- Page 1 and 2: BioinformaticsGenomics& Medicinehtt
- Page 3 and 4: Computational Goals of Bioinformati
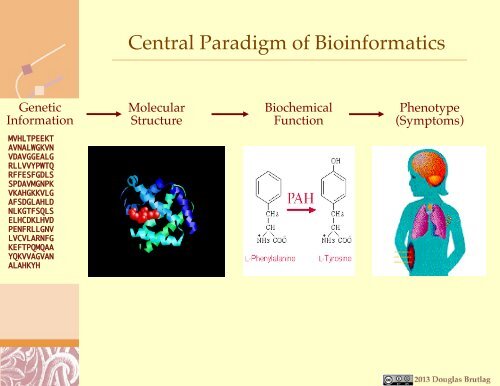
- Page 5: Central Paradigm of MedicineDNA RNA
- Page 10 and 11: Challenges Understanding Genetic In
- Page 12 and 13: Discovering Function from Protein S
- Page 14 and 15: Discovering Function from Protein S
- Page 16 and 17: Protein Motifs fromMultiple Sequenc
- Page 18 and 19: Discovering Function from Protein S
- Page 20 and 21: Protein Motifs fromMultiple Sequenc
- Page 22 and 23: Blocks or Finger Prints fromMultipl
- Page 24 and 25: Discovering Function from Protein S
- Page 26 and 27: Data Mining:The Seach for Buried Tr
- Page 28 and 29: Data Mining:The Seach for Buried Tr
- Page 30 and 31: Swiss Institute of Bioinformaticsht
- Page 32 and 33: Expasy Bioinformatics Resource Port
- Page 34 and 35: UniProt Opsin Entrieshttp://www.uni
- Page 36 and 37: UniProt Human Opsin Entries Reviewe
- Page 38 and 39: Blast UniProt Human Opsin OPN1MW En
- Page 40 and 41: NCBI BLAST Home Pagehttp://blast.nc
- Page 42 and 43: NCBI BLAST Parametershttp://blast.n
- Page 44 and 45: Entrez Gene search for Colorblindne
- Page 46 and 47: Entrez Gene search for Colorblindne
- Page 48 and 49: Entrez Gene search for Opsins
- Page 50 and 51: Choose Standard Protein-Protein BLA
- Page 52 and 53: Optional Parameters
- Page 54 and 55: Sequence Aligned with Domain
- Page 56 and 57:
Most Signifcant Similarity Hits
- Page 58 and 59:
Discovering Function from Protein S
- Page 60 and 61:
Evaluation of ProflesSpecificity=TN
- Page 62 and 63:
MyHits Local Motifs Queryhttp://myh
- Page 64 and 65:
MyHits Local Motifs Summaryhttp://m
- Page 66 and 67:
MyHits Local Motifs Hist (Cont.)htt
- Page 68 and 69:
MyHits Local Motifs Hist (Cont.)
- Page 70 and 71:
InterProScanhttp://www.ebi.ac.uk/in
- Page 72 and 73:
InterPro Scan HourGlasshttp://www.e
- Page 74 and 75:
InterPro Scan Resultshttp://www.ebi
- Page 76 and 77:
GO: Gene Ontology for Opsin OPN1MWh
- Page 78 and 79:
GO: Sequence Information for OPN1MW
- Page 80 and 81:
GO: Gene Ontology Databasehttp://ww
- Page 82 and 83:
GO: Gene Ontology Term GCRPhttp://w
- Page 84 and 85:
GO: Gene Ontology GCPR Termhttp://w
- Page 86 and 87:
Statistical vs. Biological Signifca
- Page 88 and 89:
Statistical vs. Biological Signifca
- Page 90 and 91:
Multiple Enhancer Sequences
- Page 92 and 93:
PolyAdenylation of mRNAs© Michael
- Page 94 and 95:
Splicing, Capping & polyAdenylation
- Page 96 and 97:
Alternative Splicing Generates Dist
- Page 98 and 99:
Alternative SplicingDetected in EST
- Page 100 and 101:
Gene Locihttp://www.ncbi.nlm.nih.go
- Page 102 and 103:
Genomics, Bioinformatics &Computati
- Page 104 and 105:
Redundancy in Genomic& Protein Sequ
- Page 106 and 107:
pFAM at Sanger Center (UK)http://ww
- Page 108 and 109:
pFAM at Sanger Center (UK)http://pf
- Page 110 and 111:
pFAM at Sanger Center (UK)http://pf