Molecular phylogeny of the mycorrhizal desert truffles ... - Mycologia
Molecular phylogeny of the mycorrhizal desert truffles ... - Mycologia
Molecular phylogeny of the mycorrhizal desert truffles ... - Mycologia
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
<strong>Mycologia</strong>, 94(2), 2002, pp. 247–259.<br />
2002 by The Mycological Society <strong>of</strong> America, Lawrence, KS 66044-8897<br />
Jesús Díez 1<br />
<strong>Molecular</strong> <strong>phylogeny</strong> <strong>of</strong> <strong>the</strong> <strong>mycorrhizal</strong> <strong>desert</strong> <strong>truffles</strong> (Terfezia and<br />
Tirmania), host specificity and edaphic tolerance<br />
UMR 1136 INRA-UHP ‘‘Interactions Arbres/<br />
Micro-organismes’’, INRA-Nancy, F-54280<br />
Champenoux, France<br />
José Luis Manjón<br />
Dpto. de Biología Vegetal, Universidad de Alcalá, E-<br />
28871 Alcalá de Henares (Madrid), Spain<br />
Francis Martin<br />
UMR 1136 INRA-UHP ‘‘Interactions Arbres/<br />
Micro-organismes’’, INRA -Nancy, F-54280<br />
Champenoux, France<br />
Abstract: Terfezia and Tirmania, so called <strong>desert</strong><br />
<strong>truffles</strong>, are <strong>mycorrhizal</strong> fungi mostly endemic to arid<br />
and semi-arid areas <strong>of</strong> <strong>the</strong> Mediterranean Region,<br />
where <strong>the</strong>y are associated with Helian<strong>the</strong>mum species.<br />
The aim <strong>of</strong> this work was to study <strong>the</strong> phylogenetic<br />
relationships in <strong>the</strong>se pezizalean hypogeous fungi.<br />
The restriction fragment length polymorphism<br />
(RFLP) and DNA sequences <strong>of</strong> internal transcribed<br />
spacers (ITS) <strong>of</strong> <strong>the</strong> nuclear rDNA were studied for<br />
several morphological species, Terfezia arenaria, T.<br />
boudieri, T. claveryi, T. leptoderma, T. terfezioides<br />
(Mattirolomyces terfezioides), Tirmania nivea and T.<br />
pinoyi. The sequences were analyzed with distance<br />
and parsimony methods. Phylogenetic analyses indicated<br />
a close genetic relationship between Tirmania<br />
and Terfezia. They may have arisen from a single evolutionary<br />
lineage <strong>of</strong> pezizalean fungi that developed<br />
<strong>the</strong> hypogeous habit as an adaptation to heat and<br />
drought in Mediterranean ecosystems. This analysis<br />
also supports <strong>the</strong> re-establishment <strong>of</strong> <strong>the</strong> genus Mattirolomyces.<br />
The genera Tirmania and Terfezia were<br />
monophyletic, and morphological species corresponded<br />
to phylogenetic species. The Tirmania clade<br />
comprises <strong>desert</strong> <strong>truffles</strong> with smooth spores and amyloid<br />
asci, which were found in <strong>desert</strong>s. The Terfezia<br />
clade grouped species found in semi-arid habitats<br />
having ornamented and spherical spores. These species<br />
are adapted to exploit different types <strong>of</strong> soil (ei<strong>the</strong>r<br />
acid or basic soils) in association with specific<br />
hosts (ei<strong>the</strong>r basophilous or acidophilous species).<br />
Although o<strong>the</strong>r factors might also play a role, host<br />
specialization and edaphic tolerances (fungus and/<br />
Accepted for publication August 7, 2001.<br />
1 Corresponding author, Email: diezmuriel@yahoo.com<br />
247<br />
or host tolerances) might be <strong>the</strong> key in <strong>the</strong> species<br />
diversity <strong>of</strong> <strong>the</strong>se genera.<br />
Key Words: fungal evolution, Helian<strong>the</strong>mum, internal<br />
transcribed spacer, <strong>mycorrhizal</strong> fungi<br />
INTRODUCTION<br />
Pezizales are widespread Ascomycetes with ei<strong>the</strong>r enclosed<br />
underground (hypogeous) or exposed (epigeous)<br />
fruit bodies (Trappe 1979). The hypogeous<br />
ascocarps <strong>of</strong> <strong>the</strong>se fungi are known as <strong>truffles</strong><br />
(Trappe 1992). In <strong>the</strong> Mediterranean region, hypogeous<br />
fungi colonize a variety <strong>of</strong> forests and semi-arid<br />
ecosystems. These fungi frequently establish mutualistic<br />
associations with vascular plants via specialized<br />
nutrient-ga<strong>the</strong>ring organs called mycorrhizas (Trappe<br />
1992).<br />
Many hypogeous fungi occurring in semi-arid ecosystems<br />
<strong>of</strong> <strong>the</strong> Mediterranean Basin belong to <strong>the</strong><br />
genera Tirmania, Terfezia and Picoa (Alsheikh and<br />
Trappe 1983a, b, Moreno et al 1986, 1991, 2000,<br />
2001). Most <strong>of</strong> <strong>the</strong>m are endemic to <strong>the</strong> Mediterranean<br />
region and establish <strong>mycorrhizal</strong> symbioses with<br />
members <strong>of</strong> <strong>the</strong> Cistaceae, mainly with Helian<strong>the</strong>mum<br />
species (FIG. 1) (Dexheimer et al 1985, Fortas and<br />
Chevalier 1992). These plants and <strong>the</strong>ir associated<br />
mycota may play a major role in <strong>the</strong> maintenance <strong>of</strong><br />
Mediterranean shrublands and xerophytic grasslands,<br />
and thus in preventing erosion and <strong>desert</strong>ification<br />
(Honrubia et al 1992).<br />
Morphological characters have been used to describe<br />
different species <strong>of</strong> <strong>desert</strong> <strong>truffles</strong>; i.e., spore<br />
and peridium morphology, gleba colour, odor and<br />
o<strong>the</strong>r organoleptic characters. These fungi, however,<br />
are difficult to identify at <strong>the</strong> species level. Regarding<br />
<strong>the</strong> systematics <strong>of</strong> <strong>desert</strong> <strong>truffles</strong>, <strong>the</strong> use <strong>of</strong> morphological<br />
features is problematic, because <strong>of</strong> <strong>the</strong> reduced<br />
set <strong>of</strong> morphological characters and <strong>the</strong>ir homoplasy.<br />
Ascocarp features are homoplastic as a result<br />
<strong>of</strong> parallel evolution <strong>of</strong> independent lineages <strong>of</strong><br />
epigeous/hypogeous fruit bodies during <strong>the</strong> evolutionary<br />
history <strong>of</strong> <strong>the</strong> Pezizales (Trappe 1979). This<br />
accounts for <strong>the</strong> early artificial classifications <strong>of</strong> hypogeous<br />
fungi in <strong>the</strong> order Tuberales. Tuberales included<br />
several hypogeous families with similar morphology<br />
originating by parallel or convergent evolution<br />
(O’Donnell et al 1997). Realizing that <strong>the</strong> Tub-
248 MYCOLOGIA<br />
erales artificially grouped many Pezizales, Trappe<br />
(1979) transferred some <strong>of</strong> families from Tuberales<br />
to Pezizales and amended some families in Pezizales<br />
to include related hypogeous fungi. He also amended<br />
<strong>the</strong> Pezizaceae Fries sensu Korf to accommodate<br />
several related hypogeous taxa lacking forcible spore<br />
discharge, such as Tirmania. In addition, based on a<br />
comparative morphological study, Trappe (1979) included<br />
Terfezia in <strong>the</strong> Terfeziaceae and Picoa in <strong>the</strong><br />
Balsamiaceae.<br />
The increasing amount <strong>of</strong> molecular phylogenetic<br />
