 -9<br />
Converting the data to the ASCII form:<br />
bash$ < sin.rsf sfdd form=ascii > asin.rsf<br />
bash$ < asin.rsf sfdisfil<br />
0: 0 4.794 8.415 9.975 9.093
<strong>Madagascar</strong> Documentation RSF format 95<br />
5: 5.985 1.411 -3.508 -7.568 -9.775<br />
bash$ sfin asin.rsf<br />
asin.rsf:<br />
in="/tmp/asin.rsf@"<br />
esize=0 type=float form=ascii<br />
n1=10 d1=1 o1=0<br />
10 elements<br />
bash$ cat /tmp/asin.rsf@<br />
0 4.79426 8.41471 9.97495 9.09297 5.98472 1.4112 -3.50783<br />
-7.56803 -9.7753<br />
Hypercube<br />
While RSF stores binary data in a contiguous 1-D array, the conceptual data model<br />
is a multidimensional hypercube. By convention, the dimensions of the cube are<br />
defined with parameters n1, n2, n3, etc. The fastest axis is n1. Additionally, the<br />
grid sampling can be given by parameters d1, d2, d3, etc. The axes origins are given<br />
by parameters o1, o2, o3, etc. Optionally, you can also supply the axis label strings<br />
label1, label2, label3, etc., and axis units strings unit1, unit2, unit3, etc.<br />
COMPATIBILITY WITH OTHER FILE FORMATS<br />
It is possible to exchange RSF-formatted data with other popular data formats.<br />
Compatibility with SEPlib<br />
RSF is mostly compatible with its predecessor, the SEPlib <strong>file</strong> format. However, there<br />
are several important differences:<br />
1. SEPlib program typically use the element size (esize= parameter) to distinguish<br />
between different data types: esize=4 corresponds to floating point data, while<br />
esize=8 corresponds to complex data. The typical type handling mechanism<br />
in RSF is different: RSF looks at data format= to determine the data type.<br />
2. The default data form in SEPlib programs is typically XDR and not native as<br />
it is in RSF.<br />
3. It is possible to pipe the output of RSF programs to SEPlib:<br />
bash$ sfspike n1=1 | Attr want=min<br />
minimum value = 1 at 1
96 Fomel <strong>Madagascar</strong> Documentation<br />
However, piping the output of SEPlib programs to RSF (or, for that matter,<br />
any other non-SEPlib programs) will result in an unterminated process. Do not<br />
try<br />
bash$ Spike n1=1 | sfattr want=ming<br />
That happens because SEPlib uses sockets for piping and expects a socket<br />
connection from the receiving program. RSF passes data through regular Unix<br />
pipes.<br />
4. SEP3D is an extension of SEPlib for operating with irregularly sampled data<br />
(Biondi et al., 1996). There is no equivalent of it in RSF for the reasons explained<br />
in the beginning of this guide. Operations with irregular datasets are<br />
supported through the use of auxiliary input <strong>file</strong>s that represent the geometry<br />
information.<br />
Reading and writing SEG-Y and SU <strong>file</strong>s<br />
The SEG-Y format is based on the proposal of Barry et al. (1975). It was revised in<br />
2002 2 . The SU format is a modification of SEG-Y used in Seismic Unix (Stockwell,<br />
1997).<br />
To convert <strong>file</strong>s from SEG-Y or SU format to RSF, use the sfsegyread program.<br />
Let us first manufacture an example <strong>file</strong> using SU utilities (Stockwell, 1999):<br />
bash$ suplane > plane.su<br />
bash$ segyhdrs < plane.su | segywrite tape=plane.segy<br />
To convert it to RSF, use either<br />
bash$ sfsuread < plane.su t<strong>file</strong>=t<strong>file</strong>.rsf endian=0 > plane.rsf<br />
or<br />
bash$ sfsegyread < plane.segy t<strong>file</strong>=t<strong>file</strong>.rsf \<br />
h<strong>file</strong>=h<strong>file</strong> b<strong>file</strong>=b<strong>file</strong> endian=0 > plane.rsf<br />
The endian flag is needed if the SU <strong>file</strong> originated from a little-endian machine such<br />
as Linux PC.<br />
Several <strong>file</strong>s are generated. The standard output contains an RSF <strong>file</strong> with the<br />
data (32 traces with 64 samples each):<br />
2 See http://seg.org/publications/tech-stand/seg_y_rev1.pdf.
<strong>Madagascar</strong> Documentation RSF format 97<br />
bash$ sfin plane.rsf<br />
plane.rsf:<br />
in="/tmp/plane.rsf@"<br />
esize=4 type=float form=native<br />
n1=64 d1=0.004 o1=0<br />
n2=32 d2=? o2=?<br />
2048 elements 8192 bytes<br />
The contents of this <strong>file</strong> are displayed in Figure 3. The t<strong>file</strong> is an RSF integer-type<br />
<strong>file</strong> with the trace headers (32 headers with 71 traces each):<br />
bash$ sfin t<strong>file</strong>.rsf<br />
t<strong>file</strong>.rsf:<br />
in="/tmp/t<strong>file</strong>.rsf@"<br />
esize=4 type=int form=native<br />
n1=71 d1=? o1=?<br />
n2=32 d2=? o2=?<br />
2272 elements 9088 bytes<br />
The contents of trace headers can be quickly examined with the sfheaderattr program.<br />
The h<strong>file</strong> is the ASCII header <strong>file</strong> for the whole record.<br />
bash$ head -c 242 h<strong>file</strong><br />
C This tape was made at the<br />
C<br />
C Center for Wave Phenomena<br />
The b<strong>file</strong> is the binary header <strong>file</strong>.<br />
To convert <strong>file</strong>s back from RSF to SEG-Y or SU, use the sfsegywrite program<br />
and reverse the input and output:<br />
bash$ sfsuwrite > plane.su t<strong>file</strong>=t<strong>file</strong>.rsf endian=0 < plane.rsf<br />
or<br />
bash$ sfsegywrite > plane.segy t<strong>file</strong>=t<strong>file</strong>.rsf \<br />
h<strong>file</strong>=h<strong>file</strong> b<strong>file</strong>=b<strong>file</strong> endian=0 < plane.rsf<br />
If h<strong>file</strong>= and b<strong>file</strong>= are not supplied to sfsegywrite, the corresponding headers<br />
will be either picked from the default locations (<strong>file</strong>s named header and binary) or<br />
generated on the fly. The trace header <strong>file</strong> can be generated with sfsegyheader.<br />
Here is an example:
98 Fomel <strong>Madagascar</strong> Documentation<br />
Figure 3: The output of suplane, converted to RSF and displayed with sfwiggle.<br />
rsf/format plane
<strong>Madagascar</strong> Documentation RSF format 99<br />
bash$ rm header binary<br />
bash$ sfheadermath < plane.rsf output=N+1 | sfdd type=int > tracl.rsf<br />
bash$ sfsegyheader < plane.rsf tracl=tracl.rsf > t<strong>file</strong>.rsf<br />
bash$ sfsegywrite < plane.rsf t<strong>file</strong>=t<strong>file</strong>.rsf > plane.segy<br />
Reading and writing ASCII <strong>file</strong>s<br />
Reading and writing ASCII <strong>file</strong>s can be accomplished with the sfdd program. For<br />
example, let us take an ASCII <strong>file</strong> with numbers<br />
bash$ cat <strong>file</strong>.asc<br />
1.0 1.5 3.0<br />
4.8 9.1 7.3<br />
Converting it to RSF is as simple as<br />
bash$ echo in=<strong>file</strong>.asc n1=3 n2=2 data_format=ascii_float > <strong>file</strong>.rsf<br />
bash$ sfin <strong>file</strong>.rsf<br />
<strong>file</strong>.rsf:<br />
in="<strong>file</strong>.asc"<br />
esize=0 type=float form=ascii<br />
n1=3 d1=? o1=?<br />
n2=2 d2=? o2=?<br />
6 elements<br />
For more efficient input/output operations, it might be advantageous to convert the<br />
data type to native binary, as follows:<br />
bash$ echo in=<strong>file</strong>.asc n1=3 n2=2 data_format=ascii_float | \<br />
sfdd form=native > <strong>file</strong>.rsf<br />
bash$ sfin <strong>file</strong>.rsf<br />
<strong>file</strong>.rsf:<br />
in="/tmp/<strong>file</strong>.rsf@"<br />
esize=4 type=float form=native<br />
n1=3 d1=? o1=?<br />
n2=2 d2=? o2=?<br />
6 elements 24 bytes<br />
Convert from RSF to ASCII is equally simple:<br />
bash$ sfdd form=ascii out=<strong>file</strong>.asc < <strong>file</strong>.rsf > /dev/null<br />
bash$ cat <strong>file</strong>.asc<br />
1 1.5 3 4.8 9.1 7.3
100 Fomel <strong>Madagascar</strong> Documentation<br />
You can use the line= and format= parameters in sfdd to control the ASCII formatting:<br />
bash$ sfdd form=ascii out=<strong>file</strong>.asc \<br />
line=3 format="%3.1f " < <strong>file</strong>.rsf > /dev/null<br />
bash$ cat <strong>file</strong>.asc<br />
1.0 1.5 3.0<br />
4.8 9.1 7.3<br />
An alternative is to use sfdisfil.<br />
bash$ sfdisfil > <strong>file</strong>.asc col=3 format="%3.1f " number=n < <strong>file</strong>.rsf<br />
bash$ cat <strong>file</strong>.asc<br />
1.0 1.5 3.0<br />
4.8 9.1 7.3<br />
OTHER DOCUMENTATION<br />
This note should give you a general understanding of the RSF <strong>file</strong> format. Other<br />
relevant documentation is<br />
• Introduction to RSF<br />
• Installation instructions<br />
• Self-documentation reference for RSF programs<br />
• A guide to RSF programs<br />
• A guide to RSF programming interface<br />
• A guide to programming with RSF<br />
• A tour of RSF software<br />
• A guide to SCons interface for reproducible computations<br />
REFERENCES<br />
Barry, K. M., D. A. Cavers, and C. W. Kneale, 1975, Report on recommended standards<br />
for digital tape formats: Geophysics, 40, 344–352.<br />
Biondi, B., R. Clapp, and S. Crawley, 1996, Seplib90: Seplib for 3-D prestack data,<br />
in SEP-92: Stanford Exploration Project, 343–364.<br />
Claerbout, J. F., 1991, Introduction to Seplib and SEP utility software, in SEP-70:<br />
Stanford Exploration Project, 413–436.
<strong>Madagascar</strong> Documentation RSF format 101<br />
Raymond, E. S., 2004, The art of UNIX programming: Addison-Wesley.<br />
Stockwell, J. W., 1997, Free software in education: A case study of CWP/SU: Seismic<br />
Unix: The Leading Edge, 16, 1045–1049.<br />
——–, 1999, The CWP/SU: Seismic Un*x package: Computers and Geosciences, 25,<br />
415–419.
