Review ArticleGenetics <strong>and</strong> EpilepsyThe completion of the human genome project <strong>and</strong> thefurther availability of <strong>in</strong>creas<strong>in</strong>gly detailed data onhuman chromosomes 1 promise much for our underst<strong>and</strong><strong>in</strong>gof the genetic basis of human health <strong>and</strong> disease.The entire human genome conta<strong>in</strong>s some 30,000 genes,more than half of which are expressed at some stage <strong>in</strong> thebra<strong>in</strong>. Genetic <strong>in</strong>fluences on bra<strong>in</strong> function <strong>and</strong> diseases aretherefore pervasive. In the outbred human species, epilepsyis obviously not a constitutive phenotype: there are certa<strong>in</strong>genetic variants that either directly cause or contribute toepilepsy phenotypes. Epilepsy is a heterogeneous group ofconditions, <strong>and</strong> thus genetic <strong>in</strong>fluences on different types ofepilepsy are likely to vary widely. However, whilst genesencode every prote<strong>in</strong>, many prote<strong>in</strong>s undergo additionalmodulation by factors rang<strong>in</strong>g from the post-transcriptionalto the macroenvironmental, so that we, <strong>and</strong> our diseases,are generally not products of a rigid genetic determ<strong>in</strong>ism.However, genetic factors are perhaps now morereadily determ<strong>in</strong>ed than environmental factors, so thatgenetic studies <strong>in</strong> epilepsy hold promise for a better underst<strong>and</strong><strong>in</strong>g<strong>and</strong> more rational treatment than that which currentlyexists.Genes could <strong>in</strong>fluence seizures, epileptogenesis <strong>and</strong>epilepsy at multiple levels. Genetic variation could affectthe aetiology, susceptibility, mechanisms, syndrome, treatmentresponse, prognosis <strong>and</strong> consequences of the epilepsiesto vary<strong>in</strong>g degrees <strong>in</strong> different <strong>in</strong>dividuals. Part of thepromise of genetics lies <strong>in</strong> its power to relate these facets ofthe overall cl<strong>in</strong>ical presentation to the <strong>in</strong>dividual patient.There has been considerable recent progress, though thishas focused largely on aetiology, susceptibility <strong>and</strong> treatmentresponse.More <strong>and</strong> more genetic mutations are be<strong>in</strong>g identifiedthat cause epilepsy. These mutations usually cause conditionsthat are <strong>in</strong>herited <strong>in</strong> a classical Mendelian fashion:autosomal dom<strong>in</strong>ant, autosomal recessive, X-l<strong>in</strong>ked orthrough mitochondrial <strong>in</strong>heritance. Mutations are rare<strong>and</strong>, even collectively, mutational causes of conditions <strong>in</strong>which epilepsy is the sole or ma<strong>in</strong> manifestation accountfor only a small proportion of cases of epilepsy. Perhapsunsurpris<strong>in</strong>gly given the central position of neuronalexcitability <strong>in</strong> epilepsy, most mutations so far uncoveredthat lead to epilepsy occur <strong>in</strong> genes encod<strong>in</strong>g ion channels. 2Sodium, potassium, calcium <strong>and</strong> chloride channelopathieshave all been described, <strong>and</strong> there are likely to be others tocome (Table 1). It is worth not<strong>in</strong>g, <strong>in</strong> pass<strong>in</strong>g, that acquiredepileptogenic channelopathies also exist. 3 Mutations thatcause monogenic epilepsies fall <strong>in</strong>to two other major categories:those that lead to structural bra<strong>in</strong> malformationsone cl<strong>in</strong>ical manifestation of which is epilepsy; 4 <strong>and</strong> thosethat produce the progressive myoclonic epilepsies (PMEs). 5The latter are themselves a diverse group of conditions dist<strong>in</strong>guishedby particular natural histories, pathophysiologies<strong>and</strong> <strong>in</strong>vestigational f<strong>in</strong>d<strong>in</strong>gs, but which broadly sharethe common characteristics of myoclonic jerks, otherseizure types <strong>and</strong> progressive cognitive decl<strong>in</strong>e: many PMEscan now be def<strong>in</strong>ed <strong>and</strong> dissected genetically (Table 2).Only a few other genetic mutations are known to lead toepilepsy as their major consequence. These <strong>in</strong>clude LGI1,mutations which cause a familial focal epilepsy. 