data now available has led to a continuing revision <strong>of</strong><br />
<strong>the</strong> hypogeous Ascomycetes (Gargas and Taylor 1995,<br />
Spatafora 1995, Landvik et al 1997, O’Donnell et al<br />
1997, Harrington et al 1999). O’Donnell et al (1997)<br />
provided support for <strong>the</strong> occurrence <strong>of</strong> independent<br />
lines <strong>of</strong> epigeous/hypogeous fruit body evolution in<br />
<strong>the</strong> Pezizales. This has affected classification <strong>of</strong> <strong>the</strong><br />
<strong>desert</strong> <strong>truffles</strong>. Moreover, analysis <strong>of</strong> <strong>the</strong> 18S rDNA<br />
sequences has revealed a close relationship between<br />
Terfezia and <strong>the</strong> Pezizaceae (Percudani et al 1999,<br />
Norman and Egger 1999). Picoa is not closely related<br />
to <strong>the</strong> Pezizaceae (see O’Donnell et al 1997), where<br />
Picoa carthusiana Tul. & Tul. is referred to by <strong>the</strong><br />
synonym Leucangium carthusianum (Tul.) Paol.<br />
Hence, it will be not studied in <strong>the</strong> present work.<br />
Subgeneric relationships within <strong>the</strong> Pezizales have<br />
been subjected to molecular analysis in few cases<br />
(Harrington and Potter 1997, Roux et al 1999, Norman<br />
and Egger 1996, 1999). Apart from studies <strong>of</strong><br />
<strong>the</strong> genus Tuber (Roux et al 1999), no recent work<br />
has addressed subgeneric relationships within <strong>the</strong> hypogeous<br />
genera. In particular, no work has been published<br />
on <strong>the</strong> evolutionary history <strong>of</strong> <strong>desert</strong> <strong>truffles</strong>.<br />
The phylogenetic concept <strong>of</strong> species requires that<br />
species represent a monophyletic set <strong>of</strong> organisms. In<br />
<strong>the</strong> case <strong>of</strong> <strong>desert</strong> <strong>truffles</strong>, species delimitation by<br />
morphological characters seems to be consistent.<br />
However, molecular phylogenetic analyses are needed,<br />
in order to verify whe<strong>the</strong>r morphological species<br />
<strong>of</strong> <strong>desert</strong> <strong>truffles</strong> also represent phylogenetic species.<br />
The present work focuses on two major genera <strong>of</strong><br />
pezizalean hypogeous fungi (Tirmania and Terfezia),<br />
and deals with <strong>the</strong> main morphological species that<br />
occur in <strong>the</strong> Mediterranean region. Because <strong>of</strong> <strong>the</strong><br />
limited capacity <strong>of</strong> <strong>the</strong> 18S rDNA to resolve intrageneric<br />
relationships in hypogeous ascomycetes (Percudani<br />
et al 1999), we used <strong>the</strong> faster-evolving internal<br />
transcribed spacer (ITS) <strong>of</strong> <strong>the</strong> nuclear rDNA.<br />
Restriction fragment length polymorphism (RFLP)<br />
analysis was supplemented by sequence analysis. Phylogenic<br />
analysis, toge<strong>the</strong>r with <strong>the</strong> morphological and<br />
ecological data, allowed us to propose an evolutionary<br />
history for <strong>the</strong>se genera.<br />
MATERIALS AND METHODS<br />
Fungal collections. Most specimens studied in this work<br />
were collected by <strong>the</strong> authors. Several collections deposited<br />
at <strong>the</strong> Herbarium <strong>of</strong> Alcalá University (AH) were also studied.<br />
Specimens were harvested in xerophilous shrublands<br />
and grasslands dominated by Helian<strong>the</strong>mum spp. Host and<br />
soil data are mainly based on our observations and on notes<br />
accompanying herbarium specimens. Voucher specimens <strong>of</strong><br />
new collections were deposited in AH (Alcalá University<br />
Herbarium, Spain). Specimen collections are listed in TA-<br />
BLE I.<br />
We studied a wide collection <strong>of</strong> <strong>desert</strong> <strong>truffles</strong> from <strong>the</strong><br />
Iberian Peninsula (Spain and Portugal), Morocco, Tunisia,<br />
Algeria, Israel and Kuwait. Specimens were identified as:<br />
Tirmania nivea (Des. : Fr.) Trappe, T. pinoyi (Maire) Malençon,<br />
Terfezia arenaria (Moris) Trappe, T. boudieri Chatin,<br />
T. claveryi Chatin, or T. leptoderma (described in detail in<br />
Moreno et al 1986, 1999, 2000, 2001). Variability in spore<br />
ornamentation was <strong>the</strong> most important morphological feature<br />
for discriminating between <strong>the</strong>se species. A comparative<br />
table <strong>of</strong> major morphological features is provided (TA-<br />
BLE II).<br />
Mycelial cultures. Isolates were obtained from ascomata on<br />
modified Melin-Norkrans (MMN) agar medium (Marx<br />
1969). All cultures were maintained on MMN agar medium<br />
at 25 C in <strong>the</strong> dark. Isolates were deposited at <strong>the</strong> Collection<br />
<strong>of</strong> Ecto<strong>mycorrhizal</strong> Fungi <strong>of</strong> <strong>the</strong> Plant Biology Department<br />
<strong>of</strong> <strong>the</strong> University <strong>of</strong> Alcalá.<br />
DNA extraction and ITS amplification. DNA was extracted<br />
from specimens belonging to collections <strong>of</strong> <strong>desert</strong> <strong>truffles</strong><br />
from <strong>the</strong> Iberian Peninsula, Morocco, Tunisia, Algeria, Israel,<br />
and Kuwait and two collections <strong>of</strong> Mattirolomyces terfezioides.<br />
Samples for DNA were excised ei<strong>the</strong>r from <strong>the</strong><br />
edge <strong>of</strong> a mycelial colony or from <strong>the</strong> inner part <strong>of</strong> <strong>the</strong><br />
ascocarp to avoid contamination by o<strong>the</strong>r microorganisms.<br />
Mycelial samples were preserved at 20 C pending DNA<br />
extraction. Approximately 20–50 mg <strong>of</strong> tissue was used for<br />
each DNA extraction, performed using <strong>the</strong> cetyl-trimethylammonium<br />
bromide (CTAB) protocol (Gardes et al 1991).<br />
The ITS regions <strong>of</strong> nuclear rDNA were amplified with ITS1<br />
and ITS4 primers (White 1990) as described by Henrion et<br />
al (1992) on a GeneAmp 9600 PCR <strong>the</strong>rmocycler (Perkin<br />
Elmer, Inc.). Controls with no DNA were included in every<br />
set <strong>of</strong> amplifications to test for DNA contamination in reagents<br />
and reaction buffers.<br />
RFLP analysis and DNA electrophoresis. Variation in ITS<br />
sequences was assessed among at least three specimens <strong>of</strong><br />
each collection by restriction fragment length polymorphism<br />
(RFLP) with <strong>the</strong> following enzymes: AluI, Hinf I,<br />
HhaI, and RsaI. Six to 10 L <strong>of</strong> <strong>the</strong> amplified ITS was digested<br />
with 2.5 units <strong>of</strong> each enzyme for 1 h at 37 C, according<br />
to <strong>the</strong> manufacturer’s instructions (GIBCO-BRL,<br />
Pasley, UK). The amplification and digestion products were<br />
fractionated with 1.5% regular agarose gels or 6% polyacrylamide<br />
gels in a 1 Tris-borate-EDTA buffer. The gels were<br />
stained with ethidium bromide and photographed under<br />
ultraviolet light. 174 phage DNA digested with HaeIII<br />
was included in <strong>the</strong> gels as a molecular size marker.