102 Fomel <strong>Madagascar</strong> Documentation
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Revisiting SEP tour with <strong>Madagascar</strong> and SCons<br />
Sergey Fomel 1<br />
ABSTRACT<br />
Many appreciative users were introduced to SEPlib (Claerbout, 1991) by an excellent<br />
article of Dellinger and Tálas (1992). In this paper, I show how to create<br />
a similar experience using <strong>Madagascar</strong> and SCons.<br />
GETTING STARTED<br />
Similarly to SU and SEPlib, RSF programs can be piped and executed from the<br />
command line, for example:<br />
bash$ sfspike n1=1000 k1=300 title="\s200 Welcome to \c2 RSF" | \<br />
sfbandpass fhi=2 phase=1 | sfwiggle | sfpen<br />
If you are already familiar with SEPlib, you can find most of the familiar programs<br />
with the names prepended by “sf”.<br />
Typing a command without arguments, should produce a concise self-documentation.<br />
bash$ sfbandpass<br />
The recommended way of using RSF, however, is not with the command line but<br />
with SCons and “SConstruct” <strong>file</strong>s.<br />
Setting up<br />
Open a <strong>file</strong> named “SConstruct” in your favorite editor and start it with a line<br />
5 from r s f . p r o j import ∗<br />
This line tells Python to load the RSF project module.<br />
1 e-mail: sergey.fomel@beg.utexas.edu<br />
103
104 Fomel <strong>Madagascar</strong> Documentation<br />
Obtaining the test data<br />
Add a Fetch command as follows:<br />
11 Fetch ( ’Txx .HH’ , ’ septour ’ )<br />
Now, by running<br />
bash$ scons Txx.HH<br />
you can instruct SCons to connect to an anonymous data server and extract (fetch)<br />
the data <strong>file</strong> “Txx.HH” from the “septour” directory.<br />
Displaying the data<br />
Add the following line to the SConstruct <strong>file</strong>:<br />
17 Result ( ’ wiggle0 ’ , ’Txx .HH’ , ’ wiggle ’ )<br />
Note that it does not matter if this line appears before or after the “Fetch” line.<br />
You are simply instructing SCons how to create a result plot from the input.<br />
Run<br />
bash$ scons wiggle0.view<br />
If everything is setup correctly in your environment, you should see something like<br />
the following output in your terminal:<br />
bash$ scons wiggle0.view<br />
scons: Reading SConscript <strong>file</strong>s ...<br />
scons: done reading SConscript <strong>file</strong>s.<br />
scons: Building targets ...<br />
retrieve(["Txx.HH"], [])<br />
< Txx.HH /path/to/RSF/bin/sfwiggle > Fig/wiggle0.vpl<br />
/path/to/RSF/bin/sfpen Fig/wiggle0.vpl<br />
and a figure similar to Figure 1 appearing on your screen.<br />
Windowing and plotting<br />
PROCESSING EXERCISES<br />
Our next task is to window and plot a significant portion of the data.<br />
following line to the SConstruct <strong>file</strong>:<br />
Add the
<strong>Madagascar</strong> Documentation RSF tour 105<br />
Figure 1: To see this figure on your screen, run scons wiggle0.view<br />
rsf/rsftour wiggle0
106 Fomel <strong>Madagascar</strong> Documentation<br />
23 Flow ( ’ windowed ’ , ’Txx .HH’ , ’ window n2=10 min1=0.4 max1=0.8 ’ )<br />
The window command selects the first ten traces and the time window between<br />
0.4 and 0.8 seconds.<br />
We will plot the windowed data with three different plotting programs.<br />
25 p l o t p a r = ’ ’ ’<br />
26 transp=y poly=y y r e v e r s e=y p c l i p =100 nc=20 a l l p o s=n<br />
27 unit2=km unit1=s l a b e l 1=Time l a b e l 2=O f f s e t<br />
28 ’ ’ ’<br />
29<br />
30 for p l o t in ( ’ wiggle ’ , ’ contour ’ , ’ grey ’ ) :<br />
For convenience, plotting parameters are put in a string called plotpar. A Python<br />
string can be enclosed in <strong>single</strong>, double, or triple quotes. Triple quotes allow the<br />
string to span multiple lines. In this case, we use triple quotes for convenience. Next,<br />
we loop (using Python’s for construct) through three different programs (wiggle,<br />
contour, and grey). For each program, the command portion of Result is formed<br />
by concatenating two strings with Python’s addition operator.<br />
Try running scons -Q wiggle.view. You should see something like the following<br />
output in your terminal:<br />
bash$ scons -Q wiggle.view<br />
< Txx.HH /path/to/RSF/bin/sfwindow n2=10 n1=200 f1=200 > windowed.rsf<br />
< windowed.rsf /path/to/RSF/bin/sfwiggle transp=y poly=y yreverse=y<br />
pclip=100 nc=200 > Fig/wiggle.vpl<br />
/path/to/RSF/bin/sfpen Fig/wiggle.vpl<br />
and a figure similar to Figure 2 appearing on your screen. The -Q switch tells SCons<br />
to run in a quiet mode, suppressing verbose comments. We will use it from now on<br />
to save space. You can dismiss the figure by using the “q” key on the keyboard or by<br />
hitting the “quit” button.<br />
Run scons -Q view, and you should see simply<br />
bash$ scons -Q view<br />
/path/to/RSF/bin/sfpen Fig/wiggle.vpl<br />
Since the wiggle.vpl figure is up to date, SCons does not rebuild it. After quitting<br />
the figure, SCons will resume processing with<br />
< windowed.rsf /path/to/RSF/bin/sfcontour transp=y poly=y yreverse=y<br />
pclip=100 nc=200 > Fig/contour.vpl<br />
/path/to/RSF/bin/sfpen Fig/contour.vpl
<strong>Madagascar</strong> Documentation RSF tour 107<br />
and a figure similar to Figure 3 appearing on your screen. Quitting the figure, produces<br />
< windowed.rsf /path/to/RSF/bin/sfgrey transp=y poly=y yreverse=y<br />
pclip=100 nc=200 > Fig/grey.vpl<br />
/path/to/RSF/bin/sfpen Fig/grey.vpl<br />
and Figure 4.<br />
Figure 2: To see this figure on your screen, run scons wiggle.view<br />
rsf/rsftour wiggle<br />
Resampling<br />
The next example demonstrated simple signal processing using the Fast Fourier Transform.<br />
We will first subsample the original data and then recover the data using Fourier<br />
interpolation.
108 Fomel <strong>Madagascar</strong> Documentation<br />
Figure 3: To see this figure on your screen, run scons contour.view<br />
rsf/rsftour contour
<strong>Madagascar</strong> Documentation RSF tour 109<br />
Figure 4: To see this figure on your screen, run scons grey.view rsf/rsftour grey
110 Fomel <strong>Madagascar</strong> Documentation<br />
Subsampling is accomplished with sfwindow.<br />
36<br />
37 # decimate time a x i s by two<br />
Running scons -Q subsampled.rsf produces<br />
< windowed.rsf /path/to/RSF/bin/sfwindow j1=2 > subsampled.rsf<br />
We can verify that the size of the first axis has decreased by running<br />
sfin windowed.rsf subsampled.rsf.<br />
Try also sfwiggle < subsampled.rsf | sfpen to quickly inspect the subsampled<br />
data on the screen.<br />
To interpolate the data back to the original sampling, the following sequence of<br />
steps can be applied:<br />
1. Fourier transform from time domain to frequency domain.<br />
2. Pad the frequency axis<br />
3. Inverse Fourier transform from frequency to time.<br />
All three steps are conveniently combined into one using pipes.<br />
39<br />
40 # s i n c i n t e r p o l a t i o n in the Fourier domain<br />
41 Flow ( ’ resampled ’ , ’ subsampled ’ ,<br />
Why do we pad the Fourier domain to 102? The time length of the original data<br />
is 201 samples. In the frequency domain, it can be represented with 101 positive<br />
frequencies plus the zero frequency, which amounts to 102. Note that the output of<br />
sffft1 does not contain negative frequencies.<br />
Finally, we display the result. The reconstructed data is shown in Figure 5.<br />
Comparing this result with Figure 2, we can verify a fairly accurate reconstruction.<br />
As an exercise, try subsampling the data by a factor of 4 and see if you can still<br />
reconstruct the original data with the Fourier method.
<strong>Madagascar</strong> Documentation RSF tour 111<br />
Figure 5: To see this figure on your screen, run scons resampled.view<br />
rsf/rsftour resampled
112 Fomel <strong>Madagascar</strong> Documentation<br />
Normal Moveout<br />
The next example applies a simple constant-velocity NMO correction to the windowed<br />
data and pipes the result to a wiggle plotting command:<br />
49<br />
50 Result ( ’nmo ’ , ’ windowed ’ ,<br />
51 ’ ’ ’<br />
52 nmostretch v0 =2.05 h a l f=n |<br />
53 wiggle p c l i p =100 max1=0.6 poly=y<br />
Running scons -Q nmo.view produces<br />
< windowed.rsf /path/to/RSF/bin/sfnmostretch v0=2.05 half=n |<br />
/path/to/RSF/bin/sfwiggle pclip=100 max1=0.6 poly=y > Fig/nmo.vpl<br />
/path/to/RSF/bin/sfpen Fig/nmo.vpl<br />
and Figure 6. Note that SCons does not recreate the windowed.rsf <strong>file</strong> if that <strong>file</strong> is<br />
up to date. You can experiment with the NMO velocity (2.05 km/s) or with plotting<br />
parameters to get different results. As Dellinger and Tálas (1992) point out, the<br />
NMO velocity of 2.05 km/s “appears to split the difference between two distinctly<br />
non-hyperbolic shear waves”.<br />
Advanced plotting<br />
Sometimes, we need to combine different plots either by overlaying them on top of<br />
each other or by putting them side by side. Here is an example of accomplishing it<br />
with RSF and SCons.<br />
Start by creating common plotting plotting arguments and plotting the data in<br />
greyscale.<br />
59<br />
60 p l o t p a r = p l o t p a r+’ min1=.4 max1=.8 max2=1. min2=.05 poly=n ’<br />
61<br />
62 Plot ( ’ grey ’ , ’ windowed ’ ,<br />
Next, plot the wiggle traces twice: the fist time, using thick black lines (plotcol=0<br />
plotfat=10), and the second time, using thinner white lines (plotcol=7 plotfat=5).<br />
63 ’ grey w h e r e t i t l e=t wherexlabel=b ’ + p l o t p a r )<br />
64 Plot ( ’ wiggle1 ’ , ’ windowed ’ ,<br />
65 ’ wiggle p l o t c o l =0 p l o t f a t =10 ’ + p l o t p a r )<br />
66 Plot ( ’ wiggle2 ’ , ’ windowed ’ ,
<strong>Madagascar</strong> Documentation RSF tour 113<br />
Figure 6: To see this figure on your screen, run scons nmo.view rsf/rsftour nmo
114 Fomel <strong>Madagascar</strong> Documentation<br />
The plots are combined by overlaying or by putting them side by side.<br />
68<br />
69 Result ( ’ o v e r p l o t ’ , ’ grey wiggle1 wiggle2 ’ , ’ Overlay ’ )<br />
The resultant plots are shown in Figures 7 and 8.<br />
Figure 7: To see this figure on your screen, run scons overplot.view<br />
rsf/rsftour overplot<br />
CONCLUSIONS<br />
This tour is not designed as a comprehensive manual. It simply gives a glimpse into<br />
working in a reproducible research environment with RSF and SCons. The reader is<br />
encouraged to experiment with the SConstruct <strong>file</strong> attached to this tour and included<br />
in the Appendix. For other documentation on RSF, please see<br />
• Introduction to RSF
<strong>Madagascar</strong> Documentation RSF tour 115<br />
Figure 8: To see this figure on your screen, run scons sidebyside.view<br />
rsf/rsftour sidebyside<br />
• Installation instructions<br />
• Self-documentation reference for RSF programs<br />
• A guide to RSF programs<br />
• A guide to RSF <strong>file</strong> format<br />
• A guide to RSF programming interface<br />
• A guide to programming with RSF<br />
• A guide to SCons interface for reproducible computations<br />
ACKNOWLEDGMENTS<br />
Thanks to Joe Dellinger and Sándor Tálas for creating “SEP tour” and to James<br />
Rickett for updating it. Several generations of SEP students contributed to SEPlib.<br />
We try to preserve all their good ideas when refactoring SEPlib into RSF.<br />
The test dataset used in this paper is courtesy of Beltram Nolte and L. Neil Frazer.<br />
REFERENCES<br />
Claerbout, J. F., 1991, Introduction to Seplib and SEP utility software, in SEP-70:<br />
Stanford Exploration Project, 413–436.<br />
Dellinger, J., and S. Tálas, 1992, A tour of SEPlib for new users, in SEP-73: Stanford<br />
Exploration Project, 461–502.