6 There areof course a large number of Mendelian conditions withknown underly<strong>in</strong>g gene mutations that cause multisystemicconditions <strong>in</strong> which seizures are part of a broaderphenotype: the number of these also cont<strong>in</strong>ues to grow, butaga<strong>in</strong> account for only a small proportion of the epilepsies.This progress <strong>in</strong> the genetic aetiology of the epilepsiesis remarkable, <strong>and</strong> as yet unmatched by developments <strong>in</strong>the genetics of other aspects of disease biology, such assusceptibility or treatment response, reflect<strong>in</strong>g partly thefocus of researchers so far, but also the tractability of therelevant issues.Genetic variation <strong>in</strong>fluenc<strong>in</strong>g susceptibility to non-Mendelian epilepsies has been the other major focus ofresearch. Most epilepsies are complex traits <strong>and</strong> probablyarise <strong>in</strong> <strong>in</strong>dividuals as a result of gene-gene <strong>and</strong> gene-environment<strong>in</strong>teractions. The numbers of genes <strong>in</strong>volved areunknown, <strong>and</strong> may be few or many. The result<strong>in</strong>g patternsof <strong>in</strong>heritance <strong>in</strong> the majority of cases are thus more complicated<strong>and</strong> subtle: such <strong>in</strong>heritance falls outside theMendelian patterns to which we have become accustomed.As a corollary, large cohorts of patients are needed to extractthe risks attributable to particular common genetic variants.Many positive associations between particular gene variants<strong>and</strong> def<strong>in</strong>ed epilepsy syndromes have been found; it is likelythat many more negative results have not made it to theeditor’s <strong>in</strong>-tray. However, no common genetic variants areyet accepted as genu<strong>in</strong>ely <strong>in</strong>creas<strong>in</strong>g the risk of any particularepilepsy type or syndrome. 7 Numerous problems ofmethodology dog this area of research. Perhaps the mostimportant is that any one research centre is unlikely to beable to recruit sufficient numbers of patients with an appropriatelyhomogeneous phenotype to have adequate powerto detect <strong>in</strong>dividual gene effects which are <strong>in</strong> practice likelyto be m<strong>in</strong>or: a parallel with <strong>in</strong>ternational cl<strong>in</strong>ical trialsrecruit<strong>in</strong>g thous<strong>and</strong>s to show m<strong>in</strong>or benefits from a newtreatment regime may be drawn. However, as our underst<strong>and</strong><strong>in</strong>gof the genome <strong>and</strong> its vagaries improves, <strong>and</strong> with<strong>in</strong>creased experience <strong>and</strong> <strong>in</strong>ternational collaboration, itseems likely that common genetic variants driv<strong>in</strong>g commondisease processes will emerge.The genetics of drug response may prove to be moreamenable to analysis than other aspects of genetics <strong>in</strong>epilepsy, because the prote<strong>in</strong>s that are drug targets, drugtransporters <strong>and</strong> drug metabolisers are to vary<strong>in</strong>g extentsalready known. Their encod<strong>in</strong>g genes do not need to bepicked, at r<strong>and</strong>om or otherwise, from 30,000 genes, 8 <strong>and</strong>many have been thoroughly characterised. 