Sequencing <strong>of</strong> <strong>the</strong> amplified ITS regions. ITS regions from<br />
18 specimens displaying different RFLP patterns were sequenced<br />
(TABLE III). Amplified DNA was purified with<br />
GeneClean Kit (Bio101, Carlsbad, California, USA). DNA<br />
concentrations were quantified on agarose gels with a Low<br />
DNA Mass ladder (GIBCO-BRL, Pasley, UK) and sizes calculated<br />
with a 100-bp ladder (GIBCO-BRL). Sequencing reactions<br />
were performed directly on purified PCR products<br />
with ITS1 or ITS4 primers (White et al 1990). Both strands<br />
were sequenced with <strong>the</strong> Taq DyeDeoxy Terminator Cycle<br />
Sequencing Kit (Perkin Elmer, Norwalk, Connecticut,<br />
USA). The sequence products were analyzed with an ABI<br />
model 373 DNA fluorescent sequencer (Perkin Elmer). The<br />
sequences obtained were deposited in <strong>the</strong> National Center<br />
for Biotechnology Information (NCBI) GenBank (TABLE<br />
III).<br />
Sequence alignment and analysis. The search for sequence<br />
identity in <strong>the</strong> GenBank DNA database was conducted by<br />
Gapped BLAST (NCBI) (Altschul et al 1997). Three ITS<br />
sequences <strong>of</strong> T. boudieri were retrieved from GenBank and<br />
added to <strong>the</strong> alignment data set. The ITS sequences <strong>of</strong> <strong>the</strong><br />
closely related species <strong>of</strong> Peziza (Norman and Egger 1999)<br />
were also retrieved from GenBank in order to verify whe<strong>the</strong>r<br />
<strong>the</strong> Tirmania/Terfezia clade is monophyletic after introduction<br />
<strong>of</strong> Peziza species. Sequences were aligned with<br />
MultAlin (Corpet 1988) at <strong>the</strong> ProDom web site (http://<br />
www.toulouse.inra.fr/multalin.html) (INRA-CNRS, Toulouse).<br />
The sequence alignment was hand-edited and deposited<br />
in TreeBASE (accession number SN908) (http://<br />
www.herbaria.harvard.edu/treebase/).<br />
Aligned sequences were analyzed by means <strong>of</strong> distance<br />
and parsimony methods. Tuber melanosporum was used as<br />
outgroup (GenBank accession no. U89359). Distances were<br />
calculated according to <strong>the</strong> Junkes and Cantor model, as<br />
D AB (3/4)ln[1 4/3(Su/I Su)][1 G/T] G/T.<br />
D AB is <strong>the</strong> distance between sequences A and B, I <strong>the</strong> number<br />
<strong>of</strong> identical nucleotides, Su <strong>the</strong> number <strong>of</strong> positions<br />
showing a substitution, G <strong>the</strong> number <strong>of</strong> gaps in one sequence<br />
with respect to <strong>the</strong> o<strong>the</strong>r, and T <strong>the</strong> sum <strong>of</strong> I, S and<br />
G. Insertions and deletions were taken into account. Tree<br />
topology was inferred by <strong>the</strong> Neighbor-Joining (NJ) method<br />
(Saitou and Nei 1987). The Bootstrap method (Felsenstein<br />
1985) was performed with 1000 replications to evaluate <strong>the</strong><br />
reliability <strong>of</strong> tree topologies. Distance analysis and tree<br />
drawing were carried out with TreeCon (Van de Peer and<br />
De Wachter 1993).<br />
Maximum parsimony (MP) trees were inferred with <strong>the</strong><br />
heuristic method with <strong>the</strong> aid <strong>of</strong> PAUP 3.1.1 (Sw<strong>of</strong>ford<br />
1993). Validity <strong>of</strong> <strong>the</strong> clades was tested by bootstrap analysis<br />
(Felsenstein 1985). The analysis was conducted with 1000<br />
bootstrap replications, retaining groups compatible with <strong>the</strong><br />
50% majority-rule in <strong>the</strong> bootstrap consensus tree, with <strong>the</strong><br />
Nearest-Neighbor Interchange (NNI) branch-swapping option,<br />
and no more than 600 trees (700 length) saved each<br />
replication. The 50% majority rule consensus tree is available<br />
on line at TreeBASE (SN908). Parsimony trees were<br />
drawn with <strong>the</strong> aid <strong>of</strong> TreeView (Page 1996).<br />
DíEZ ET AL: DESERT TRUFFLES PHYLOGENY<br />
RESULTS<br />
249<br />
RFLP pr<strong>of</strong>iles <strong>of</strong> amplified ITS. The ITS region was<br />
successfully amplified for 34 collections. In contrast,<br />
four old herbarium collections (nos. 6, 21, 26, and<br />
27) did not amplify, probably as a result <strong>of</strong> DNA degradation.<br />
Each species produced a characteristic<br />
RFLP pattern (A to G, TABLE IV), except for Terfezia<br />
arenaria and T. leptoderma. Terfezia arenaria was polymorphic<br />
for AluI and produced two RFLP pr<strong>of</strong>iles.<br />
Terfezia leptoderma was polymorphic for Hinf I and<br />
RsaI. Three sequences <strong>of</strong> T. boudieri retrieved from<br />
GenBank (AF092096–AF092098; strain type-1, type-2<br />
and type-3; Western Negev, Israel) gave predicted<br />
RFLP patterns for this species.<br />
Sequence comparisons. Search for similar sequences<br />
in <strong>the</strong> GenBank DNA database produced significant<br />
alignments with <strong>the</strong> ITS sequences <strong>of</strong> Terfezia boudieri<br />
(AF092096–AF092098) and Mattirolomyces terfezioides<br />
(AJ272442–AJ272445). The Genbank sequences <strong>of</strong><br />
M. terfezioides showed 99% similarity with <strong>the</strong> sequence<br />
AF276681 <strong>of</strong> Mattirolomyces terfezioides (collection<br />
38, present study). Surprisingly, a search for<br />
sequence similarity using <strong>the</strong> BLASTn algorithm did<br />
not produce significant alignments with <strong>the</strong> Peziza/<br />
Plicaria sequences reported by Norman and Egger<br />
(1999). In addition, <strong>the</strong> ITS sequences <strong>of</strong> P. badia<br />
and P. griseo-rosea (U40475 and U4047) were shorter<br />
and very different from those <strong>of</strong> Terfezia and Tirmania.<br />
This finding was unexpected, because <strong>the</strong>y have<br />
been suggested to be closely related species to Terfezia<br />
arenaria (Norman and Egger 1999). Fur<strong>the</strong>rmore,<br />
no unambiguous alignment was achieved after<br />
<strong>the</strong> addition <strong>of</strong> <strong>the</strong>se sequences <strong>of</strong> Peziza to <strong>the</strong> set<br />
<strong>of</strong> Terfezia and Tirmania ITS sequences.<br />
The variations in <strong>the</strong> length <strong>of</strong> <strong>the</strong> ITS sequences<br />
were <strong>of</strong>ten attributable to deletions and insertions.<br />
Gaps were <strong>the</strong>refore introduced in order to align <strong>the</strong><br />
sequences. The total length <strong>of</strong> <strong>the</strong> alignment was 660<br />
positions. They comprised a small portion <strong>of</strong> <strong>the</strong><br />
flanking 18S and 28S rDNA genes (11 and 36 bp<br />
respectively), <strong>the</strong> ITS1 region (nucleotides 12–247),<br />
<strong>the</strong> 5.8S rDNA (nucleotides 248–403), and <strong>the</strong> ITS2<br />
sequence (nucleotides 404–624). Sequence variability<br />
was most prominent within <strong>the</strong> ITS regions, which<br />
had several indels (e.g., all Terfezia and Tirmania species<br />
contained a 6-bp deletion between <strong>the</strong> nucleotide<br />
positions 33 and 39). O<strong>the</strong>r deletions were specific<br />