116 Fomel <strong>Madagascar</strong> Documentation<br />
SCONSTRUCT FILE<br />
Here is a complete listing of the SConstruct <strong>file</strong> used in this example.<br />
1 #########################################################<br />
2 # S e t t i n g up<br />
3 #########################################################<br />
4<br />
5 from r s f . p r o j import ∗<br />
6<br />
7 #########################################################<br />
8 # Obtaining the t e s t data<br />
9 #########################################################<br />
10<br />
11 Fetch ( ’Txx .HH’ , ’ septour ’ )<br />
12<br />
13 #########################################################<br />
14 # D i s p l a y i n g the data<br />
15 #########################################################<br />
16<br />
17 Result ( ’ wiggle0 ’ , ’Txx .HH’ , ’ wiggle ’ )<br />
18<br />
19 #########################################################<br />
20 # Windowing and p l o t t i n g<br />
21 #########################################################<br />
22<br />
23 Flow ( ’ windowed ’ , ’Txx .HH’ , ’ window n2=10 min1=0.4 max1=0.8 ’ )<br />
24<br />
25 p l o t p a r = ’ ’ ’<br />
26 transp=y poly=y y r e v e r s e=y p c l i p =100 nc=20 a l l p o s=n<br />
27 unit2=km unit1=s l a b e l 1=Time l a b e l 2=O f f s e t<br />
28 ’ ’ ’<br />
29<br />
30 for p l o t in ( ’ wiggle ’ , ’ contour ’ , ’ grey ’ ) :<br />
31 Result ( plot , ’ windowed ’ , p l o t + p l o t p a r )<br />
32<br />
33 #########################################################<br />
34 # Resampling<br />
35 #########################################################<br />
36<br />
37 # decimate time a x i s by two<br />
38 Flow ( ’ subsampled ’ , ’ windowed ’ , ’ window j 1=2 ’ )<br />
39<br />
40 # s i n c i n t e r p o l a t i o n in the Fourier domain<br />
41 Flow ( ’ resampled ’ , ’ subsampled ’ ,
<strong>Madagascar</strong> Documentation RSF tour 117<br />
42 ’ f f t 1 | pad n1=102 | f f t 1 inv=y opt=n | window max1=0.8 ’ )<br />
43<br />
44 Result ( ’ resampled ’ , ’ wiggle t i t l e=Resampled ’ + p l o t p a r )<br />
45<br />
46 #########################################################<br />
47 # V e l o c i t y a n a l y s i s and NMO<br />
48 #########################################################<br />
49<br />
50 Result ( ’nmo ’ , ’ windowed ’ ,<br />
51 ’ ’ ’<br />
52 nmostretch v0 =2.05 h a l f=n |<br />
53 wiggle p c l i p =100 max1=0.6 poly=y<br />
54 ’ ’ ’ )<br />
55<br />
56 #########################################################<br />
57 # Advanced p l o t t i n g<br />
58 #########################################################<br />
59<br />
60 p l o t p a r = p l o t p a r+’ min1=.4 max1=.8 max2=1. min2=.05 poly=n ’<br />
61<br />
62 Plot ( ’ grey ’ , ’ windowed ’ ,<br />
63 ’ grey w h e r e t i t l e=t wherexlabel=b ’ + p l o t p a r )<br />
64 Plot ( ’ wiggle1 ’ , ’ windowed ’ ,<br />
65 ’ wiggle p l o t c o l =0 p l o t f a t =10 ’ + p l o t p a r )<br />
66 Plot ( ’ wiggle2 ’ , ’ windowed ’ ,<br />
67 ’ wiggle p l o t c o l =7 p l o t f a t =3 ’ + p l o t p a r )<br />
68<br />
69 Result ( ’ o v e r p l o t ’ , ’ grey wiggle1 wiggle2 ’ , ’ Overlay ’ )<br />
70 Result ( ’ s i d e b y s i d e ’ , ’ grey wiggle2 ’ , ’ SideBySideIso ’ )<br />
71<br />
72 #########################################################<br />
73 # Wrapping up<br />
74 #########################################################<br />
75<br />
76 End ( )
118 Fomel <strong>Madagascar</strong> Documentation
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Guide to RSF API<br />
Sergey Fomel 1<br />
ABSTRACT<br />
This guide explains the RSF programming interface.<br />
INTRODUCTION<br />
To work with RSF <strong>file</strong>s in your own programs, you may need to use an appropriate<br />
programming interface. We will demonstrate the interface in different languages using<br />
a simple example. The example is a clipping program. It reads and writes RSF <strong>file</strong>s<br />
and accesses parameters both from the input <strong>file</strong> and the command line. The input<br />
is processed trace by trace. This is not necessarily the most efficient approach 2 but<br />
it suffices for a simple demonstration.<br />
The C clip function is listed below.<br />
1 /∗ Clip the data . ∗/<br />
2<br />
3 #include <br />
4<br />
C INTERFACE<br />
5 int main ( int argc , char∗ argv [ ] )<br />
6 {<br />
7 int n1 , n2 , i1 , i 2 ;<br />
8 float c l i p , ∗ t r a c e ;<br />
9 s f f i l e in , out ; /∗ Input and output f i l e s ∗/<br />
10<br />
11 /∗ I n i t i a l i z e RSF ∗/<br />
12 s f i n i t ( argc , argv ) ;<br />
13 /∗ standard input ∗/<br />
14 in = s f i n p u t ( ” in ” ) ;<br />
15 /∗ standard output ∗/<br />
16 out = s f o u t p u t ( ” out ” ) ;<br />
17<br />
1 e-mail: sergey.fomel@beg.utexas.edu<br />
2 Compare with the library clip program.<br />
119
120 Fomel <strong>Madagascar</strong> Documentation<br />
18 /∗ check t h a t the input i s f l o a t ∗/<br />
19 i f (SF FLOAT != s f g e t t y p e ( in ) )<br />
20 s f e r r o r ( ”Need f l o a t input ” ) ;<br />
21<br />
22 /∗ n1 i s the f a s t e s t dimension ( t r a c e l e n g t h ) ∗/<br />
23 i f ( ! s f h i s t i n t ( in , ”n1”,&n1 ) )<br />
24 s f e r r o r ( ”No n1= in input ” ) ;<br />
25 /∗ l e f t s i z e g e t s n2∗n3∗n4 ∗ . . . ( the number of t r a c e s ) ∗/<br />
26 n2 = s f l e f t s i z e ( in , 1 ) ;<br />
27<br />
28 /∗ parameter from the command l i n e ( i . e . c l i p =1.5 ) ∗/<br />
29 i f ( ! s f g e t f l o a t ( ” c l i p ”,& c l i p ) ) s f e r r o r ( ”Need c l i p=” ) ;<br />
30<br />
31 /∗ a l l o c a t e f l o a t i n g p o i n t array ∗/<br />
32 t r a c e = s f f l o a t a l l o c ( n1 ) ;<br />
33<br />
34 /∗ loop over t r a c e s ∗/<br />
35 for ( i 2 =0; i 2 < n2 ; i 2++) {<br />
36<br />
37 /∗ read a t r a c e ∗/<br />
38 s f f l o a t r e a d ( trace , n1 , in ) ;<br />
39<br />
40 /∗ loop over samples ∗/<br />
41 for ( i 1 =0; i 1 < n1 ; i 1++) {<br />
42 i f ( t r a c e [ i 1 ] > c l i p ) t r a c e [ i 1 ]= c l i p ;<br />
43 else i f ( t r a c e [ i 1 ] < −c l i p ) t r a c e [ i 1 ]=− c l i p ;<br />
44 }<br />
45<br />
46 /∗ w r i t e a t r a c e ∗/<br />
47 s f f l o a t w r i t e ( trace , n1 , out ) ;<br />
48 }<br />
49<br />
50<br />
51 e x i t ( 0 ) ;<br />
52 }<br />
Let us examine it in detail.<br />
3 #include <br />
The include preprocessing directive is required to access the RSF interface.<br />
9 s f f i l e in , out ; /∗ Input and output f i l e s ∗/<br />
RSF data <strong>file</strong>s are defined with an abstract sf <strong>file</strong> data type. An abstract data<br />
type means that the contents of it are not publicly declared, and all operations on
<strong>Madagascar</strong> Documentation RSF API 121<br />
sf <strong>file</strong> objects should be performed with library functions. This is analogous to<br />
FILE * data type used in stdio.h and as close as C gets to an object-oriented style<br />
of programming (Roberts, 1998).<br />
11 /∗ I n i t i a l i z e RSF ∗/<br />
12 s f i n i t ( argc , argv ) ;<br />
Before using any of the other functions, you must call sf init. This function parses<br />
the command line and initializes an internally stored table of command-line parameters.<br />
13 /∗ standard input ∗/<br />
14 in = s f i n p u t ( ” in ” ) ;<br />
15 /∗ standard output ∗/<br />
16 out = s f o u t p u t ( ” out ” ) ;<br />
The input and output RSF <strong>file</strong> objects are created with sf input and sf output<br />
constructor functions. Both these functions take a string argument. The string<br />
may refer to a <strong>file</strong> name or a <strong>file</strong> tag. For example, if the command line contains<br />
vel=velocity.rsf, then both sf input("velocity.rsf") and sf input("vel")<br />
are acceptable. Two tags are special: "in" refers to the <strong>file</strong> in the standard input<br />
and "out" refers to the <strong>file</strong> in the standard output.<br />
18 /∗ check t h a t the input i s f l o a t ∗/<br />
19 i f (SF FLOAT != s f g e t t y p e ( in ) )<br />
20 s f e r r o r ( ”Need f l o a t input ” ) ;<br />
RSF <strong>file</strong>s can store data of different types (character, integer, floating point, complex).<br />
We extract the data type of the input <strong>file</strong> with the library sf gettype function and<br />
check if it represents floating point numbers. If not, the program is aborted with an<br />
error message, using the sf error function. It is generally a good idea to check the<br />
input for user errors and, if they cannot be corrected, to take a safe exit.<br />
22 /∗ n1 i s the f a s t e s t dimension ( t r a c e l e n g t h ) ∗/<br />
23 i f ( ! s f h i s t i n t ( in , ”n1”,&n1 ) )<br />
24 s f e r r o r ( ”No n1= in input ” ) ;<br />
25 /∗ l e f t s i z e g e t s n2∗n3∗n4 ∗ . . . ( the number of t r a c e s ) ∗/<br />
26 n2 = s f l e f t s i z e ( in , 1 ) ;<br />
Conceptually, the RSF data model is a multidimensional hypercube. By convention,<br />
the dimensions of the cube are stored in n1=, n2=, etc. parameters. The n1 parameter<br />
refers to the fastest axis. If the input dataset is a collection of traces, n1 refers to the<br />
trace length. We extract it using the sf histint function (integer parameter from<br />
history) and abort if no value for n1 is found. We could proceed in a similar fashion,<br />
extracting n2, n3, etc. If we are interested in the total number of traces, like in the clip<br />
example, a shortcut is to use the sf leftsize function. Calling sf leftsize(in,0)
122 Fomel <strong>Madagascar</strong> Documentation<br />
returns the total number of elements in the hypercube (the product of n1, n2, etc.),<br />
calling sf leftsize(in,1) returns the number of traces (the product of n2, n3,<br />
etc.), calling sf leftsize(in,2) returns the product of n3, n4, etc. By calling<br />
sf leftsize, we avoid the need to extract additional parameters for the hypercube<br />
dimensions that we are not interested in.<br />
28 /∗ parameter from the command l i n e ( i . e . c l i p =1.5 ) ∗/<br />
29 i f ( ! s f g e t f l o a t ( ” c l i p ”,& c l i p ) ) s f e r r o r ( ”Need c l i p=” ) ;<br />
The clip parameter is read from the command line, where it can be specified, for<br />
example, as clip=10. The parameter has the float type, therefore we read it with<br />
the sf getfloat function. If no clip= parameter is found among the command<br />
line arguments, the program is aborted with an error message using the sf error<br />
function.<br />
31 /∗ a l l o c a t e f l o a t i n g p o i n t array ∗/<br />
32 t r a c e = s f f l o a t a l l o c ( n1 ) ;<br />
Next, we allocate an array of floating-point numbers to store a trace with the library<br />
sf floatalloc function. Unlike the standard malloc the RSF allocation function<br />
checks for errors and either terminates the program or returns a valid pointer.<br />
34 /∗ loop over t r a c e s ∗/<br />
35 for ( i 2 =0; i 2 < n2 ; i 2++) {<br />
36<br />
37 /∗ read a t r a c e ∗/<br />
38 s f f l o a t r e a d ( trace , n1 , in ) ;<br />
39<br />
40 /∗ loop over samples ∗/<br />
41 for ( i 1 =0; i 1 < n1 ; i 1++) {<br />
42 i f ( t r a c e [ i 1 ] > c l i p ) t r a c e [ i 1 ]= c l i p ;<br />
43 else i f ( t r a c e [ i 1 ] < −c l i p ) t r a c e [ i 1 ]=− c l i p ;<br />
44 }<br />
45<br />
46 /∗ w r i t e a t r a c e ∗/<br />
47 s f f l o a t w r i t e ( trace , n1 , out ) ;<br />
48 }<br />
The rest of the program is straightforward. We loop over all available traces, read<br />
each trace, clip it and right the output out. The syntax of sf floatread and<br />
sf floatwrite functions is similar to the syntax of the C standard fread and fwrite<br />
function except that the type of the element is specified explicitly in the function name<br />
and that the input and output <strong>file</strong>s have the RSF type sf <strong>file</strong>.
<strong>Madagascar</strong> Documentation RSF API 123<br />
Compiling<br />
To compile the clip program, run<br />
cc clip.c -I$RSFROOT/include -L$RSFROOT/lib -lrsf -lm<br />
Change cc to the C compiler appropriate for your system and include additional<br />
compiler flags if necessary. The flags that RSF typically uses are in<br />
$RSFROOT/share/madagascar/etc/config.py.<br />
The C++ clip function is listed below.<br />
1 /∗ Clip the data . ∗/<br />
2<br />
3 #include <br />
4 #include <br />
5<br />
C++ INTERFACE<br />
6 int main ( int argc , char∗ argv [ ] )<br />
7 {<br />
8 s f i n i t ( argc , argv ) ; // I n i t i a l i z e RSF<br />
9<br />
10 iRSF par ( 0 ) , in ; // input parameter , f i l e<br />
11 oRSF out ; // output f i l e<br />
12<br />
13 int n1 , n2 ; // t r a c e length , number of t r a c e s<br />
14 float c l i p ;<br />
15<br />
16 in . get ( ”n1” , n1 ) ;<br />
17 n2=in . s i z e ( 1 ) ;<br />
18<br />
19 par . get ( ” c l i p ” , c l i p ) ; // parameter from the command l i n e<br />
20<br />
21 std : : valarray t r a c e ( n1 ) ;<br />
22<br />
23 for ( int i 2 =0; i 2 < n2 ; i 2++) { // loop over t r a c e s<br />
24 in >> t r a c e ; // read a t r a c e<br />
25<br />
26 for ( int i 1 =0; i 1 < n1 ; i 1++) { // loop over samples<br />
27 i f ( t r a c e [ i 1 ] > c l i p ) t r a c e [ i 1 ]= c l i p ;<br />
28 else i f ( t r a c e [ i 1 ] < −c l i p ) t r a c e [ i 1 ]=− c l i p ;<br />
29 }<br />
30
124 Fomel <strong>Madagascar</strong> Documentation<br />
31 out > t r a c e ; // read a t r a c e<br />
25<br />
26 for ( int i 1 =0; i 1 < n1 ; i 1++) { // loop over samples<br />
27 i f ( t r a c e [ i 1 ] > c l i p ) t r a c e [ i 1 ]= c l i p ;
<strong>Madagascar</strong> Documentation RSF API 125<br />
28 else i f ( t r a c e [ i 1 ] < −c l i p ) t r a c e [ i 1 ]=− c l i p ;<br />
29 }<br />
30<br />
31 out 1000) then<br />
18 c a l l s f e r r o r ( ”n1 i s too long ” )<br />
19 end i f<br />
20 n2 = s f l e f t s i z e ( in , 1 )
126 Fomel <strong>Madagascar</strong> Documentation<br />
21<br />
22 i f ( . not . s f g e t f l o a t ( ” c l i p ” , c l i p ) )<br />
23 & c a l l s f e r r o r ( ”Need c l i p=” )<br />
24<br />