9 For example, itis well established that <strong>in</strong>dividuals who possess certa<strong>in</strong> allelesof the CYP2C9 gene, that encodes the major metabolis<strong>in</strong>genzyme of phenyto<strong>in</strong>, have significantly reduced ratesof metabolism of phenyto<strong>in</strong>, necessitat<strong>in</strong>g lower ma<strong>in</strong>tenancedoses, 10 although prospective genotyp<strong>in</strong>g is not yetundertaken <strong>in</strong> practice. Whether variants <strong>in</strong> the ABCB1gene, that encodes the broad-spectrum multidrug transporterP-glycoprote<strong>in</strong>, <strong>in</strong>fluence resistance to antiepilepticdrugs or not rema<strong>in</strong>s a hotly-debated po<strong>in</strong>t. 11,12 Gene variants<strong>in</strong>fluenc<strong>in</strong>g the sensitivity of targets to antiepilepticdrugs are also be<strong>in</strong>g uncovered: for example, a splice sitevariation <strong>in</strong> the SCN1A gene that encodes the cerebral neuronaltarget of many antiepileptic drugs, has been associatedwith dos<strong>in</strong>g of these drugs. 10 Such pharmacogeneticadvances, if substantiated by further studies <strong>and</strong> proven <strong>in</strong>structured trials to be cl<strong>in</strong>ically significant, may permitcloser modell<strong>in</strong>g of treatment to the <strong>in</strong>dividual patient.The genetics of most other biological facets of epilepsyhave not yet even been considered <strong>in</strong> any detail, but itrema<strong>in</strong>s possible that genetic variation will also have animpact of cl<strong>in</strong>ical relevance <strong>in</strong> these other areas (eg biologicalconsequences of epilepsy <strong>in</strong> an <strong>in</strong>dividual patient).Much is still expected of genetic research: careful evaluation<strong>in</strong> the cl<strong>in</strong>ical sett<strong>in</strong>g will rema<strong>in</strong> critical to establish<strong>in</strong>g thepractical utility of such research. Close collaborationbetween epilepsy cl<strong>in</strong>ical centres, <strong>and</strong> between cl<strong>in</strong>ical <strong>and</strong>laboratory scientists, <strong>and</strong> accurate def<strong>in</strong>ition of phenotypes,will be the key to br<strong>in</strong>g<strong>in</strong>g epilepsy genetics to thecl<strong>in</strong>ic for the benefit of patients <strong>and</strong> society.Dr Sisodiya works <strong>in</strong> the Departmentof Cl<strong>in</strong>ical <strong>and</strong> ExperimentalEpilepsy, Institute of Neurology,UCL, London. His cl<strong>in</strong>ical <strong>and</strong>research <strong>in</strong>terests are <strong>in</strong> the neurogenetic,neuropathological <strong>and</strong>developmental biological aspectsof epilepsy, recently focus<strong>in</strong>g ongenetic variation, disease susceptibility<strong>and</strong> treatment response.Correspondence to:SM Sisodiya MA, PhD, FRCP,Reader <strong>in</strong> Neurology,Department of Cl<strong>in</strong>ical <strong>and</strong>Experimental Epilepsy,National Hospital for Neurology<strong>and</strong> Neurosurgery, Box 29,Queen Square, London WC1N 3BG.Tel: 020 7837 3611,Fax: 020 7837 3941,Email: sisodiya@ion.ucl.ac.uk10 I <strong>ACNR</strong> • VOLUME 5 NUMBER 6 • JANUARY/FEBRUARY 2006
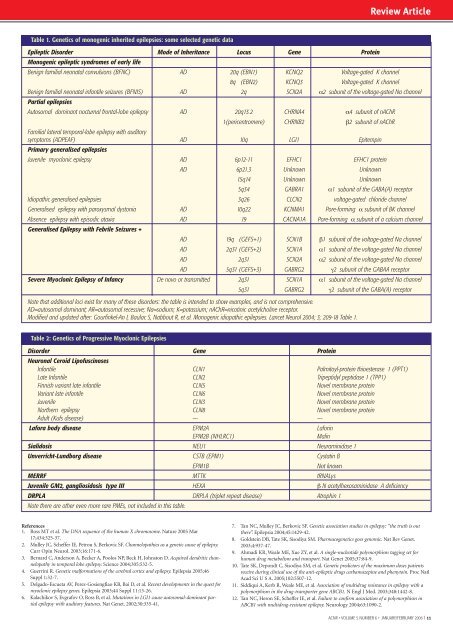
Review ArticleTable 1. Genetics of monogenic <strong>in</strong>herited epilepsies: some selected genetic dataEpileptic Disorder Mode of Inheritance Locus Gene Prote<strong>in</strong>Monogenic epileptic syndromes of early lifeBenign familial neonatal convulsions (BFNC) AD 20q (EBN1) KCNQ2 Voltage-gated K channel8q (EBN2) KCNQ3 Voltage-gated K channelBenign familial neonatal <strong>in</strong>fantile seizures (BFNIS) AD 2q SCN2A α2 subunit of the voltage-gated Na channelPartial epilepsiesAutosomal dom<strong>in</strong>ant nocturnal frontal-lobe epilepsy AD 20q13.2 CHRNA4 α4 subunit of nAChR1(pericentromere) CHRNB2 β2 subunit of nAChRFamilial lateral temporal-lobe epilepsy with auditorysymptoms (ADPEAF) AD 10q LGI1 Epitemp<strong>in</strong>Primary generalised epilepsiesJuvenile myoclonic epilepsy AD 6p12-11 EFHC1 EFHC1 prote<strong>in</strong>AD 6p21.3 Unknown Unknown15q14 Unknown Unknown5q34 GABRA1 α1 subunit of the GABA(A) receptorIdiopathic generalised epilepsies 3q26 CLCN2 voltage-gated chloride channelGeneralised epilepsy with paroxysmal dystonia AD 10q22 KCNMA1 Pore-form<strong>in</strong>g α subunit of BK channelAbsence epilepsy with episodic ataxia AD 19 CACNA1A Pore-form<strong>in</strong>g α subunit