to a set <strong>of</strong> species or just one species; e.g., <strong>the</strong><br />
ITS1 sequence <strong>of</strong> Terfezia arenaria had a large deletion<br />
between positions 101–114 in <strong>the</strong> consensus.<br />
Phylogenetic inferred trees. The phylogenetic trees inferred<br />
by both distance-based (FIG. 2) and cladistic<br />
methods (FIG. 3) showed <strong>the</strong> same topology, in spite
250 MYCOLOGIA<br />
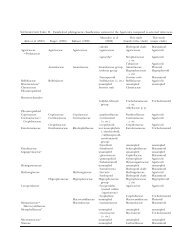
TABLE I. Collections <strong>of</strong> <strong>desert</strong> <strong>truffles</strong> used in <strong>the</strong> present work<br />
Collection<br />
no. a Origin Habitat b Host<br />
Spore<br />
morphology Phenetic species<br />
1* Kuwait Basic soil, AR Helian<strong>the</strong>mum salici- Smooth Tirmania nivea<br />
folium (L.) Mill.<br />
Chatin<br />
2** Spain, <strong>the</strong> Tabernas Sandy basic soil,<br />
— Smooth T. nivea (Des. : Fr)<br />
Desert, Almeria AR<br />
Trappe<br />
3** Tunisia, market <strong>of</strong><br />
Tunis<br />
— — Smooth T. nivea<br />
4* Tunisia, market <strong>of</strong><br />
Tunis<br />
— — Smooth T. nivea<br />
5* Algeria Acid soil, AR H. guttatum (L.) Mill. Subsmooth T. pinoyi (Maire)<br />
Malençon<br />
6** Morocco Acid soil, AR H. guttatum Subsmooth T. pinoyi<br />
7* Spain, Navalmoral, Sandy acid soils, H. guttatum Warty Terfezia arenaria<br />
Cáceres<br />
SAR<br />
(Moris) Trappe<br />
8* Spain, Navalmoral, Sandy acid soils, H. guttatum Warty Terfezia arenaria<br />
Cáceres<br />
SAR<br />
(Moris) Trappe<br />
9** Spain, Navalmoral, Sandy acid soils, H. guttatum Warty Terfezia arenaria<br />
Cáceres<br />
SAR<br />
(Moris) Trappe<br />
10* Spain, Campo Ar- Sandy acid soil, H. guttatum Warty T. arenaria<br />
añuelo, Cáceres SAR<br />
11** Spain, Campo Ar- Sandy acid soil, H. guttatum Warty T. arenaria<br />
añuelo, Cáceres SAR<br />
12* Spain, Albuquerque,<br />
Cáceres<br />
Acid soil, SAR H. guttatum Warty T. arenaria<br />
13* Morocco, Mamora Acid soil, SAR H. macrosepalum (L.)<br />
Mill.<br />
Warty T. arenaria<br />
14* Morocco, Mamora,<br />
bought in a local<br />
market<br />
— — Warty T. arenaria<br />
15** Morocco, Mamora,<br />
bought in a local<br />
market<br />
— — Warty T. arenaria<br />
16* Portugal Acid soils, SAR H. guttatum Warty T. arenaria<br />
17** Algeria Basic soil, SAR H. salicifolium Reticulate<br />
and warty<br />
T. boudieri Chatin<br />
18* Kuwait Basic soil, SAR H. salicifolium Reticulate<br />
and warty<br />
T. boudieri<br />
19* Spain, Fuentidueña, Gypsiferous H. squamatum (L.) Reticulate T. boudieri<br />
Madrid<br />
shrubland,<br />
SAR<br />
Dum. Cours.<br />
and warty<br />
20** Spain, Valdeguerra, Calcareous soil, H. salicifolium Reticulate T. boudieri<br />
Madrid<br />
SAR<br />
and Warty<br />
21* Spain, Cabezamesa- Basic soil, SAR H. ledifolium and H. Reticulate T. boudieri<br />
da, Toledo<br />
salicifolium<br />
and warty<br />
22** Spain, The Toledo Basic soil, SAR H. salicifolium Reticulate T. boudieri<br />
Mountains<br />
and warty<br />
23* Morocco, market <strong>of</strong><br />
— — Reticulate T. boudieri<br />
Tangier<br />
and warty<br />
24** Morocco, market <strong>of</strong><br />
— — Reticulate T. boudieri<br />
Casablanca<br />
and warty<br />
25** Spain, Torres de la Calcareous soils Helian<strong>the</strong>mum spp. Reticulate T. claveryi Chatin<br />
Alameda, Madrid SAR<br />
basphilous<br />
26** Spain, Guadix, Gra- Basic soil, SAR Helian<strong>the</strong>mum spp. Reticulate T. claveryi<br />
nada<br />
basophilous<br />
27** Spain, Llanos de Basic soil, SAR Helian<strong>the</strong>mum spp. Reticulate T. claveryi<br />
Armilla, Granada<br />
basophilous
TABLE I. Continued<br />
DíEZ ET AL: DESERT TRUFFLES PHYLOGENY<br />
Collection<br />
no. a Origin Habitat b Host<br />
Spore<br />
morphology Phenetic species<br />
28* Spain, Navalmoral, Sandy acid soil, H. guttatum Spiny T. leptoderma Tul. &<br />
Cáceres<br />
SAR<br />
C. Tul.<br />
29* Spain, Navalmoral, Sandy acid soil, H. guttatum Spiny T. leptoderma<br />
Cáceres<br />
SAR<br />
30** Spain, Campo Ar- Sandy acid soil, H. guttatum Spiny T. leptoderma<br />
añuelo, Cáceres SAR<br />
31** Spain, Campo Ar- Sandy acid soil, H. guttatum Spiny T. leptoderma<br />
añuelo, Cáceres SAR<br />
32** Spain, Cáceres Sandy acid soil,<br />
SAR<br />
H. guttatum Spiny T. leptoderma<br />
33** Spain, Trujillo, Cá- Sandy acid soil, H. guttatum Spiny T. leptoderma<br />
ceres<br />
SAR<br />
34** Spain, Cañaveral de Clayey slate de- Cistus ladanifer L. Spiny T. leptoderma<br />
León, Huelva rived soil, SAR<br />
35** Spain, Valencia Siliceous soil,<br />
SAR<br />
Pinus halepensis Mill. Spinyc T. leptoderma<br />
36** France, Auge-Fourvieille<br />
Clayey soil, SAR Quercus ilex L. Spinyc T. leptoderma<br />
37** Hungary, Csomad Forest plantation Robinia pseudoacacia Reticulate Mattirolomyces terfezioides<br />
(Matt.) Fischer<br />
38** Hungary, Surany Garden Ribes rubrum Reticulate M. terfezioides<br />
a DNA was extracted from mycelium (*) or fruit body (**).<br />
b Habitat: AR arid regions; SAR semi-arid regions.<br />
c Slightly smaller spore having shorter spines.<br />
TABLE II. Pezizalean <strong>desert</strong> <strong>truffles</strong> that naturally occur in <strong>the</strong> Mediterranean Region. Diagnostic morphological features,<br />
hosts and soil types<br />
Morphological<br />
species<br />
Main ascocarp<br />
features<br />
T. nivea Ascocarp <strong>of</strong>f-white<br />
when young<br />
T. pinoyi Ascocarp white-yellowish<br />
when young<br />
T. arenaria Gleba with pinkish<br />
tones at maturity<br />
T. boudieri Thick peridium gleba<br />
red tones at maturity<br />
T. claveryi Gleba with red tones<br />
at maturity<br />
T. leptoderma Thin peridium, gleba<br />
with olivaceous<br />
tones at maturity<br />
Melzer’s reaction<br />
<strong>of</strong> asci<br />
Spore ornamentation<br />
and size<br />
Amyloid Smooth oval to ellipsoidal,<br />
16–18 12–14<br />
m<br />
Amyloid Subsmooth (minute<br />
warts) spherical, 15–<br />
20 m<br />
Non amyloid Warty spherical, 20–26<br />
m<br />
Non amyloid Reticulate-warty spherical,<br />
18–22 m<br />
Non amyloid Reticulate spherical, 18–<br />
22 m<br />
Non amyloid Spiny spherical, 17–25<br />
m a<br />
a Inmature ascocarps present slightly smaller spores (15–22 m) and shorter spines.<br />
b Habitat: AR arid regions, SAR semi-arid regions.<br />
Soil type, habitat b<br />
and host<br />
251<br />
Basic soils, AR, basophilousHelian-<br />
<strong>the</strong>mum spp.<br />
Acid soils, AR H.<br />
guttatum<br />
Acid soils, SAR H.<br />
guttatum<br />
Basic soils, SAR basophilus<br />
spp. <strong>of</strong><br />
Helian<strong>the</strong>mum<br />
Basic soils, SAR acidophilous<br />
spp. <strong>of</strong><br />
Helian<strong>the</strong>mum<br />
Acid soils, SAR H.<br />
guttatum and Cistus<br />
ladanifer,<br />
Quercus ilex, Pinus<br />
halepensis
252 MYCOLOGIA<br />
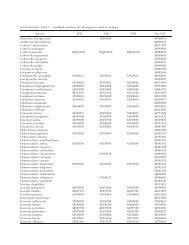
TABLE III. Specimens used for sequencing<br />
Phenetic species a<br />
Tirmania nivea (03)<br />
T. pinoyi<br />
Terfezia claveryi<br />
(02)<br />
(04)<br />
(01)<br />
(05)<br />
(25)<br />
(26)<br />
T. boudieri (18)<br />
(17)<br />
T. arenaria (10)<br />
(11)<br />
T. leptoderma (30)<br />
(31)<br />
(34)<br />
(35)<br />
(36)<br />
Mattirolomyces terfezioides (37)<br />
(38)<br />