25 do 10 i 2 =1, n2<br />
26 c a l l s f f l o a t r e a d ( trace , n1 , in )<br />
27<br />
28 do 20 i 1 =1, n1<br />
29 i f ( t r a c e ( i 1 ) > c l i p ) then<br />
30 t r a c e ( i 1 )= c l i p<br />
31 else i f ( t r a c e ( i 1 ) < −c l i p ) then<br />
32 t r a c e ( i 1)=− c l i p<br />
33 end i f<br />
34 20 continue<br />
35<br />
36 c a l l s f f l o a t w r i t e ( trace , n1 , out )<br />
37 10 continue<br />
38<br />
39 stop<br />
40 end<br />
Let us examine it in detail.<br />
8 c a l l s f i n i t ( )<br />
The program starts with a call to sf init, which initializes the command-line interface.<br />
9 in = s f i n p u t ( ” in ” )<br />
10 out = s f o u t p u t ( ” out ” )<br />
The input and output <strong>file</strong>s are created with calls to sf input and sf output. Because<br />
of the absence of derived types in Fortran-77, we use simple integer pointers to<br />
represent RSF <strong>file</strong>s. Both sf input and sf output accept a character string, which<br />
may refer to a <strong>file</strong> name or a <strong>file</strong> tag. For example, if the command line contains<br />
vel=velocity.rsf, then both sf input("velocity.rsf") and sf input("vel")<br />
are acceptable. Two tags are special: "in" refers to the <strong>file</strong> in the standard input<br />
and "out" refers to the <strong>file</strong> in the standard output.<br />
12 i f (3 . ne . s f g e t t y p e ( in ) )<br />
13 & c a l l s f e r r o r ( ”Need f l o a t input ” )<br />
RSF <strong>file</strong>s can store data of different types (character, integer, floating point, complex).<br />
The function sf gettype checks the type of data stored in the RSF <strong>file</strong>. We make<br />
sure that the type corresponds to floating-point numbers. If not, the program is<br />
aborted with an error message, using the sf error function. It is generally a good
<strong>Madagascar</strong> Documentation RSF API 127<br />
idea to check the input for user errors and, if they cannot be corrected, to take a safe<br />
exit.<br />
15 i f ( . not . s f h i s t i n t ( in , ”n1” , n1 ) ) then<br />
16 c a l l s f e r r o r ( ”No n1= in input ” )<br />
17 else i f ( n1 > 1000) then<br />
18 c a l l s f e r r o r ( ”n1 i s too long ” )<br />
19 end i f<br />
20 n2 = s f l e f t s i z e ( in , 1 )<br />
Conceptually, the RSF data model is a multidimensional hypercube. By convention,<br />
the dimensions of the cube are stored in n1=, n2=, etc. parameters. The n1 parameter<br />
refers to the fastest axis. If the input dataset is a collection of traces, n1 refers to<br />
the trace length. We extract it using the sf histint function (integer parameter<br />
from history) and abort if no value for n1 is found. Since Fortran-77 cannot easily<br />
handle dynamic allocation, we also need to check that n1 is not larger than the size<br />
of the statically allocated array. We could proceed in a similar fashion, extracting n2,<br />
n3, etc. If we are interested in the total number of traces, like in the clip example,<br />
a shortcut is to use the sf leftsize function. Calling sf leftsize(in,0) returns<br />
the total number of elements in the hypercube (the product of n1, n2, etc.), calling<br />
sf leftsize(in,1) returns the number of traces (the product of n2, n3, etc.), calling<br />
sf leftsize(in,2) returns the product of n3, n4, etc. By calling sf leftsize, we<br />
avoid the need to extract additional parameters for the hypercube dimensions that<br />
we are not interested in.<br />
22 i f ( . not . s f g e t f l o a t ( ” c l i p ” , c l i p ) )<br />
23 & c a l l s f e r r o r ( ”Need c l i p=” )<br />
The clip parameter is read from the command line, where it can be specified, for<br />
example, as clip=10. The parameter has the float type, therefore we read it with<br />
the sf getfloat function. If no clip= parameter is found among the command<br />
line arguments, the program is aborted with an error message using the sf error<br />
function.<br />
25 do 10 i 2 =1, n2<br />
26 c a l l s f f l o a t r e a d ( trace , n1 , in )<br />
27<br />
28 do 20 i 1 =1, n1<br />
29 i f ( t r a c e ( i 1 ) > c l i p ) then<br />
30 t r a c e ( i 1 )= c l i p<br />
31 else i f ( t r a c e ( i 1 ) < −c l i p ) then<br />
32 t r a c e ( i 1)=− c l i p<br />
33 end i f<br />
34 20 continue<br />
35<br />
36 c a l l s f f l o a t w r i t e ( trace , n1 , out )
128 Fomel <strong>Madagascar</strong> Documentation<br />
37 10 continue<br />
Finally, we do the actual work: loop over input traces, reading, clipping, and writing<br />
out each trace.<br />
Compiling<br />
To compile the Fortran-77 program, run<br />
f77 clip.f -L$RSFROOT/lib -lrsff -lrsf -lm<br />
Change f77 to the Fortran compiler appropriate for your system and include additional<br />
compiler flags if necessary. The flags that RSF typically uses are in<br />
$RSFROOT/share/madagascar/etc/config.py.<br />
FORTRAN-90 INTERFACE<br />
The Fortran-90 clip function is listed below.<br />
1 program C l i p i t<br />
2 use r s f<br />
3<br />
4 implicit none<br />
5 type ( f i l e ) : : in , out<br />
6 integer : : n1 , n2 , i1 , i 2<br />
7 real : : c l i p<br />
8 real , dimension ( : ) , allocatable : : t r a c e<br />
9<br />
10 c a l l s f i n i t ( ) ! i n i t i a l i z e RSF<br />
11 in = r s f i n p u t ( )<br />
12 out = r s f o u t p u t ( )<br />
13<br />
14 i f ( s f f l o a t /= gettype ( in ) ) c a l l s f e r r o r ( ”Need f l o a t s ” )<br />
15<br />
16 c a l l from par ( in , ”n1” , n1 )<br />
17 n2 = f i l e s i z e ( in , 1 )<br />
18<br />
19 c a l l from par ( ” c l i p ” , c l i p ) ! command−l i n e parameter<br />
20<br />
21 allocate ( t r a c e ( n1 ) )<br />
22<br />
23 do i 2 =1, n2 ! loop over t r a c e s<br />
24 c a l l r s f r e a d ( in , t r a c e )
<strong>Madagascar</strong> Documentation RSF API 129<br />
25<br />
26 where ( t r a c e > c l i p ) t r a c e = c l i p<br />
27 where ( t r a c e < −c l i p ) t r a c e = −c l i p<br />
28<br />
29 c a l l r s f w r i t e ( out , t r a c e )<br />
30 end do<br />
31 end program C l i p i t<br />
Let us examine it in detail.<br />
2 use r s f<br />
The program starts with importing the rsf module.<br />
10 c a l l s f i n i t ( ) ! i n i t i a l i z e RSF<br />
A call to sf init is needed to initialize the command-line interface.<br />
11 in = r s f i n p u t ( )<br />
12 out = r s f o u t p u t ( )<br />
The standard input and output <strong>file</strong>s are initialized with rsf input and rsf output<br />
functions. Both functions accept optional arguments. For example, if the command<br />
line contains vel=velocity.rsf, then both rsf input("velocity.rsf") and<br />
rsf input("vel") are acceptable.<br />
14 i f ( s f f l o a t /= gettype ( in ) ) c a l l s f e r r o r ( ”Need f l o a t s ” )<br />
A call to from par extracts the “n1” parameter from the input <strong>file</strong>. Conceptually,<br />
the RSF data model is a multidimensional hypercube. The n1 parameter refers to<br />
the fastest axis. If the input dataset is a collection of traces, n1 corresponds to the<br />
trace length. We could proceed in a similar fashion, extracting n2, n3, etc. If we are<br />
interested in the total number of traces, like in the clip example, a shortcut is to use<br />
the <strong>file</strong>size function. Calling <strong>file</strong>size(in) returns the total number of elements<br />
in the hypercube (the product of n1, n2, etc.), calling <strong>file</strong>size(in,1) returns the<br />
number of traces (the product of n2, n3, etc.), calling <strong>file</strong>size(in,2) returns the<br />
product of n3, n4, etc. By calling <strong>file</strong>size, we avoid the need to extract additional<br />
parameters for the hypercube dimensions that we are not interested in.<br />
17 n2 = f i l e s i z e ( in , 1 )<br />
The clip parameter is read from the command line, where it can be specified, for<br />
example, as clip=10. If we knew a good default value for clip, we could specify it<br />
with an optional argument, i.e. call from par("clip",clip,default).<br />
21 allocate ( t r a c e ( n1 ) )<br />
22
130 Fomel <strong>Madagascar</strong> Documentation<br />
23 do i 2 =1, n2 ! loop over t r a c e s<br />
24 c a l l r s f r e a d ( in , t r a c e )<br />
25<br />
26 where ( t r a c e > c l i p ) t r a c e = c l i p<br />
27 where ( t r a c e < −c l i p ) t r a c e = −c l i p<br />
Finally, we do the actual work: loop over input traces, reading, clipping, and writing<br />
out each trace.<br />
Compiling<br />
To compile the Fortran-90 program, run<br />
f90 clip.f90 -I$RSFROOT/include -L$RSFROOT/lib -lrsff90 -lrsf -lm<br />
Change f90 to the Fortran-90 compiler appropriate for your system and include additional<br />
compiler flags if necessary. The flags that RSF typically uses are in<br />
$RSFROOT/share/madagascar/etc/config.py.<br />
The Python clip script is listed below.<br />
1 #!/ usr / bin /env python<br />
2<br />
3 import numpy<br />
4 import m8r<br />
5<br />
6 par = m8r . Par ( )<br />
7 inp = m8r . Input ( )<br />
8 output = m8r . Output ( )<br />
9 a s s e r t ’ f l o a t ’ == inp . type<br />
10<br />
11 n1 = inp . i n t ( ”n1” )<br />
12 n2 = inp . s i z e ( 1 )<br />
13 a s s e r t n1<br />
14<br />
15 c l i p = par . f l o a t ( ” c l i p ” )<br />
16 a s s e r t c l i p<br />
17<br />
18 t r a c e = numpy . z e r o s ( n1 , ’ f ’ )<br />
19<br />
PYTHON INTERFACE<br />
20 for i 2 in xrange ( n2 ) : # loop over t r a c e s
<strong>Madagascar</strong> Documentation RSF API 131<br />
21 inp . read ( t r a c e )<br />
22 t r a c e = numpy . c l i p ( trace ,− c l i p , c l i p )<br />
23 output . w r i t e ( t r a c e )<br />
Let us examine it in detail.<br />
3 import numpy<br />
4 import m8r<br />
The script starts with importing the numpy and rsf modules.<br />
6 par = m8r . Par ( )<br />
7 inp = m8r . Input ( )<br />
8 output = m8r . Output ( )<br />
9 a s s e r t ’ f l o a t ’ == inp . type<br />
Next, we initialize the command line interface and the standard input and output<br />
<strong>file</strong>s. We also make sure that the input <strong>file</strong> type is floating point.<br />
11 n1 = inp . i n t ( ”n1” )<br />
12 n2 = inp . s i z e ( 1 )<br />
13 a s s e r t n1<br />
We extract the “n1” parameter from the input <strong>file</strong>. Conceptually, the RSF data<br />
model is a multidimensional hypercube. The n1 parameter refers to the fastest axis.<br />
If the input dataset is a collection of traces, n1 corresponds to the trace length. We<br />
could proceed in a similar fashion, extracting n2, n3, etc. If we are interested in the<br />
total number of traces, like in the clip example, a shortcut is to use the size method<br />
of the Input class1. Calling size(0) returns the total number of elements in the<br />
hypercube (the product of n1, n2, etc.), calling size(1) returns the number of traces<br />
(the product of n2, n3, etc.), calling size(2) returns the product of n3, n4, etc.<br />
15 c l i p = par . f l o a t ( ” c l i p ” )<br />
16 a s s e r t c l i p<br />
The clip parameter is read from the command line, where it can be specified, for<br />
example, as clip=10.<br />
20 for i 2 in xrange ( n2 ) : # loop over t r a c e s<br />
21 inp . read ( t r a c e )<br />
22 t r a c e = numpy . c l i p ( trace ,− c l i p , c l i p )<br />
23 output . w r i t e ( t r a c e )<br />
Finally, we do the actual work: loop over input traces, reading, clipping, and writing<br />
out each trace.
132 Fomel <strong>Madagascar</strong> Documentation<br />
Compiling<br />
The python script does not require compilation. Simply make sure to set PYTHONPATH<br />
and LD LIBRARY PATH according to<br />
$RSFROOT/etc/madagascar/env.sh or $RSFROOT/etc/madagascar/env.csh.<br />
MATLAB INTERFACE<br />
The MATLAB clip function is listed below.<br />
1 function c l i p ( in , out , c l i p )<br />
2 %CLIP Clip the data<br />
3<br />
4 dims = r s f d i m ( in ) ;<br />
5 n1 = dims ( 1 ) ; % t r a c e l e n g t h<br />
6 n2 = prod( dims ( 2 : end ) ) ; % number of t r a c e s<br />
7 trace = 1 : n1 ; % a l l o c a t e t r a c e<br />
8 r s f c r e a t e ( out , in ) % c r e a t e an output f i l e<br />
9<br />
10 for i 2 = 1 : n2 % loop over t r a c e s<br />
11 r s f r e a d ( trace , in , ’ same ’ ) ;<br />
12 trace ( trace > c l i p ) = c l i p ;<br />
13 trace ( trace < − c l i p ) = −c l i p ;<br />
14 r s f w r i t e ( trace , out , ’ same ’ ) ;<br />
15 end<br />
Let us examine it in detail.<br />
4 dims = r s f d i m ( in ) ;<br />
We start by figuring out the input <strong>file</strong> dimensions.<br />
5 n1 = dims ( 1 ) ; % t r a c e l e n g t h<br />
6 n2 = prod( dims ( 2 : end ) ) ; % number of t r a c e s<br />
The first dimension is the trace length, the product of all other dimensions correspond<br />
to the number of traces.<br />
7 trace = 1 : n1 ; % a l l o c a t e t r a c e<br />
8 r s f c r e a t e ( out , in ) % c r e a t e an output f i l e<br />
Next, we allocate the trace array and create an output <strong>file</strong>.<br />
10 for i 2 = 1 : n2 % loop over t r a c e s<br />
11 r s f r e a d ( trace , in , ’ same ’ ) ;<br />
12 trace ( trace > c l i p ) = c l i p ;
<strong>Madagascar</strong> Documentation RSF API 133<br />
13 trace ( trace < − c l i p ) = −c l i p ;<br />
14 r s f w r i t e ( trace , out , ’ same ’ ) ;<br />
15 end<br />
Finally, we do the actual work: loop over input traces, reading, clipping, and writing<br />
out each trace.<br />
Compiling<br />
The MATLAB script does not require compilation. Simply make sure that $RSFROOT/lib<br />
is in MATLABPATH and LD LIBRARY PATH.<br />
INSTALLATION<br />
To install the interface to a particular language, use API= parameter in the RSF configuration.<br />
For example, to to install C++ and Fortran-90 API bindings in addition<br />
to the basic package, run<br />
scons API=c++,f90 config<br />
Only the C interface is configured by default. The configuration parameters are stored<br />
in<br />
$RSFROOT/share/madagascar/etc/config.py.<br />
REFERENCES<br />
Roberts, E. S., 1998, Programming abstractions in C: Addison-Wesley.