of a calcium channelGeneralised Epilepsy with Febrile Seizures +AD 19q (GEFS+1) SCN1B β1 subunit of the voltage-gated Na channelAD 2q31 (GEFS+2) SCN1A α1 subunit of the voltage-gated Na channelAD 2q31 SCN2A α2 subunit of the voltage-gated Na channelAD 5q31 (GEFS+3) GABRG2 γ2 subunit of the GABAA receptorSevere Myoclonic Epilepsy of Infancy De novo or transmitted 2q31 SCN1A α1 subunit of the voltage-gated Na channel5q31 GABRG2 γ2 subunit of the GABA(A) receptorNote that additional loci exist for many of these disorders: the table is <strong>in</strong>tended to show examples, <strong>and</strong> is not comprehensive.AD=autosomal dom<strong>in</strong>ant; AR=autosomal recessive; Na=sodium; K=potassium; nAChR=nicot<strong>in</strong>ic acetylchol<strong>in</strong>e receptor.Modified <strong>and</strong> updated after: Gourf<strong>in</strong>kel-An I, Baulac S, Nabbout R, et al. Monogenic idiopathic epilepsies. Lancet Neurol 2004; 3; 209-18 Table 1.Table 2: Genetics of Progressive Myoclonic EpilepsiesDisorder Gene Prote<strong>in</strong>Neuronal Ceroid Lipofusc<strong>in</strong>osesInfantile CLN1 Palmitoyl-prote<strong>in</strong> thioesterase 1 (PPT1)Late Infantile CLN2 Tripeptidyl peptidase 1 (TPP1)F<strong>in</strong>nish variant late <strong>in</strong>fantile CLN5 Novel membrane prote<strong>in</strong>Variant late <strong>in</strong>fantile CLN6 Novel membrane prote<strong>in</strong>Juvenile CLN3 Novel membrane prote<strong>in</strong>Northern epilepsy CLN8 Novel membrane prote<strong>in</strong>Adult (Kufs disease) —- —-Lafora body disease EPM2A Lafor<strong>in</strong>EPM2B (NHLRC1)Mal<strong>in</strong>Sialidosis NEU1 Neuram<strong>in</strong>idase 1Unverricht-Lundborg disease CSTB (EPM1) Cystat<strong>in</strong> BEPM1BNot knownMERRF MTTK tRNALysJuvenile GM2, gangliosidosis type III HEXA β N acetylhexosam<strong>in</strong>idase A deficiencyDRPLA DRPLA (triplet repeat disease) Atroph<strong>in</strong> 1Note there are other even more rare PMEs, not <strong>in</strong>cluded <strong>in</strong> this table.References1. Ross MT et al. The DNA sequence of the human X chromosome. Nature 2005 Mar17;434:325-37.2. Mulley JC, Scheffer IE, Petrou S, Berkovic SF. Channelopathies as a genetic cause of epilepsy.Curr Op<strong>in</strong> Neurol. 2003;16:171-6.3. Bernard C, Anderson A, Becker A, Poolos NP, Beck H, Johnston D. Acquired dendritic channelopathy<strong>in</strong> temporal lobe epilepsy. Science 2004;305:532-5.4. Guerr<strong>in</strong>i R. Genetic malformations of the cerebral cortex <strong>and</strong> epilepsy. Epilepsia 2005;46Suppl 1:32-7.5. Delgado-Escueta AV, Perez-Gosiengfiao KB, Bai D, et al. Recent developments <strong>in</strong> the quest formyoclonic epilepsy genes. Epilepsia 2003;44 Suppl 11:13-26.6. Kalachikov S, Evgrafov O, Ross B, et al. Mutations <strong>in</strong> LGI1 cause autosomal-dom<strong>in</strong>ant partialepilepsy with auditory features. Nat Genet. 2002;30:335-41.7. Tan NC, Mulley JC, Berkovic SF. Genetic association studies <strong>in</strong> epilepsy: “the truth is outthere”. Epilepsia 2004;45:1429-42.8. Goldste<strong>in</strong> DB, Tate SK, Sisodiya SM. Pharmacogenetics goes genomic. Nat Rev Genet.2003;4:937-47.9. Ahmadi KR, Weale ME, Xue ZY, et al. A s<strong>in</strong>gle-nucleotide polymorphism tagg<strong>in</strong>g set forhuman drug metabolism <strong>and</strong> transport. Nat Genet 2005;37:84-9.10. Tate SK, Depondt C, Sisodiya SM, et al. Genetic predictors of the maximum doses patientsreceive dur<strong>in</strong>g cl<strong>in</strong>ical use of the anti-epileptic drugs carbamazep<strong>in</strong>e <strong>and</strong> phenyto<strong>in</strong>. Proc NatlAcad Sci U S A. 2005;102:5507-12.11. Siddiqui A, Kerb R, Weale ME, et al. Association of multidrug resistance <strong>in</strong> epilepsy with apolymorphism <strong>in</strong> the drug-transporter gene ABCB1. N Engl J Med. 2003;348:1442-8.12. Tan NC, Heron SE, Scheffer IE, et al. Failure to confirm association of a polymorphism <strong>in</strong>ABCB1 with multidrug-resistant epilepsy. Neurology 2004;63:1090-2.<strong>ACNR</strong> • VOLUME 5 NUMBER 6 • JANUARY/FEBRUARY 2006 I 11