ITS-RFLP<br />
patterns<br />
A<br />
A<br />
A<br />
A<br />
B<br />
C<br />
C<br />
D<br />
D<br />
E1<br />
E2<br />
F1<br />
F1<br />
F2<br />
F3<br />
F4<br />
G<br />
G<br />
Sample<br />
no. b Origin Soil type Host<br />
tun08*<br />
tab04**<br />
tun06*<br />
niv05*<br />
pin01*<br />
cly06**<br />
alc03**<br />
bou04*<br />
mot08**<br />
are19*<br />
are20**<br />
lpt12*<br />
lpt08**<br />
jar02**<br />
val06**<br />
enc07**<br />
rob01**<br />
rib02**<br />
Tunisia<br />
Spain<br />
Tunisia<br />
Kuwait<br />
Algeria<br />
Morocco<br />
Spain<br />
Kuwait<br />
Algeria<br />
Spain<br />
Spain<br />
Spain<br />
Spain<br />
Spain<br />
Spain<br />
France<br />
Hungary<br />
Hungary<br />
—<br />
Basic<br />
—<br />
—<br />
—<br />
Basic<br />
Basic<br />
Basic<br />
Basic<br />
Acid<br />
a In brackets collection no. according to TABLE I.<br />
b The ITS regions were amplified from mycelium (*) or fruit bodies (**).<br />
<strong>of</strong> slight differences in branch stability among equivalent<br />
branches. Trees branched into two main clades,<br />
which were well supported by bootstrap values: <strong>the</strong><br />
Mattirolomyces clade (100% NJ and MP) and <strong>the</strong> Tirmania/Terfezia<br />
clade (99% NJ and MP). Terminal<br />
clades <strong>of</strong> <strong>the</strong> inferred trees corresponded exactly to<br />
morphological species (FIG. 2) and were related to<br />
ecological features and hosts (FIG. 3).<br />
The Tirmania clade comprised <strong>desert</strong> <strong>truffles</strong> with<br />
smooth spores and amyloid asci, which were found<br />
in <strong>desert</strong>s. The Tirmania nivea terminal clade (100%<br />
NJ and MP) was composed <strong>of</strong> collections with oval to<br />
ellipsoidal spores, found under basophilous species<br />
<strong>of</strong> Helian<strong>the</strong>mum (e.g., H. salicifolium). Tirmania pinoyi,<br />
which has spherical spores and occurs under <strong>the</strong><br />
acidophilous plant H. guttatum, was a sister species<br />
<strong>of</strong> T. nivea.<br />
The Terfezia clade (97% NJ, 85% MP) grouped species<br />
found in semi-arid habitats and with ornamented<br />
and spherical spores. The first subclade comprises<br />
well-supported terminal clades corresponding to Terfezia<br />
spp. However, relationships between <strong>the</strong>m were<br />
not well resolved. Terfezia arenaria, which displays<br />
warty spores and occurs in sandy acid soils with H.<br />
guttatum (FIG. 1), was monophyletic and <strong>the</strong> corresponding<br />
terminal clade was well supported (100%<br />
NJ and MP). Terfezia boudieri and T. claveryi, which<br />
occur in basic soils under basophilous Helian<strong>the</strong>mum<br />
and possess reticulated and warty-reticulated spores,<br />
respectively, were similarly monophyletic (100% NJ<br />
Acid<br />
Acid<br />
Acid<br />
Acid<br />
Basic<br />
Basic<br />
—<br />
—<br />
Helian<strong>the</strong>mum salicifolium<br />
—<br />
—<br />
—<br />
H. guttatum<br />
H. ledifolium<br />
H. salicifolium<br />
H. salicifolium<br />
—<br />
H. guttatum<br />
H. guttatum<br />
H. guttatum<br />
H. guttatum<br />
Cistus ladanifer<br />
Pinus halepensis<br />
Quercus ilex<br />
Robinia<br />
—<br />
Genbank<br />
acc. no.<br />
AF276665<br />
AF276666<br />
AF276667<br />
AF276668<br />
AF276669<br />
AF276670<br />
AF276671<br />
AF276672<br />
AF276673<br />
AF276674<br />
AF276675<br />
AF276678<br />
AF276679<br />
AF396862<br />
AF396863<br />
AF396863<br />
AF276680<br />
AF276681<br />
and MP). The second subclade includes specimens<br />
with spherical spiny spores belonging to T. leptoderma<br />
(89% NJ, 95% MP). Terfezia leptoderma clade comprised<br />
specimens associated with several different<br />
hosts (Helian<strong>the</strong>mum guttatum, Cistus ladanider,<br />
Quercus ilex and Pinus halepensis), and were collected<br />
in different type <strong>of</strong> soils (ei<strong>the</strong>r basic or acid soils).<br />
Thus, <strong>the</strong> molecular phylogenetic analysis indicated<br />
that morphological species represent phylogenetic<br />
species.<br />
DISCUSSION<br />
Previous studies have noted that <strong>desert</strong> <strong>truffles</strong> are<br />
sometimes difficult to distinguish on <strong>the</strong> basis <strong>of</strong> <strong>the</strong>ir<br />
morphology (Moreno et al 2000, 2001). In <strong>the</strong> present<br />
work, <strong>the</strong> RFLP pr<strong>of</strong>iles <strong>of</strong> <strong>the</strong> nuclear rDNA ITS<br />
sequences were consistent with species delimitation<br />
based on known morphological characters. In addition,<br />
ITS sequences were nearby homogenous within<br />
species (e.g., 0.3% nucleotide divergence within Terfezia<br />
arenaria) to polymorphic (up to 7% within T.<br />
leptoderma), and clearly polymorphic at <strong>the</strong> interspecific<br />
level (8%).<br />
The general morphological characteristics <strong>of</strong> Tirmania<br />
and Terfezia are described in Trappe (1979),<br />
and Alsheikh and Trappe (1983a) authored a monograph<br />
<strong>of</strong> Tirmania. Both genera are characterized by<br />
globose to turbinate ascocarps, with solid glebae<br />
formed <strong>of</strong> fertile pockets separated by pale sterile tra-
TABLE IV. Restriction size polymorphism in <strong>desert</strong> <strong>truffles</strong> (Tirmania and Terfezia spp.) and Mattirolomyces terfezioides. Size <strong>of</strong> <strong>the</strong> uncut ITS and restriction fragments<br />
in base pairs (bp)<br />
RFLP<br />
patterns<br />
Size <strong>of</strong> RFLP fragments<br />
AluI HhaI HinfI b RsaI<br />
Uncut<br />
ITS<br />
Collection nos. a Phenetic species<br />
1, 2, 3, 4 Tirmania nivea 620 390, 230 350, 270 210, 170, 120, 100 620 type A<br />
5, 6 T. pinoyi 630 380, 250 340, 290 200, 140, 130, 100, 40 480, 150 type B<br />
24, 25, 26, 27 Terfezia claveryi 630 380, 250 340, 290 300, 310 490, 90, 50 type C<br />
17, 18, 19, 20, 21, 22,<br />
23 T. boudieri 640 390, 250 350, 290 310, 230, 80 500, 90, 50 type D<br />
7, 8, 10, 11, 12, 13 T. arenaria (are19) 640 370, 270 330, 310 300, 320 490, 90, 60 type E1<br />
9, 11, 12, 13, 14, 15, 16 (are20) 640 640 330, 310 300, 320 490, 90, 60 type E2<br />
28, 29, 30, 31, 32, 33 T. leptoderma (lpt08) (lpt12) 620 390, 230 350, 270 310, 290 530, 90 type F1<br />
34 (jar02) 630 380, 250 340, 290 300, 310 400, 140, 90 type F2<br />
35 (val06) 640 390, 250 350, 290 310, 310 400, 140, 90 type F3<br />
36 (enc07) 620 380, 240 340, 280 300, 190, 110 530, 90 type F4<br />
37, 38 Mattirolomyces terfezioides 670 390, 280 370, 300 330, 140, 110, 80 580, 90 type G<br />
DíEZ ET AL: DESERT TRUFFLES PHYLOGENY<br />
a See Table I for specimen data.<br />
b This enzyme also produced one or two 20 bp fragments.<br />
253<br />
mal veins. They have saccate to globose asci with up<br />
to eight clustered spores. Based on <strong>the</strong>se morphological<br />
data, similar habitat and associated hosts, <strong>the</strong>y<br />
have been considered as closely related genera. Our<br />
molecular analyses confirmed <strong>the</strong>ir close relationship.<br />
Moreover, <strong>the</strong> phylogenetic analyses indicate<br />
that both genera are monophyletic. Such a monophyly<br />
is in agreement with morphological data. Terfezia<br />