134 Fomel <strong>Madagascar</strong> Documentation
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Guide to programming using RSF<br />
Paul Sava 1<br />
ABSTRACT<br />
This guide demonstrates a simple time-domain finite-differences modeling code<br />
in RSF.<br />
INTRODUCTION<br />
This section presents time-domain finite-difference modeling 2 written with the RSF<br />
library. The program is demonstrated with the C, C++ and Fortran 90 interfaces.<br />
The acoustic wave-equation<br />
∆U − 1 ∂ 2 U<br />
= f(t) (1)<br />
v 2 ∂t2 can be written as<br />
[∆U − f(t)] v 2 = ∂2 U<br />
∂t . (2)<br />
2<br />
∆ is the Laplacian symbol, f(t) is the source wavelet, v is the velocity, and U is a<br />
scalar wavefield.<br />
A discrete time-step involves the following computations:<br />
U i+1 = [∆U − f(t)] v 2 ∆t 2 + 2U i − U i−1 , (3)<br />
where U i−1 , U i and U i+1 represent the propagating wavefield at various time steps.<br />
1 e-mail: paul.sava@beg.utexas.edu<br />
2 “Hello world” of seismic imaging.<br />
135
136 Sava <strong>Madagascar</strong> Documentation<br />
C PROGRAM<br />
1 /∗ time−domain a c o u s t i c FD m o d e l i n g ∗/<br />
2 #include < r s f . h><br />
3 int main ( int argc , char∗ argv [ ] )<br />
4 {<br />
5 /∗ L a p l a c i a n c o e f f i c i e n t s ∗/<br />
6 f l o a t c0 = −30./12. , c1 =+16./12. , c2=− 1 . / 1 2 . ;<br />
7<br />
8 b o o l verb ; /∗ v e r b o s e f l a g ∗/<br />
9 s f f i l e Fw=NULL, Fv=NULL, Fr=NULL, Fo=NULL; /∗ I /O f i l e s ∗/<br />
10 s f a x i s at , az , ax ; /∗ c u b e a x e s ∗/<br />
11 int i t , i z , i x ; /∗ i n d e x v a r i a b l e s ∗/<br />
12 int nt , nz , nx ;<br />
13 f l o a t dt , dz , dx , idx , idz , dt2 ;<br />
14<br />
15 f l o a t ∗ww, ∗ ∗ vv , ∗ ∗ r r ; /∗ I /O a r r a y s ∗/<br />
16 f l o a t ∗∗um, ∗ ∗ uo , ∗ ∗ up , ∗ ∗ ud ; /∗ tmp a r r a y s ∗/<br />
17<br />
18 s f i n i t ( argc , argv ) ;<br />
19 i f ( ! s f g e t b o o l ( ” verb ” ,& verb ) ) verb =0; /∗ v e r b o s e f l a g ∗/<br />
20<br />
21 /∗ s e t u p I /O f i l e s ∗/<br />
22 Fw = s f i n p u t ( ” i n ” ) ;<br />
23 Fo = s f o u t p u t ( ” out ” ) ;<br />
24 Fv = s f i n p u t ( ” v e l ” ) ;<br />
25 Fr = s f i n p u t ( ” r e f ” ) ;<br />
26<br />
27 /∗ Read / Write a x e s ∗/<br />
28 at = s f i a x a (Fw , 1 ) ; nt = s f n ( at ) ; dt = s f d ( at ) ;<br />
29 az = s f i a x a ( Fv , 1 ) ; nz = s f n ( az ) ; dz = s f d ( az ) ;<br />
30 ax = s f i a x a ( Fv , 2 ) ; nx = s f n ( ax ) ; dx = s f d ( ax ) ;<br />
31<br />
32 s f o a x a ( Fo , az , 1 ) ;<br />
33 s f o a x a ( Fo , ax , 2 ) ;<br />
34 s f o a x a ( Fo , at , 3 ) ;<br />
35<br />
36 dt2 = dt ∗ dt ;<br />
37 i d z = 1/( dz ∗ dz ) ;<br />
38 i d x = 1/( dx∗dx ) ;<br />
39<br />
40 /∗ r e a d w a v e l e t , v e l o c i t y & r e f l e c t i v i t y ∗/<br />
41 ww= s f f l o a t a l l o c ( nt ) ; s f f l o a t r e a d (ww , nt ,Fw ) ;<br />
42 vv= s f f l o a t a l l o c 2 ( nz , nx ) ; s f f l o a t r e a d ( vv [ 0 ] , nz ∗nx , Fv ) ;<br />
43 r r= s f f l o a t a l l o c 2 ( nz , nx ) ; s f f l o a t r e a d ( r r [ 0 ] , nz ∗nx , Fr ) ;<br />
44<br />
45 /∗ a l l o c a t e t e m p o r a r y a r r a y s ∗/<br />
46 um= s f f l o a t a l l o c 2 ( nz , nx ) ;<br />
47 uo= s f f l o a t a l l o c 2 ( nz , nx ) ;<br />
48 up= s f f l o a t a l l o c 2 ( nz , nx ) ;<br />
49 ud= s f f l o a t a l l o c 2 ( nz , nx ) ;<br />
50<br />
51 f o r ( i x =0; ix
<strong>Madagascar</strong> Documentation RSF DEMO 137<br />
90<br />
91 /∗ t i m e s t e p ∗/<br />
92 f o r ( i z =0; i z
138 Sava <strong>Madagascar</strong> Documentation<br />
• Compute Laplacian: ∆U.<br />
66 f o r ( i z =2; i z
<strong>Madagascar</strong> Documentation RSF DEMO 139<br />
C++ PROGRAM<br />
1 // time−domain a c o u s t i c FD m o d e l i n g<br />
2 #include <br />
3 #include <br />
4 #include < r s f . hh><br />
5 #include <br />
6 #include <br />
7 using namespace s t d ;<br />
8<br />
9 int main ( int argc , char∗ argv [ ] )<br />
10 {<br />
11 // L a p l a c i a n c o e f f i c i e n t s<br />
12 f l o a t c0 = −30./12. , c1 =+16./12. , c2=− 1 . / 1 2 . ;<br />
13<br />
14 s f i n i t ( argc , argv ) ; // i n i t RSF<br />
15 bool verb ; // v e b o s e f l a g<br />
16 i f ( ! s f g e t b o o l ( ” verb ” ,& verb ) ) verb =0;<br />
17<br />
18 // s e t u p I /O f i l e s<br />
19 CUB Fw( ” i n ” , ” i ” ) ; Fw . headin ( ) ; //Fw . r e p o r t ( ) ;<br />
20 CUB Fv ( ” v e l ” , ” i ” ) ; Fv . headin ( ) ; // Fv . r e p o r t ( ) ;<br />
21 CUB Fr ( ” r e f ” , ” i ” ) ; Fr . headin ( ) ; // Fr . r e p o r t ( ) ;<br />
22 CUB Fo ( ” out ” , ”o” ) ; Fo . s e t u p ( 3 , Fv . e s i z e ( ) ) ;<br />
23<br />
24 // Read / Write a x e s<br />
25 s f a x i s at = Fw . g e t a x ( 0 ) ; int nt = s f n ( at ) ; f l o a t dt = s f d ( at ) ;<br />
26 s f a x i s az = Fv . g e t a x ( 0 ) ; int nz = s f n ( az ) ; f l o a t dz = s f d ( az ) ;<br />
27 s f a x i s ax = Fv . g e t a x ( 1 ) ; int nx = s f n ( ax ) ; f l o a t dx = s f d ( ax ) ;<br />
28<br />
29 Fo . putax ( 0 , az ) ;<br />
30 Fo . putax ( 1 , ax ) ;<br />
31 Fo . putax ( 2 , at ) ;<br />
32 Fo . headou ( ) ;<br />
33<br />
34 f l o a t dt2 = dt ∗ dt ;<br />
35 f l o a t i d z = 1/( dz ∗ dz ) ;<br />
36 f l o a t i d x = 1/( dx∗dx ) ;<br />
37<br />
38 // r e a d w a v e l e t , v e l o c i t y and r e f l e c t i v i t y<br />
39 v a l a r r a y ww( nt ) ; ww=0; Fw >> ww;<br />
40 v a l a r r a y vv ( nz ∗nx ) ; vv =0; Fv >> vv ;<br />
41 v a l a r r a y r r ( nz ∗nx ) ; r r =0; Fr >> r r ;<br />
42<br />
43 // a l l o c a t e t e m p o r a r y a r r a y s<br />
44 v a l a r r a y um( nz ∗nx ) ; um=0;<br />
45 v a l a r r a y uo ( nz ∗nx ) ; uo =0;<br />
46 v a l a r r a y up ( nz ∗nx ) ; up=0;<br />
47 v a l a r r a y ud ( nz ∗nx ) ; ud=0;<br />
48<br />
49 // i n i t ValArray I n d e x c o u n t e r<br />
50 VAI k ( nz , nx ) ;<br />
51<br />
52 // MAIN LOOP<br />
53 i f ( verb ) c e r r
140 Sava <strong>Madagascar</strong> Documentation<br />
1. Declare input, output and auxiliary <strong>file</strong> cubes (of type CUB).<br />
19 CUB Fw( ” i n ” , ” i ” ) ; Fw . headin ( ) ; //Fw . r e p o r t ( ) ;<br />
20 CUB Fv ( ” v e l ” , ” i ” ) ; Fv . headin ( ) ; // Fv . r e p o r t ( ) ;<br />
21 CUB Fr ( ” r e f ” , ” i ” ) ; Fr . headin ( ) ; // Fr . r e p o r t ( ) ;<br />
22 CUB Fo ( ” out ” , ”o” ) ; Fo . s e t u p ( 3 , Fv . e s i z e ( ) ) ;<br />
2. Declare, read and write RSF cube axes: at time axis, ax space axis, az depth<br />
axis.<br />
25 s f a x i s at = Fw . g e t a x ( 0 ) ; int nt = s f n ( at ) ; f l o a t dt = s f d ( at ) ;<br />
26 s f a x i s az = Fv . g e t a x ( 0 ) ; int nz = s f n ( az ) ; f l o a t dz = s f d ( az ) ;<br />
27 s f a x i s ax = Fv . g e t a x ( 1 ) ; int nx = s f n ( ax ) ; f l o a t dx = s f d ( ax ) ;<br />
28<br />
29 Fo . putax ( 0 , az ) ;<br />
30 Fo . putax ( 1 , ax ) ;<br />
31 Fo . putax ( 2 , at ) ;<br />
32 Fo . headou ( ) ;<br />
3. Declare multi-dimensional valarrays for input, output and read data.<br />
39 v a l a r r a y ww( nt ) ; ww=0; Fw >> ww;<br />
40 v a l a r r a y vv ( nz ∗nx ) ; vv =0; Fv >> vv ;<br />
41 v a l a r r a y r r ( nz ∗nx ) ; r r =0; Fr >> r r ;<br />
4. Declare multi-dimensional valarrays for temporary storage.<br />
44 v a l a r r a y um( nz ∗nx ) ; um=0;<br />
45 v a l a r r a y uo ( nz ∗nx ) ; uo =0;<br />
46 v a l a r r a y up ( nz ∗nx ) ; up=0;<br />
47 v a l a r r a y ud ( nz ∗nx ) ; ud=0;<br />
5. Initialize multidimensional valarray index counter (of type VAI).<br />
50 VAI k ( nz , nx ) ;<br />
6. Loop over time.<br />
54 f o r ( int i t =0; i t
<strong>Madagascar</strong> Documentation RSF DEMO 141<br />
FORTRAN 90 PROGRAM<br />
1 ! time−domain a c o u s t i c FD m o d e l i n g<br />
2 program AFDMf90<br />
3 use r s f<br />
4<br />
5 i m p l i c i t none<br />
6<br />
7 ! L a p l a c i a n c o e f f i c i e n t s<br />
8 r e a l : : c0 = −30./12. , c1 =+16./12. , c2=− 1 . / 1 2 .<br />
9<br />
10 l o g i c a l : : verb ! v e r b o s e f l a g<br />
11 type ( f i l e ) : : Fw, Fv , Fr , Fo ! I /O f i l e s<br />
12 type ( axa ) : : at , az , ax ! c u b e a x e s<br />
13 integer : : i t , i z , i x ! i n d e x v a r i a b l e s<br />
14 r e a l : : idx , idz , dt2<br />
15<br />
16 real , a l l o c a t a b l e : : vv ( : , : ) , r r ( : , : ) , ww ( : ) ! I /O a r r a y s<br />
17 real , a l l o c a t a b l e : : um ( : , : ) , uo ( : , : ) , up ( : , : ) , ud ( : , : ) ! tmp a r r a y s<br />
18<br />
19 c a l l s f i n i t ( ) ! i n i t RSF<br />
20 c a l l f r o m p a r ( ” verb ” , verb , . f a l s e . )<br />
21<br />
22 ! s e t u p I /O f i l e s<br />
23 Fw=r s f i n p u t ( ” i n ” )<br />
24 Fv=r s f i n p u t ( ” v e l ” )<br />
25 Fr=r s f i n p u t ( ” r e f ” )<br />
26 Fo=r s f o u t p u t ( ” out ” )<br />
27<br />
28 ! Read / Write a x e s<br />
29 c a l l i a x a (Fw, at , 1 ) ; c a l l i a x a ( Fv , az , 1 ) ; c a l l i a x a ( Fv , ax , 2 )<br />
30 c a l l oaxa ( Fo , az , 1 ) ; c a l l oaxa ( Fo , ax , 2 ) ; c a l l oaxa ( Fo , at , 3 )<br />
31<br />
32 dt2 = at%d∗ at%d<br />
33 i d z = 1/( az%d∗ az%d )<br />
34 i d x = 1/( ax%d∗ax%d )<br />
35<br />
36 ! r e a d w a v e l e t , v e l o c i t y & r e f l e c t i v i t y<br />
37 a l l o c a t e (ww( at%n ) ) ; ww= 0 . ; c a l l r s f r e a d (Fw,ww)<br />
38 a l l o c a t e ( vv ( az%n , ax%n ) ) ; vv = 0 . ; c a l l r s f r e a d ( Fv , vv )<br />
39 a l l o c a t e ( r r ( az%n , ax%n ) ) ; r r = 0 . ; c a l l r s f r e a d ( Fr , r r )<br />
40<br />
41 ! a l l o c a t e t e m p o r a r y a r r a y s<br />
42 a l l o c a t e (um( az%n , ax%n ) ) ; um=0.<br />
43 a l l o c a t e ( uo ( az%n , ax%n ) ) ; uo =0.<br />
44 a l l o c a t e ( up ( az%n , ax%n ) ) ; up=0.<br />
45 a l l o c a t e ( ud ( az%n , ax%n ) ) ; ud=0.<br />
46<br />
47 ! MAIN LOOP<br />
48 do i t =1, at%n<br />
49 i f ( verb ) write ( 0 , ∗ ) i t<br />
50<br />
51 ! 4 t h o r d e r l a p l a c i a n<br />
52 do i z =2, az%n−2<br />
53 do i x =2,ax%n−2<br />
54 ud ( i z , i x ) = &<br />
55 c0 ∗ uo ( i z , i x ) ∗ ( i d x + i d z ) + &<br />
56 c1 ∗( uo ( i z , ix −1) + uo ( i z , i x +1))∗ i d x + &<br />
57 c2 ∗( uo ( i z , ix −2) + uo ( i z , i x +2))∗ i d x + &<br />
58 c1 ∗( uo ( i z −1, i x ) + uo ( i z +1, i x ) ) ∗ i d z + &<br />
59 c2 ∗( uo ( i z −2, i x ) + uo ( i z +2, i x ) ) ∗ i d z<br />
60 end do<br />
61 end do<br />
62<br />
63 ! i n j e c t w a v e l e t<br />
64 ud = ud − ww( i t ) ∗ r r<br />
65<br />
66 ! s c a l e by v e l o c i t y<br />
67 ud= ud ∗vv∗vv<br />
68<br />
69 ! t i m e s t e p<br />
70 up = 2∗ uo − um + ud ∗ dt2<br />
71 um = uo<br />
72 uo = up<br />
73<br />
74 ! w r i t e w a v e f i e l d t o o u t p u t<br />
75 c a l l r s f w r i t e ( Fo , uo )<br />
76 end do<br />
77<br />
78 c a l l e x i t ( 0 )<br />
79 end program AFDMf90
142 Sava <strong>Madagascar</strong> Documentation<br />
• Declare input, output and auxiliary <strong>file</strong> tags.<br />
11 type ( f i l e ) : : Fw, Fv , Fr , Fo ! I /O f i l e s<br />
• Declare RSF cube axes: at time axis, ax space axis, az depth axis.<br />
12 type ( axa ) : : at , az , ax ! c u b e a x e s<br />
• Declare multi-dimensional arrays for input, output and computations.<br />
16 real , a l l o c a t a b l e : : vv ( : , : ) , r r ( : , : ) , ww ( : ) ! I /O a r r a y s<br />
17 real , a l l o c a t a b l e : : um ( : , : ) , uo ( : , : ) , up ( : , : ) , ud ( : , : ) ! tmp a r r a y s<br />
• Open <strong>file</strong>s for input/output.<br />
23 Fw=r s f i n p u t ( ” i n ” )<br />
24 Fv=r s f i n p u t ( ” v e l ” )<br />
25 Fr=r s f i n p u t ( ” r e f ” )<br />
26 Fo=r s f o u t p u t ( ” out ” )<br />
• Read axes from input <strong>file</strong>s; write axes to output <strong>file</strong>.<br />
29 c a l l i a x a (Fw, at , 1 ) ; c a l l i a x a ( Fv , az , 1 ) ; c a l l i a x a ( Fv , ax , 2 )<br />
30 c a l l oaxa ( Fo , az , 1 ) ; c a l l oaxa ( Fo , ax , 2 ) ; c a l l oaxa ( Fo , at , 3 )<br />
• Allocate arrays and read wavelet, velocity and reflectivity.<br />
37 a l l o c a t e (ww( at%n ) ) ; ww= 0 . ; c a l l r s f r e a d (Fw,ww)<br />
38 a l l o c a t e ( vv ( az%n , ax%n ) ) ; vv = 0 . ; c a l l r s f r e a d ( Fv , vv )<br />
39 a l l o c a t e ( r r ( az%n , ax%n ) ) ; r r = 0 . ; c a l l r s f r e a d ( Fr , r r )<br />
• Allocate temporary arrays.<br />
42 a l l o c a t e (um( az%n , ax%n ) ) ; um=0.<br />
43 a l l o c a t e ( uo ( az%n , ax%n ) ) ; uo =0.<br />