has non-amyloid asci and ornamented spores. In<br />
contrast, Tirmania has amyloid asci and smooth<br />
spores, both <strong>of</strong> which are features regarded as diagnostic<br />
characters at different taxonomic ranks.<br />
Trappe (1971) used Melzer’s reagent to distinguish<br />
Tirmania from Terfezia. Later, Trappe (1979) used<br />
this character to transfer Tirmania from <strong>the</strong> Terfeziaceae<br />
(non-amyloid asci) to <strong>the</strong> Pezizaceae (amyloid<br />
asci). Amyloid reaction seems to be diagnostic at <strong>the</strong><br />
genus level. However, <strong>the</strong> strong statistical support<br />
for <strong>the</strong> Terfezia/Tirmania molecular clade (present<br />
work), toge<strong>the</strong>r with <strong>the</strong> lack <strong>of</strong> amyloid asci in Matteriolomyces,<br />
suggest that this biochemical reaction is<br />
<strong>of</strong> limited value as diagnostic character at <strong>the</strong> family<br />
level.<br />
To reconstruct <strong>the</strong> molecular <strong>phylogeny</strong> <strong>of</strong> <strong>the</strong>se<br />
fungi, we included <strong>the</strong> related hypogeous fungus<br />
Mattirolomyces terfezioides Fischer (Percudani et al<br />
1999). This fungus does not occur in <strong>the</strong> same habitats<br />
as Terfezia and Tirmania, but in temperate forests.<br />
This species seems to be specifically associated<br />
with Robinia pseudoacacia (Montecchi and Lazzari<br />
1993, Bratek et al 1996). It was probably co-introduced<br />
to Europe with Robinia pseudoacacia. The collection<br />
<strong>of</strong> M. terfezioides under Ribes rubrum in a garden<br />
in Hungary (collection no. 38, TABLE I) does not<br />
provide reliable information on its <strong>mycorrhizal</strong> status,<br />
because Robinia pseudoacacia is a common garden<br />
tree in Hungary. Mattirolomyces terfezioides was<br />
described by Fischer (1938) and <strong>the</strong>n assigned to <strong>the</strong><br />
genus Terfezia by Trappe (1971). Later, based on <strong>the</strong><br />
analysis <strong>of</strong> <strong>the</strong> 18S rDNA sequence, Percudani (1999)<br />
suggested <strong>the</strong> re-establishment <strong>of</strong> <strong>the</strong> genus Mattirolomyces.<br />
The lack <strong>of</strong> nesting <strong>of</strong> Mattirolomyces terfezioides<br />
in <strong>the</strong> Terfezia clade (FIGS. 2 and 3) and <strong>the</strong><br />
study <strong>of</strong> Norman and Egger (1999) support this reestablishment.<br />
According to molecular data (O’Donnell et al<br />
1997, Norman and Egger 1999, Percudani et al<br />
1999), Terfezia and o<strong>the</strong>r pezizalean ascomycetes<br />
such as Pachyphloeus, Mattirolomyces, and Cazia, may<br />
have evolved from ancestral epigeous pezizas towards<br />
a hypogeous habit. Some species <strong>of</strong> Pachyphloeus and<br />
Mattirolomyces combine a hypogeous ascoma with biseriate<br />
spores. This combination might represent an<br />
intermediate step between <strong>the</strong> uniseriate asci <strong>of</strong> epigeous<br />
Pezizaceae and globose asci <strong>of</strong> <strong>the</strong> genera Ter-
254 MYCOLOGIA<br />
FIG. 1. A common <strong>desert</strong> truffle in sandy acid soils <strong>of</strong> <strong>the</strong> semiarid regions in <strong>the</strong> Iberian Peninsula (Terfezia arenaria)<br />
and its <strong>mycorrhizal</strong> host plant Helian<strong>the</strong>mum guttatum.<br />
fezia and Tirmania. Analysis <strong>of</strong> <strong>the</strong> ITS sequences<br />
showed <strong>the</strong> separation <strong>of</strong> <strong>the</strong> Mattirolomyces from <strong>the</strong><br />
Terfezia-Tirmania clade. They seem to represent two<br />
different evolutionary pezizalean lineages. This hypo<strong>the</strong>sis<br />
is in agreement with <strong>the</strong> analyses <strong>of</strong> <strong>the</strong> 18S<br />
rDNA sequences by Norman and Egger (1999) and<br />
Percudani et al (1999), who found that Terfezia arenaria<br />
and Mattirolomyces terfezioides are not sister<br />
genera. Indeed, Pachyphloeus and Mattirolomyces species<br />
occur in temperate forests with cold winters and<br />
are associated with North Temperate trees (Trappe<br />
1979). In addition, M. terfezioides and Pachyphloeus<br />
melanoxanthus were sister species in Percudani<br />
(1999). They perhaps evolved hypogeous ascocarps<br />
to protect fruitbody development from frost, and rely<br />
on animals for <strong>the</strong>ir spore dispersal. This might be<br />
<strong>the</strong> case with most common forest-dwelling <strong>truffles</strong><br />
fruiting in winter in Central Europe (e.g., Tuber melanosporum).<br />
In contrast, Terfezia and Tirmania occur<br />
in arid and semi-arid ecosystems in <strong>the</strong> Mediterranean<br />
region, associated with dwarf shrubs and herbaceous<br />
plants (Helian<strong>the</strong>mum spp.). The <strong>desert</strong> <strong>truffles</strong><br />
fruit are vernal fungi. The Terfezia-Tirmana evolutionary<br />
lineages have probably evolved towards hypogeous<br />
fruitbodies as protection from <strong>the</strong> heat and<br />
drought <strong>of</strong> late spring in semi-arid and arid habitats.<br />
The evolution <strong>of</strong> epigeous fruitbodies towards hypogeous<br />
habits seems to have happened in several lineages<br />
<strong>of</strong> ecto<strong>mycorrhizal</strong> fungi; selection for reduction<br />
<strong>of</strong> water loss has been proposed to explain <strong>the</strong><br />
accelerated evolution <strong>of</strong> suilloid basidiocaps towards<br />
false <strong>truffles</strong> (e.g., Rhizopogon) through secotioid<br />
forms (Bruns et al 1989).<br />
Relationships between Peziza/Plicaria and Terfezia
DíEZ ET AL: DESERT TRUFFLES PHYLOGENY<br />
FIG. 2. Neighbor-joining tree <strong>of</strong> 30 ITS/5.8S rDNA sequences <strong>of</strong> pezizalean <strong>truffles</strong> constructed with <strong>the</strong> Jukes and<br />
Cantor’s one-parameter distance method. Numbers in branches are <strong>the</strong> bootstrap values as percentage bootstrap replication<br />
from a 1000 replicate analysis. Scale represents <strong>the</strong> distance between isolates. Outlines delimit clusters, which correspond to<br />
morphological species. Square brackets delimit genera. Major diagnostic morphological characters are indicated.<br />
255
256 MYCOLOGIA<br />
FIG. 3. Rooted 50% majority rule consensus tree resulting from 1000 bootstrap replications <strong>of</strong> <strong>the</strong> parsimony analysis <strong>of</strong><br />
<strong>the</strong> ITS/5.8 rDNA sequences (607 steps, consistency index, CI 0.81; retention index, RI 0.86; rescaled consistency index,<br />
RC 0.70; homoplasy index, HI 0.19). Analysis was conducted using <strong>the</strong> heuristic search algorithm <strong>of</strong> PAUP 3.1.1.<br />
(Sw<strong>of</strong>ford 1993). Numbers on <strong>the</strong> branches are <strong>the</strong> bootstrap values (%). Scale represents steps. Outlines indicate <strong>the</strong><br />
different clades observed, which coincide with <strong>the</strong> different habitats (e.g., temperate forests, <strong>desert</strong>s and semi-arid areas).<br />
Habitat, soil types and hosts are indicated.