44 a l l o c a t e ( up ( az%n , ax%n ) ) ; up=0.<br />
45 a l l o c a t e ( ud ( az%n , ax%n ) ) ; ud=0.<br />
• Loop over time.<br />
48 do i t =1, at%n<br />
• Compute Laplacian: ∆U.<br />
52 do i z =2, az%n−2<br />
53 do i x =2,ax%n−2<br />
54 ud ( i z , i x ) = &<br />
55 c0 ∗ uo ( i z , i x ) ∗ ( i d x + i d z ) + &<br />
56 c1 ∗( uo ( i z , ix −1) + uo ( i z , i x +1))∗ i d x + &<br />
57 c2 ∗( uo ( i z , ix −2) + uo ( i z , i x +2))∗ i d x + &<br />
58 c1 ∗( uo ( i z −1, i x ) + uo ( i z +1, i x ) ) ∗ i d z + &<br />
59 c2 ∗( uo ( i z −2, i x ) + uo ( i z +2, i x ) ) ∗ i d z<br />
60 end do<br />
61 end do<br />
• Inject source wavelet: [∆U − f(t)]<br />
64 ud = ud − ww( i t ) ∗ r r<br />
• Scale by velocity: [∆U − f(t)] v 2<br />
67 ud= ud ∗vv∗vv<br />
• Time step: U i+1 = [∆U − f(t)] v 2 ∆t 2 + 2U i − U i−1<br />
70 up = 2∗ uo − um + ud ∗ dt2<br />
71 um = uo<br />
72 uo = up
<strong>Madagascar</strong> Documentation, RSF, July 19, 2012<br />
Reproducible computational experiments<br />
using SCons<br />
Sergey Fomel 1 and Gilles Hennenfent 21<br />
ABSTRACT<br />
SCons (from Software Construction) is a well-known open-source program designed<br />
primarily for building software. In this paper, we describe our method of<br />
extending SCons for managing data processing flows and reproducible computational<br />
experiments. We demonstrate our usage of SCons with a couple of simple<br />
examples.<br />
INTRODUCTION<br />
This paper introduces an environment for reproducible computational experiments<br />
developed as part of the “<strong>Madagascar</strong>” software package. To reproduce the example<br />
experiments in this paper, you can download <strong>Madagascar</strong> from http://www.ahay.<br />
org/. At the moment, the main <strong>Madagascar</strong> interface is the Unix shell command<br />
line so that you will need a Unix/POSIX system (Linux, Mac OS X, Solaris, etc.) or<br />
Unix emulation under Windows (Cygwin, SFU, etc.)<br />
Our focus, however, is not only on particular tools we use in our research but also<br />
on the general philosophy of reproducible computations.<br />
Reproducible research philosophy<br />
Peer review is the backbone of scientific progress. From the ancient alchemists, who<br />
worked in secret on magic solutions to insolvable problems, the modern science has<br />
come a long way to become a social enterprise, where hypotheses, theories, and experimental<br />
results are openly published and verified by the community. By reproducing<br />
and verifying previously published research, a researcher can take new steps to advance<br />
the progress of science.<br />
Traditionally, scientific disciplines are divided into theoretical and experimental<br />
studies. Reproduction and verification of theoretical results usually requires only<br />
1 University of Texas at Austin, E-mail: sergey.fomel@beg.utexas.edu<br />
2 Earth & Ocean Sciences, University of British Columbia, E-mail: ghennenfent@eos.ubc.ca<br />
1 e-mail: paul.sava@beg.utexas.edu<br />
143
144 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
imagination (apart from pencils and paper), experimental results are verified in laboratories<br />
using equipment and materials similar to those described in the publication.<br />
During the last century, computational studies emerged as a new scientific discipline.<br />
Computational experiments are carried out on a computer by applying numerical<br />
algorithms to digital data. How reproducible are such experiments? On one hand,<br />
reproducing the result of a numerical experiment is a difficult undertaking. The reader<br />
needs to have access to precisely the same kind of input data, software and hardware<br />
as the author of the publication in order to reproduce the published result. It is often<br />
difficult or impossible to provide detailed specifications for these components. On the<br />
other hand, basic computational system components such as operating systems and<br />
<strong>file</strong> formats are getting increasingly standardized, and new components can be shared<br />
in principle because they simply represent digital information transferable over the<br />
Internet.<br />
The practice of software sharing has fueled the miraculously efficient development<br />
of Linux, Apache, and many other open-source software projects. Its proponents<br />
often refer to this ideology as an analog of the scientific peer review tradition. Eric<br />
Raymond, a well-known open-source advocate, writes (Raymond, 2004):<br />
Abandoning the habit of secrecy in favor of process transparency and peer<br />
review was the crucial step by which alchemy became chemistry. In the<br />
same way, it is beginning to appear that open-source development may<br />
signal the long-awaited maturation of software development as a discipline.<br />
While software development is trying to imitate science, computational science needs<br />
to borrow from the open-source model in order to sustain itself as a fully scientific<br />
discipline. In words of Randy LeVeque, a prominent mathematician (LeVeque, 2006),<br />
Within the world of science, computation is now rightly seen as a third<br />
vertex of a triangle complementing experiment and theory. However, as<br />
it is now often practiced, one can make a good case that computing is<br />
the last refuge of the scientific scoundrel [...] Where else in science can<br />
one get away with publishing observations that are claimed to prove a<br />
theory or illustrate the success of a technique without having to give a<br />
careful description of the methods used, in sufficient detail that others<br />
can attempt to repeat the experiment? [...] Scientific and mathematical<br />
journals are filled with pretty pictures these days of computational experiments<br />
that the reader has no hope of repeating. Even brilliant and<br />
well intentioned computational scientists often do a poor job of presenting<br />
their work in a reproducible manner. The methods are often very vaguely<br />
defined, and even if they are carefully defined, they would normally have<br />
to be implemented from scratch by the reader in order to test them.<br />
In computer science, the concept of publishing and explaining computer programs<br />
goes back to the idea of literate programming promoted by Knuth (1984) and ex-
<strong>Madagascar</strong> Documentation Reproducible research 145<br />
pended by many other researchers (Thimbleby, 2003). In his 2004 lecture on “better<br />
programming”, Harold Thimbleby notes 2<br />
We want ideas, and in particular programs, that work in one place to work<br />
elsewhere. One form of objectivity is that published science must work<br />
elsewhere than just in the author’s laboratory or even just in the author’s<br />
imagination; this requirement is called reproducibility.<br />
Nearly ten years ago, the technology of reproducible research in geophysics was<br />
pioneered by Jon Claerbout and his students at the Stanford Exploration Project<br />
(SEP). SEP’s system of reproducible research requires the author of a publication to<br />
document creation of numerical results from the input data and software sources to<br />
let others test and verify the result reproducibility (Claerbout, 1992a; Schwab et al.,<br />
2000). The discipline of reproducible research was also adopted and popularized in<br />
the statistics and wavelet theory community by Buckheit and Donoho (1995). It<br />
is referenced in several popular wavelet theory books (Hubbard, 1998; Mallat, 1999).<br />
Pledges for reproducible research appear nowadays in fields as diverse as bioinformatics<br />
(Gentleman et al., 2004), geoinformatics (Bivand, 2006), and computational wave<br />
propagation (LeVeque, 2006). However, the adoption or reproducible research practice<br />
by computational scientists has been slow. Partially, this is caused by difficult<br />
and inadequate tools.<br />
Tools for reproducible research<br />
The reproducible research system developed at Stanford is based on “make (Stallman<br />
et al., 2004)”, a Unix software construction utility. Originally, SEP used “cake”, a<br />
dialect of “make” (Nichols and Cole, 1989; Claerbout and Nichols, 1990; Claerbout,<br />
1992b; Claerbout and Karrenbach, 1993). The system was converted to “GNU make”,<br />
a more standard dialect, by Schwab and Schroeder (1995). The “make” program<br />
keeps track of dependencies between different components of the system and the<br />
software construction targets, which, in the case of a reproducible research system,<br />
turn into figures and manuscripts. The targets and commands for their construction<br />
are specified by the author in “make<strong>file</strong>s”, which serve as databases for defining source<br />
and target dependencies. A dependency-based system leads to rapid development,<br />
because when one of the sources changes, only parts that depend on this source get<br />
recomputed. Buckheit and Donoho (1995) based their system on MATLAB, a popular<br />
integrated development environment produced by MathWorks (Sigmon and Davis,<br />
2001). While MATLAB is an adequate tool for prototyping numerical algorithms, it<br />
may not be sufficient for large-scale computations typical for many applications in<br />
computational geophysics.<br />
“Make” is an extremely useful utility employed by thousands of software development<br />
projects. Unfortunately, it is not well designed from the user experience<br />
2 http://www.uclic.ucl.ac.uk/harold/
146 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
prospective. “Make” employs an obscure and limited special language (a mixture of<br />
Unix shell commands and special-purpose commands), which often appears confusing<br />
to unexperienced users. According to Peter van der Linden, a software expert from<br />
Sun Microsystems (van der Linden, 1994),<br />
“Sendmail” and “make” are two well known programs that are pretty<br />
widely regarded as originally being debugged into existence. That’s why<br />
their command languages are so poorly thought out and difficult to learn.<br />
It’s not just you – everyone finds them troublesome.<br />
The inconvenience of “make” command language is also in its limited capabilities.<br />
The reproducible research system developed by Schwab et al. (2000) includes not<br />
only custom “make” rules but also an obscure and hardly portable agglomeration of<br />
shell and Perl scripts that extend “make” (Fomel et al., 1997).<br />
Several alternative systems for dependency-checking software construction have<br />
been developed in recent years. One of the most promising new tools is SCons, enthusiastically<br />
endorsed by Dubois (2003). The SCons initial design won the Software<br />
Carpentry competition sponsored by Los Alamos National Laboratory in 2000 in the<br />
category of “a dependency management tool to replace make”. Some of the main<br />
advantages of SCons are:<br />
• SCons configuration <strong>file</strong>s are Python scripts. Python is a modern programming<br />
language praised for its readability, elegance, simplicity, and power (Rossum,<br />
2000a,b). Scales and Ecke (2002) recommend Python as the first programming<br />
language for geophysics students.<br />
• SCons offers reliable, automatic, and extensible dependency analysis and creates<br />
a global view of all dependencies – no more “make depend”, “make clean”,<br />
or multiple build passes of touching and reordering targets to get all of the<br />
dependencies.<br />
• SCons has built-in support for many programming languages and systems: C,<br />
C++, Fortran, Java, LaTeX, and others.<br />
• While “make” relies on timestamps for detecting <strong>file</strong> changes (creating numerous<br />
problems on platforms with different system clocks), SCons uses by default a<br />
more reliable detection mechanism employing MD5 signatures. It can detect<br />
changes not only in <strong>file</strong>s but also in commands used to build them.<br />
• SCons provides integrated support for parallel builds.<br />
• SCons provides configuration support analogous to the “autoconf” utility for<br />
testing the environment on different platforms.<br />
• SCons is designed from the ground up as a cross-platform tool. It is known to<br />
work equally well on both POSIX systems (Linux, Mac OS X, Solaris, etc.) and<br />
Windows.
<strong>Madagascar</strong> Documentation Reproducible research 147<br />
• The stability of SCons is assured by an incremental development methodology<br />
utilizing comprehensive regression tests.<br />
• SCons is publicly released under a liberal open-source license 3<br />
In this paper, we propose to adopt SCons as a new platform for reproducible<br />
research in scientific computing.<br />
Paper organization<br />
We first give a brief overview of “<strong>Madagascar</strong>” software package and define the different<br />
levels of user interactions. To demonstrate our adoption of SCons for reproducible<br />
research, we then describe a couple of simple examples of computational experiments<br />
and finally show how SCons helps us document our computational results.<br />
MADAGASCAR SOFTWARE PACKAGE OVERVIEW<br />
Report/paper<br />
(SCons + LaTeX)<br />
Book<br />
(SCons + LaTeX)<br />
Report/paper<br />
(SCons + LaTeX)<br />
Documention<br />
(<strong>PDF</strong> & HTML)<br />
Processing flow<br />
(SCons + Python)<br />
Processing flow<br />
(SCons + Python)<br />
Processing flow<br />
(SCons + Python)<br />
Processing flows<br />
Program<br />
(Matlab)<br />
Program<br />
(Mathematica)<br />
Program<br />
(Python)<br />
Program<br />
(C++)<br />
Program<br />
(Fortran)<br />
Program<br />
(C)<br />
Program<br />
(SU)<br />
Program<br />
(SEP)<br />
Program<br />
(Delphi)<br />
Command line<br />
Figure 1: caption scons/. diag<br />
“<strong>Madagascar</strong>” is a multi-layered software package (Fig. 1). Users can thus use it<br />
in different ways:<br />
3 As of time of this writing, SCons is in a beta version 0.96 approaching the 1.0 official release.<br />
See http://www.scons.org/.
148 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
• command line: “<strong>Madagascar</strong>” is first of all a collection of command line<br />
programs. Most programs act as filters on input data and can be chained in a<br />
Unix pipeline, e.g.<br />
sfspike n1=200 n2=50 | sfnoise rep=y >noise.rsf<br />
Although these programs mainly focus at this point on geophysical applications,<br />
users can use the API (application programmer’s interface) for writing their own<br />
software to manipulate Regularly Sampled Format (RSF) <strong>file</strong>s, “<strong>Madagascar</strong>”<br />
<strong>file</strong> format. The main software language of “<strong>Madagascar</strong>” is C. Interfaces to<br />
other languages (C++, Fortran-77, Fortran-90, Python) are also provided.<br />
• processing flows: “<strong>Madagascar</strong>” is also an environment for reproducible numerical<br />
experiments in a very broad sense. These numerical experiments (or<br />
“computational recipes”) can be done not only using “<strong>Madagascar</strong>” command<br />
line programs but also Matlab, Mathematica, Python, or other seismic packages<br />
(e.g. SEP, Seismic Unix). We adopted SCons for this part as we shall<br />
demonstrate later.<br />
• documentation: the most upper layer of “<strong>Madagascar</strong>” and maybe the most<br />
critical for reproducible research is documentation. “<strong>Madagascar</strong>” establishes<br />
a direct link between the figures of a paper or a report and the codes that<br />
were used to generate them. This layer uses SCons in combination with L A TEX<br />
to generate <strong>PDF</strong>, HTML, and MediaWiki <strong>file</strong>s real easy and undoubtly makes<br />
“<strong>Madagascar</strong>” an environment of choice for technology transfer, report, thesis,<br />
and peer-reviewed publication writing.<br />
EXAMPLE EXPERIMENTS<br />
The main SConstruct commands defined in our reproducible research environment<br />
are collected in Table 1.<br />
These commands are defined in $RSFROOT/lib/rsfproj.py where RSFROOT is the<br />
environmental variable to the <strong>Madagascar</strong> installation directory. The source of this<br />
<strong>file</strong> is in python/rsfproj.py.<br />
Example 1<br />
To follow the first example, select a working project directory and copy the following<br />
code to a <strong>file</strong> named SConstruct 4 .<br />
1 from r s f . p r o j import ∗<br />
2<br />
4 The source of this <strong>file</strong> is also accessible at book/rsf/scons/easystart/SConstruct.
<strong>Madagascar</strong> Documentation Reproducible research 149<br />
Fetch(data <strong>file</strong>,dir[,ftp server info])<br />
A rule to download from a specific directory of an<br />
FTP server .<br />
Flow(target[s],source[s],command[s][,stdin][,stdout])<br />
A rule to generate from using command[s]<br />
Plot(intermediate plot[,source],plot command) or<br />
Plot(intermediate plot,intermediate plots,combination)<br />
A rule to generate in the working directory.<br />
Result(plot[,source],plot command) or<br />
Result(plot,intermediate plots,combination)<br />
A rule to generate a final in the special Fig folder of the working<br />
directory.<br />
End()<br />
A rule to collect default targets.<br />
Table 1: Basic methods of an rsf.proj object.<br />
3 # Download the input data f i l e<br />
4 Fetch ( ’ lena . img ’ , ’ imgs ’ )<br />
5<br />
6 # Create RSF header<br />
7 Flow ( ’ lena . hdr ’ , ’ lena . img ’ ,<br />
8 ’ echo n1=512 n2=513 in=$SOURCE data format=n a t i v e u c h a r ’ ,<br />
9 s t d i n =0)<br />
10<br />
11 # Convert to f l o a t i n g p o i n t and window out f i r s t t r a c e<br />
12 Flow ( ’ lena ’ , ’ lena . hdr ’ , ’ dd type=f l o a t | window f 2=1 ’ )<br />
13<br />
14 # Display<br />
15 Result ( ’ lena ’ ,<br />
16 ’ ’ ’<br />
17 s f g r e y t i t l e =”Hello , World ! ” transp=n c o l o r=b b i a s =128<br />
18 c l i p =100 s c r e e n r a t i o =1<br />
19 ’ ’ ’ )<br />
20<br />
21 # Wrap up<br />
22 End ( )
150 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
This is our “hello world” example that illustrates the basic use of some of the<br />
commands presented in Table 1. The plan for this experiment is simply to download<br />
data from a public data server, to convert it to an appropriate <strong>file</strong> format and to<br />
generate a figure for publication. But let us have a closer look at the SConstruct<br />
script and try to decorticate it.<br />
1 from r s f . p r o j import ∗<br />
is a standard Python command that loads the <strong>Madagascar</strong> project management module<br />
rsf.proj.py which provides our extension to SCons.<br />
4 Fetch ( ’ lena . img ’ , ’ imgs ’ )<br />
instructs SCons to connect to a public data server (the default server if no FTP server<br />
information is provided) and to fetch the data <strong>file</strong> lena.img from the data/imgs<br />
directory. Try running “scons lena.img” on the command line. The successful<br />
output should look like<br />
bash$ scons lena.img<br />
scons: Reading SConscript <strong>file</strong>s ...<br />
scons: done reading SConscript <strong>file</strong>s.<br />
scons: Building targets ...<br />
retrieve(["lena.img"], [])<br />
scons: done building targets.<br />
with the target <strong>file</strong> lena.img appearing in your directory. In the following examples,<br />
we will use -Q (quiet) option of scons to suppress the verbose output.<br />
7 Flow ( ’ lena . hdr ’ , ’ lena . img ’ ,<br />
8 ’ echo n1=512 n2=513 in=$SOURCE data format=n a t i v e u c h a r ’ ,<br />
9 s t d i n =0)<br />
prepares the <strong>Madagascar</strong> header <strong>file</strong> lena.hdr using the standard Unix command<br />
echo.<br />
bash$ scons -Q lena.hdr<br />
echo n1=512 n2=513 in=lena.img data_format=native_uchar > lena.hdr<br />
Since echo does not take a standard input, stdin is set to 0 in the Flow command<br />
otherwise the first source is the standard input. Likewise, the first target is the<br />
standard output unless otherwise specified. Note that lena.img is referred as $SOURCE<br />
in the command. This allows us to change the name of the source <strong>file</strong> without changing<br />
the command.
<strong>Madagascar</strong> Documentation Reproducible research 151<br />
The data format of the lena.img image <strong>file</strong> is uchar (unsigned character), the<br />
image consists of 513 traces with 512 samples per trace. Our next step is to convert the<br />
image representation to floating point numbers and to window out the first trace so<br />
that the final image is a 512 by 512 square. The two transformations are conveniently<br />
combined into one with the help of a Unix pipe.<br />
12 Flow ( ’ lena ’ , ’ lena . hdr ’ , ’ dd type=f l o a t | window f 2=1 ’ )<br />
bash$ scons -Q lena<br />
scons: *** Do not know how to make target ‘lena’.<br />
Stop.<br />
What happened? In the absence of the <strong>file</strong> suffix, the Flow command assumes that<br />
the target <strong>file</strong> suffix is “.rsf”. Let us try again.<br />
scons -Q lena.rsf<br />
< lena.hdr /RSF/bin/sfdd type=float | /RSF/bin/sfwindow f2=1 > lena.rsf<br />
Notice that <strong>Madagascar</strong> modules sfdd and sfwindow get substituted for the corresponding<br />
short names in the SConstruct <strong>file</strong>. The <strong>file</strong> lena.rsf is in a regularly<br />
sampled format 5 and can be examined, for example, with sfin lena.rsf 6 .<br />
bash$ sfin lena.rsf<br />
lena.rsf:<br />
in="/datapath/lena.rsf@"<br />
esize=4 type=float form=native<br />
n1=512 d1=1 o1=0<br />
n2=512 d2=1 o2=1<br />
262144 elements 1048576 bytes<br />
In the last step, we will create a plot <strong>file</strong> for displaying the image on the screen and<br />
for including it in the publication.<br />
15 Result ( ’ lena ’ ,<br />
16 ’ ’ ’<br />
17 s f g r e y t i t l e =”Hello , World ! ” transp=n c o l o r=b b i a s =128<br />
18 c l i p =100 s c r e e n r a t i o =1<br />
19 ’ ’ ’ )<br />
Notice that we broke the long command string into multiple lines by using Python’s<br />
triple quote syntax. All the extra white space will be ignored when the multiple line<br />
string gets translated into the command line. The Result command has special targets<br />
associated with it. Try, for example, “scons lena.view” to observe the figure<br />
Fig/lena.vpl generated in a specially created Fig directory and displayed on the
152 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
Figure 2: The output of the<br />
first numerical experiment.<br />
scons/easystart lena<br />
screen. The output should look like Figure 2.<br />
The reproducible script ends with<br />
22 End ( )<br />
Ready to experiment? Try some of the following:<br />
1. Run scons -c. The -c (clean) option tells SCons to remove all default targets<br />
(the Fig/lena.vpl image <strong>file</strong> in our case) and also all intermediate targets that<br />
it generated.<br />
bash$ scons -c -Q<br />
Removed lena.img<br />
Removed lena.hdr<br />
Removed lena.rsf<br />
Removed /datapath/lena.rsf@<br />
Removed Fig/lena.vpl<br />
Run scons again, and the default target will be regenerated.<br />
bash$ scons -Q<br />
retrieve(["lena.img"], [])<br />
echo n1=512 n2=513 in=lena.img data_format=native_uchar > lena.hdr<br />
< lena.hdr /RSF/bin/sfdd type=float | /RSF/bin/sfwindow f2=1 > lena.rsf<br />
< lena.rsf /RSF/bin/sfgrey title="Hello, World!" transp=n color=b \<br />
bias=128 clip=100 screenratio=1 > Fig/lena.vpl<br />
2. Edit your SConstruct <strong>file</strong> and change some of the plotting parameters. For<br />
example, change the value of clip from clip=100 to clip=50. Run scons<br />
again and observe that only the last part of the processing flow (precisely, the<br />