have been recently studied by 18S rDNA sequence<br />
analysis (Norman and Egger 1999, Percudani et al<br />
1999). Whereas Percudani et al (1999) indicated that<br />
P. badia is not a sister species <strong>of</strong> Terfezia, Norman and<br />
Egger (1999) reported Peziza badia and P. griseo-rosea<br />
as closely related to Terfezia. In <strong>the</strong> present study, Terfezia,<br />
Matteriolomyces, and Tirmania ITS did not produce<br />
significant sequence similarities with ITS <strong>of</strong> Peziza<br />
deposited in GenBank. This result is in agreement<br />
with Percudani et al (1999). The ITS sequences<br />
<strong>of</strong> P. badia and P. griseo-rosea (U40475 and U4047)<br />
are shorter and very different from those <strong>of</strong> Terfezia<br />
and Tirmania. In <strong>the</strong> present study, we wanted to verify<br />
whe<strong>the</strong>r <strong>the</strong> Tirmania/Terfezia clade was still<br />
monophyletic after introduction <strong>of</strong> Peziza species in<br />
<strong>the</strong> phylogenetic analyses. The lack <strong>of</strong> satisfactory<br />
alignments among ITS sequences <strong>of</strong> Tirmania/Terfezia<br />
and Peziza supports <strong>the</strong> divergence <strong>of</strong> <strong>the</strong>se lineages<br />
and <strong>the</strong> monophyly <strong>of</strong> <strong>the</strong> Terfezia/Tirmania<br />
clade.<br />
The Tirmania-Terfezia lineage presented two wellsupported<br />
branching clades, one with smooth spores<br />
and amyloid asci (Tirmania spp.) and <strong>the</strong> o<strong>the</strong>r with<br />
ornamented spores and nonamyloid asci (Terfezia<br />
spp.). Species with smooth spores occur in <strong>desert</strong> areas<br />
<strong>of</strong> <strong>the</strong> Mediterranean region (Alsheikh and<br />
Trappe 1983b). In <strong>the</strong> Western Mediterranean Basin,<br />
its nor<strong>the</strong>rnmost populations seem to be in <strong>the</strong> Tabernas<br />
Desert in Sou<strong>the</strong>rn Spain (Moreno et al<br />
2000). In contrast, Terfezia species (ornamented<br />
spores) occur in semiarid areas and thus have a more<br />
nor<strong>the</strong>rly distribution; i.e., <strong>the</strong>y are well represented<br />
all over <strong>the</strong> Mediterranean area <strong>of</strong> <strong>the</strong> Iberian Peninsula<br />
(Alvarez et al 1993, Moreno et al 2001). Although<br />
regarded as hypogeous fungi, <strong>the</strong> <strong>desert</strong> <strong>truffles</strong><br />
eventually emerge above ground, resulting in a<br />
semi-hypogeous habit. Fur<strong>the</strong>rmore, it has been<br />
claimed that <strong>the</strong>ir ascocarps <strong>the</strong>n dry in situ from <strong>the</strong><br />
heat and drought <strong>of</strong> late spring (Trappe 1992). Trappe<br />
(1992) fur<strong>the</strong>r suggested that in Tirmania <strong>the</strong> peridium<br />
collapses when dried and spores are <strong>the</strong>n disseminated<br />
by dry winds. We have observed fruitbodies<br />
<strong>of</strong> Terfezia bitten by rodents, suggesting a potential<br />
role <strong>of</strong> animals in spore dissemination.<br />
Because today’s species <strong>of</strong> <strong>desert</strong> <strong>truffles</strong> seem to<br />
represent different species adapted to different types<br />
<strong>of</strong> soils, differences in <strong>the</strong> edaphic tolerance may account<br />
for species diversity. Terfezia boudieri and T.<br />
claveryi occur in marl-gypsum soils. In contrast, T. arenaria<br />
lives in siliceous sands. Terfezia leptoderma was<br />
found in association with Helian<strong>the</strong>mun guttatum in<br />
acid soil and with Cistus ladanifer in slate-derived<br />
soils. This fungus was also associated with Quercus ilex<br />
and Pinus halepensis in basic soils. We observed some<br />
differences in <strong>the</strong> spore morphology <strong>of</strong> <strong>the</strong> T. lepto-<br />
DíEZ ET AL: DESERT TRUFFLES PHYLOGENY<br />
257<br />
derma. The small spores <strong>of</strong> specimens collected under<br />
pine and Q. ilex would fit those <strong>of</strong> T. olbiensis.<br />
Terfezia olbiensis was described with similar morphology<br />
to T. leptoderma, except for slightly smaller spores<br />
and shorter spines. However, <strong>the</strong>re is a certain consensus<br />
that T. olbiensis is an immature form and a<br />
synonym <strong>of</strong> T. leptoderma (Moreno et al 1986, 2001,<br />
Alvarez et al 1993). The morphological species T. leptoderma<br />
might be ei<strong>the</strong>r a species with wide edaphic<br />
tolerance and host range or a species complex; isolates<br />
occurring in Cistus scrubs and pine and Quercus<br />
sclerophilous woodlands could belong to distinct species<br />
with different host or/and edaphic specialization.<br />
Our sampling in <strong>the</strong>se ecosystems is scanty and<br />
fur<strong>the</strong>r study is necessary.<br />
In <strong>the</strong> Mediterranean region, o<strong>the</strong>r <strong>mycorrhizal</strong><br />
taxa seem to present different edaphic tolerances<br />
and host ranges, i.e., <strong>the</strong> Pisolithus species complex<br />
(Díez et al 2000, 2001). As in <strong>the</strong> case <strong>of</strong> Terfezia<br />
species, ITS analyses suggested <strong>the</strong> presence <strong>of</strong> several<br />
Pisolithus species occurring in different soil types<br />
(basic, acid, and clayey slate-derived soils) and with<br />
specificity for particular indigenous hosts.<br />
In <strong>the</strong> case <strong>of</strong> Tirmania spp., Tirmania nivea collections<br />
analyzed in <strong>the</strong> present work were found in<br />
basic soils (e.g., collection no. 2) and associated with<br />
<strong>the</strong> basophilous plant H. salicifolium (e.g., no. 1). In<br />
contrast, T. pinoyi specimens were collected under<br />
<strong>the</strong> acidophilous plant H. guttatum (e.g., collections<br />
nos. 5–6). Larger surveys are needed to confirm<br />
whe<strong>the</strong>r T. nivea and T. pinoyi also have different<br />
edaphic tolerances and/or host adaptations.<br />
The distribution pattern <strong>of</strong> <strong>the</strong> <strong>desert</strong> <strong>truffles</strong> species<br />
seems to correlate so strongly with host, that <strong>the</strong><br />
two factors (host specialization and soil pH) might<br />
have played a key role in <strong>the</strong>ir speciation. Fur<strong>the</strong>rmore,<br />
most species <strong>of</strong> Helian<strong>the</strong>mum forming mycorrhizas<br />
with <strong>desert</strong> <strong>truffles</strong> in <strong>the</strong> Mediterranean region<br />
show different edaphic tolerances. Whereas H.<br />
salicifolium and H. ledifolium occur in basic soils, o<strong>the</strong>r<br />
species <strong>of</strong> Helian<strong>the</strong>mum occur only in acid soils,<br />
e.g., H. guttatum. Soil features <strong>the</strong>refore have an impact<br />
on <strong>the</strong> distribution <strong>of</strong> <strong>the</strong> host plants. Distribution<br />