part affected by the parameter change) is being run:<br />
5 See http://rsf.sourceforge.net/wiki/index.php/Format<br />
6 See http://rsf.sourceforge.net/wiki/index.php/Programs#sfin.
<strong>Madagascar</strong> Documentation Reproducible research 153<br />
bash$ scons -Q view<br />
< lena.rsf /RSF/bin/sfgrey title="Hello, World!" transp=n color=b \<br />
bias=128 clip=50 screenratio=1 > Fig/lena.vpl<br />
/RSF/bin/xtpen Fig/lena.vpl<br />
SCons is smart enough to recognize that your editing did not affect any of the<br />
previous results in the data flow chain! Keeping track of dependencies is the<br />
main feature that separates data processing and computational experimenting<br />
with SCons from using linear shell scripts. For computationally demanding<br />
data processing, this feature can save you a lot of time and can make your<br />
experiments more interactive and enjoyable.<br />
3. A special parameter to SCons (defined in rsf.proj.py) can time the execution<br />
of each step in the processing flow. Try running scons TIMER=y.<br />
4. The rsf.proj module has direct access to the database that stores parameters<br />
of all <strong>Madagascar</strong> modules. Try running scons CHECKPAR=y to see parameter<br />
checking enforced before computations 7 .<br />
The summary of our SCons commands is given in Table 2.<br />
Example 2<br />
The plan for this experiment is to add random noise to the test “Lena” image and then<br />
to attempt removing it by low-pass filtering and by hard thresholding of coefficients<br />
in the Fourier domain. The result images are shown in Figure 3 and 4.<br />
Since the SConstruct| <strong>file</strong> is a Python script, we can also use all the flexibility and<br />
power of the Python language in our <strong>Madagascar</strong> reproducible scripts. A demo script<br />
is available in the rsf/scons/rsfpy subdirectory of the <strong>Madagascar</strong> book directory.<br />
Rather than commenting it line-by-line, we select some parts of interest.<br />
In the SConstruct script, we can declare Python variables<br />
3 b i a s = 128<br />
and use them later, for example, to define our customized plot command as a Python<br />
function<br />
5 def grey ( t i t l e , transp=’n ’ , b i a s=b i a s ) :<br />
6 return ’ ’ ’<br />
7 s f g r e y t i t l e=”%s ” transp=%s b i a s=%g c l i p =100<br />
8 s c r e e n h t =10 screenwd=10 crowd2 =0.85 crowd1 =0.8<br />
9 l a b e l 1= l a b e l 2=<br />
10 ’ ’ ’ % ( t i t l e , transp , b i a s )<br />
7 This feature is new and experimental and may not work properly yet
154 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
scons <br />
Generate (usually requires .rsf suffix for Flow targets and .vpl<br />
suffix for Plot targets.)<br />
scons<br />
Generate default targets (usually figures specified in Result.)<br />
scons view or scons .view<br />
Generate Result figures and display them on the screen.<br />
scons print or scons .print<br />
Generate Result figures and print them.<br />
scons lock or scons .lock<br />
Generate Result figures and install them in a separate location.<br />
scons test or scons .test<br />
Generate Result figures and compare them with the corresponding “locked”<br />
figures stored in a separate location (regression testing).<br />
scons .flip<br />
Generate the figure and compare it with the corresponding<br />
“locked” figure stored in a separate location by flipping between the two<br />
figures on the screen.<br />
scons TIMER=y ...<br />
Time the execution of each step in the processing flow (using the Unix time<br />
utility.)<br />
scons CHECKPAR=y ...<br />
Check the names and values of all parameters supplied to <strong>Madagascar</strong> modules<br />
in the processing flows before executing anything (guards against incorrect<br />
input.) This option is new and experimental.<br />
Table 2: SCons commands and options defined in rsf.proj.
<strong>Madagascar</strong> Documentation Reproducible research 155<br />
Figure 3: Top left: original image. Top right: random noise added. Bottom left:<br />
original image spectrum in the Fourier (F -X) domain. Bottom right: noisy image<br />
spectrum in the Fourier (F -X) domain. scons/rsfpy panel1<br />
Figure 4: Left: denoising by low-pass filtering. Right: denoising by hard thresholding<br />
in the Fourier domain. scons/rsfpy panel2
156 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
This Python function, named grey(), can then be called in Plot or Result commands,<br />
e.g.<br />
48 Plot ( ’ l p l e n a ’ , grey ( ’ Noisy Lena LP f i l t e r e d ’ ) )<br />
We can define a Python dictionary, e.g.<br />
34 t i t l e s = { ’ lena ’ : ’ Lena ’ ,<br />
35 ’ nlena ’ : ’ Noisy Lena ’ }<br />
and loop over its entries, e.g.<br />
36 for name in t i t l e s . keys ( ) :<br />
37 Plot (name , grey ( t i t l e s [ name ] ) )<br />
38 c f t i t l e = t i t l e s [ name]+ ’ in FX domain ’<br />
39 Flow ( ’ fx ’+name , name , ’ s f s p e c t r a ’ )<br />
40 Plot ( ’ fx ’+name , grey ( c f t i t l e , ’ y ’ , 1 0 0 ) )<br />
Note that the title of the plots is obtained by concatenating Python strings.<br />
Python strings can also be used to define sequences of commands used in several<br />
Flows, e.g.<br />
65 # 2−D FFT<br />
66 f f t 2 = ’ s f f f t 1 sym=y | s f f f t 3 sym=y ’<br />
67 Flow ( ’ f n l e n a ’ , ’ nlena ’ , f f t 2 )<br />
Finally, in our <strong>Madagascar</strong> reproducible script, we may want the option to pass<br />
command line arguments when running SCons or use default values otherwise, e.g.<br />
69 # d e n o i s i n g using t h r e s h o l d i n g in the Fourier domain<br />
70 f t h r = f l o a t (ARGUMENTS. get ( ’ f t h r ’ , 70))<br />
71 Flow ( ’ f t h r l e n a ’ , ’ f n l e n a ’ , ’ s f t h r thr=%f mode=”hard ” ’ % f t h r )<br />
Running scons only, the default value set for fthr (i.e. 70) is used whereas running<br />
scons fthr=68 set fthr to a command line specified value.<br />
This is by no mean an exhaustive list of options but, hopefully, it gives you a<br />
flavor of the powerful tool you have in hands. Enjoy!<br />
CREATING REPRODUCIBLE DOCUMENTATION<br />
You are done with computational experiments and want to communicate them in a<br />
paper. SCons helps us create high-quality papers, where computational results (figures)<br />
are integrated with papers written in LA TEXṪhe corresponding SCons extension<br />
is defined in $RSFROOT/lib/rsftex.py where RSFROOT is the environmental variable<br />
to the <strong>Madagascar</strong> installation directory. The source of this <strong>file</strong> is in python/rsftex.py.<br />
We summarize the basic methods and commands in Tables 3 and 4.
<strong>Madagascar</strong> Documentation Reproducible research 157<br />
Paper(,[,lclass][,use][,include][,options])<br />
A rule to compile .tex L A TEX document using the L A TEX2e<br />
class specified in lclass (default is geophysics.cls from the SEGTeX package)<br />
with additional options specified in options, additional packages specified<br />
in use, and additional preamble specified in include<br />
End()<br />
A rule to collect default targets (referring to paper.tex document).<br />
Table 3: Basic methods of an rsf.tex object.<br />
scons<br />
Generate the default target (usually the <strong>PDF</strong> <strong>file</strong> paper.pdf from the source<br />
L A TEX <strong>file</strong> paper.tex.)<br />
scons pdf or scons .pdf<br />
Generate <strong>PDF</strong> <strong>file</strong>s from L A TEX sources paper.tex or .tex.<br />
scons read or scons .read<br />
Generate <strong>PDF</strong> <strong>file</strong>s from L A TEX sources paper.tex or .tex<br />
and display them on the screen.<br />
scons print or scons .print<br />
Generate <strong>PDF</strong> <strong>file</strong>s from L A TEX sources paper.tex or .tex<br />
and print them.<br />
scons html or scons .html<br />
Generate HTML <strong>file</strong>s from L A TEX sources paper.tex or .tex<br />
using L A TEXtoHTML. The directory html gets created.<br />
scons install or scons .install<br />
Generate <strong>PDF</strong> and HTML <strong>file</strong>s from L A TEX sources paper.tex or<br />
.tex and install them in a separate location (used for publishing<br />
on a web site).<br />
scons wiki or scons .wiki<br />
Convert L A TEX sources paper.tex or .tex to the MediaWiki<br />
format (used for publishing on a Wiki web site).<br />
Table 4: SCons commands defined in rsf.tex.
158 Fomel & Hennenfent <strong>Madagascar</strong> Documentation<br />
Example<br />
This paper by itself is an example of a reproducible document. It is generated using<br />
the following SConstruct <strong>file</strong> which is place in the directory above the projects<br />
directories.<br />
1 from r s f . tex import ∗<br />
2 Paper ( ’ velan ’ , use=’ hyperref , l i s t i n g s , c o l o r ’ )<br />
3 End( use=’ hyperref , l i s t i n g s , c o l o r ’ ,<br />
4 c o l o r=’ modl modl2 cdp1500 cdp2000 cdp2500 cdp3000 cdp3500 pick ’ )<br />
This SConstruct generates this paper but it can also compile velan.tex in the<br />
same directory. Note that there is no Paper command for paper.tex since it is the<br />
default documentation name. Optional L A TEX packages and style used in paper.tex<br />
are passed in the End command.<br />
Let’s now have a closer look at paper.tex to understand how the figures of<br />
the documentation are linked to the reproducible scripts that created them. First<br />
of all, note that paper.tex is not a regular L A TEX document but only its body<br />
(no \documentclass, \usepackage, etc.). In our paper, Fig. 2 was created in the<br />
project folder easystart (sub-folder of our documentation folder) by the result plot<br />
lena.vpl. In the L A TEX source code, it translates as<br />
432 \ i n p u t d i r { e a s y s t a r t }<br />
433 \ s i d e p l o t { lena }{ height =.25\ textheight }{The output o f the f i r s t<br />
434 numerical experiment . }<br />
The \inputdir command points to the project directory and the \sideplot command<br />
calls . The L A TEX tag of the figure is fig:.<br />
The first time the paper is compiled, the result <strong>file</strong> is automatically converted to the<br />
<strong>PDF</strong> <strong>file</strong> format.<br />
REFERENCES<br />
Bivand, R., 2006, Implementing spatial data analysis software tools in r: Geographical<br />
Analysis, 38, 23–40.<br />
Buckheit, J., and D. L. Donoho, 1995, Wavelab and reproducible research, in Wavelets<br />
and Statistics: Springer-Verlag, 103, 55–81.<br />
Claerbout, J., 1992a, Electronic documents give reproducible research a new meaning:<br />
62nd Ann. Internat. Mtg, Soc. of Expl. Geophys., 601–604.<br />
——–, 1992b, How to use Cake with interactive documents, in SEP-73: Stanford<br />
Exploration Project, 451–460.<br />
Claerbout, J. F., and M. Karrenbach, 1993, How to use cake with interactive documents,<br />
in SEP-77: Stanford Exploration Project, 427–444.
<strong>Madagascar</strong> Documentation Reproducible research 159<br />
Claerbout, J. F., and D. Nichols, 1990, Why active documents need cake, in SEP-67:<br />
Stanford Exploration Project, 145–148.<br />
Dubois, P. F., 2003, Why Johnny can’t build: Computing in Science & Engineering,<br />
5, 83–88.<br />
Fomel, S., M. Schwab, and J. Schroeder, 1997, Empowering SEP’s documents, in<br />
SEP-94: Stanford Exploration Project, 339–361.<br />
Gentleman, R. C., V. J. Carey, D. M. Bates, B. Bolstad, M. Dettling, S. Dudoit, B.<br />
Ellis, L. Gautier, Y. Ge, J. Gentry, K. Hornik, T. Hothorn, W. Huber, S. Iacus, R.<br />
Irizarry, F. Leisch, C. Li, M. Maechler, A. J. Rossini, G. Sawitzki, C. Smith, G.<br />
Smyth, L. Tierney, J. Y. Yang, and J. Zhang, 2004, Bioconductor: open software<br />
development for computational biology and bioinformatics: Genome Biology, 5,<br />
R80.<br />
Hubbard, B. B., 1998, The world according to wavelets: The story of a mathematical<br />
technique in the making: AK Peters.<br />
Knuth, D. E., 1984, Literate programming: Computer Journal, 27, 97–111.<br />
LeVeque, R. J., to appear, 2006, Wave propagation software, computational science,<br />
and reproducible research: Presented at the Proc. International Congress of Mathematicians.<br />
Mallat, S., 1999, A wavelet tour of signal processing: Academic Press.<br />
Nichols, D., and S. Cole, 1989, Device independent software installation with CAKE,<br />
in SEP-61: Stanford Exploration Project, 341–344.<br />
Raymond, E. S., 2004, The art of UNIX programming: Addison-Wesley.<br />
Rossum, G. V., 2000a, Python reference manual: Iuniverse Inc.<br />
——–, 2000b, Python tutorial: Iuniverse Inc.<br />
Scales, J. A., and H. Ecke, 2002, What programming languages should we teach our<br />
undergraduates?: The Leading Edge, 21, 260–267.<br />
Schwab, M., M. Karrenbach, and J. Claerbout, 2000, Making scientific computations<br />
reproducible: Computing in Science & Engineering, 2, 61–67.<br />
Schwab, M., and J. Schroeder, 1995, Reproducible research documents using GNUmake,<br />
in SEP-89: Stanford Exploration Project, 217–226.<br />
Sigmon, K., and T. A. Davis, 2001, MATLAB primer, sixth edition: Chapman &<br />
Hall.<br />
Stallman, R. M., R. McGrath, and P. D. Smith, 2004, GNU make: A program for<br />
directing recompilation: GNU Press.<br />
Thimbleby, H., 2003, Explaining code for publication: Software - Practice & Experience,<br />
33, 975–908.<br />
van der Linden, P., 1994, Expert C programming: Prentice Hall.
160 Fomel & Hennenfent <strong>Madagascar</strong> Documentation