<strong>of</strong> <strong>desert</strong> <strong>truffles</strong> species according to soil features<br />
might <strong>the</strong>refore just reflect <strong>the</strong> edaphic tolerance<br />
<strong>of</strong> <strong>the</strong>ir hosts.<br />
The present phylogenetic analyses confirm that<br />
<strong>the</strong>se morphological species are also phylogenetic<br />
species. Distance analysis indicated that <strong>the</strong>y are well<br />
separated. Distance values within and among species<br />
are low, suggesting that <strong>the</strong>se species diverged a short<br />
time ago. Estimation <strong>of</strong> reliable divergence-time rates<br />
for ITS sequences from <strong>mycorrhizal</strong> fungi would be<br />
invaluable in dating this divergence. A larger survey<br />
using multilocus molecular analyses is needed to de-
258 MYCOLOGIA<br />
termine <strong>the</strong> genetic structure <strong>of</strong> Terfezia and Tirmania<br />
populations.<br />
ACKNOWLEDGMENTS<br />
We thank Dr. James M. Trappe (Forest Service, Corvallis,<br />
Oregon), Dr. Thomas D. Bruns (University <strong>of</strong> California,<br />
Berkeley, California), and anonymous reviewers for critical<br />
comments on <strong>the</strong> manuscript. We are grateful to Dr. G.<br />
Chevalier and C. Dupré (INRA, Clermont-Ferrand,<br />
France), L. Khabar (University <strong>of</strong> Rabat, Morocco) and<br />
Romero de la Osa (Forest Service, Junta de Andalucía,<br />
Spain) for providing several strains and specimens. We also<br />
thank Dr. E. Jakucs (Eotvos Lorand University, Budapest,<br />
Hungary) for his comments on Mattirolomyces terfezioides,<br />
and Dr. G. Moreno (Alcalá University) for assistance in <strong>the</strong><br />
taxonomic identification. We are also grateful to INIA (project<br />
SC98–030) and ‘‘Vicerrectorado de Investigación de la<br />
Univ. de Alcalá’’ (EO28/98) for <strong>the</strong>ir financial support.<br />
This work was supported by a postdoctoral fellowship (EU,<br />
Contract HPMF-CT-1999-00174) and a ‘‘Ramón y Cajal’’<br />
contract from <strong>the</strong> MCyT (Spain) to J. Díez.<br />
LITERATURED CITED<br />
Alsheikh M, Trappe JM. 1983a. Desert <strong>truffles</strong>: <strong>the</strong> genus<br />
Tirmania. Trans Br Mycol Soc 81:83–90.<br />
———, ———. 1983b. Taxonomy <strong>of</strong> Phaeangium lefebvrei,<br />
a <strong>desert</strong> truffle eaten by birds. Can J Bot 61:1919–1925.<br />
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z,<br />
Miller W, Lipman DJ. 1997. Gapped BLAST and PSI-<br />
BLAST, a new generation <strong>of</strong> protein database search<br />
programs. Nuc Acids Res 25:3389–3402.<br />
Alvarez IF, Parladé X, Trappe JM, Castellano MA. 1993. Hypogeous<br />
<strong>mycorrhizal</strong> fungi <strong>of</strong> Spain. Mycotaxon 47:<br />
201–217.<br />
Bratek Z, Jakucs E, Boka K, Szedlayu G. 1996. Mycorrhizae<br />
between black locust (Robinia pseudoacacia) and Terfezia<br />
terfezioides. Mycorrhiza 6:271–274.<br />
Bruns TD, Fogel R, White T, Palmer JD. 1989. Accelerated<br />
evolution <strong>of</strong> a false-truffle from a mushroom ancestor.<br />
Nature 339:140–142.<br />
Corpet F. 1988. Multiple sequence alignment with hierarchical<br />
clustering. Nucl. Acids Res 16:10881–10890.<br />
Díez J, Anta B, Manjón JL, Honrubia M. 2001. Genetic variability<br />
<strong>of</strong> Pisolithus isolates associated with native hosts<br />
and exotic eucalyptus in <strong>the</strong> western Mediterranean region.<br />
New Phytol 149:577–588.<br />
———, Manjón JL, Kovacs G, Celestino C, Toribio M. 2000.<br />
Mycorrhization <strong>of</strong> vitroplants raised from somatic embryos<br />
<strong>of</strong> cork oak (Quercus suber L.). Appl Soil Ecol 15:<br />
137–144.<br />
Dexheimer J, Gérard J, Leduc JP, Chevalier G. 1985. Etude<br />
ultrastructurale comparée des associations symbiotiques<br />
mycorhiziennes Helian<strong>the</strong>mum salicifolium–Terfezia<br />
claveryi et Helian<strong>the</strong>mum salicifolium–Terfezia leptoderma.<br />
Can J Bot 63:582–591.<br />
Felsenstein J. 1985. Confidence limits on phylogenies: an<br />
approach using <strong>the</strong> bootstrap. Evolution 39:783–791.<br />
Fischer E. 1938. Die Naturlichen Pflazenfamilien, Engler-<br />
Prantl-Harms, Leipzig. 42 p.<br />
Fortas Z, Chevalier G. 1992. Effet des conditions de culture<br />
sur la mycorhization de l’Helian<strong>the</strong>mum guttatum par<br />
trois espèces de terfez des genres Terfezia et Tirmania<br />
d’Algérie. Can J Bot 70:2453–2460.<br />
Gardes M, White TJ, Fortin JA, Bruns TD, Taylor JW. 1991.<br />
Identification <strong>of</strong> indigenous and introduced symbiotic<br />
fungi in ectomycorrhizae by amplification <strong>of</strong> nuclear<br />
and mitochondrial rDNA. Can J Bot 69:189–190.<br />
Gargas A, Taylor JW. 1995. Phylogeny <strong>of</strong> Discomycetes and<br />
early radiations <strong>of</strong> <strong>the</strong> apo<strong>the</strong>cial Ascomycotina inferred<br />
from SSU rDNA sequence data. Exp Mycol 19:<br />
7–15.<br />
Harrington FA, Pfister DH, Potter D, Donoghue MJ. 1999.<br />
Phylogenetic studies within <strong>the</strong> Pezizales. I. 18S rDNA<br />
sequence data and classification. <strong>Mycologia</strong> 91:41–50.<br />
———, Potter D. 1997. Phylogenetic relationships within<br />
Sarcoscypha based upon nucleotide sequence <strong>of</strong> <strong>the</strong> internal<br />
transcribed spacer <strong>of</strong> nuclear ribosomal DNA.<br />
<strong>Mycologia</strong> 89:258–267.<br />
Henrion B, Le Tacon F, Martin F. 1992. Rapid identification<br />
<strong>of</strong> genetic variation <strong>of</strong> ecto<strong>mycorrhizal</strong> fungi by amplification<br />
<strong>of</strong> ribosomal RNA genes. New Phytol 122:289–<br />
298.<br />
Honrubia M, Cano A, Molina-Niñirola C. 1992. Hypogeous<br />
fungi from Sou<strong>the</strong>rn Spanish semi-arid lands. Persoonia<br />
14:647–653.<br />
Landvik S, Egger KN, Schumacher T. 1997. Towards a subordinal<br />
classification <strong>of</strong> <strong>the</strong> Pezizales Ascomyceta). Phylogenetic<br />
analyses <strong>of</strong> SSU rDNA sequences. Nord J Bot<br />
17:403–418.<br />
Marx DH. 1969. The influence <strong>of</strong> ectotrophic <strong>mycorrhizal</strong><br />
fungi on <strong>the</strong> resistance <strong>of</strong> pine roots to pathogenic infections.<br />
I. Antagonism <strong>of</strong> <strong>mycorrhizal</strong> fungi to root<br />
pathogenic fungi and soil bacteria. Phytopathology 59:<br />
153–163.<br />
Montecchi A, Lazzari G. 1993. Atlante fotografico di funghi<br />
ipogei. Associazione Micologica Bresadola, Centro Studi<br />
Micologici. Vicenza. 490 p.<br />
Moreno G, Díez J, Manjón JL. 2000. Picoa lefebvrei and Tirmania<br />
nivea, two rare hypogeous fungi from Spain. Mycol<br />
Res 104:378–381.<br />
———, ———, ———. 2001. Terfezia boudieri, first records<br />
from Europe <strong>of</strong> a rare vernal hypogeous <strong>mycorrhizal</strong><br />
fungus. Persoonia 17: (In press).<br />
———, Galán R, Montecchi A. 1991. Hypogeous fungi from<br />
peninsular Spain II. Mycotaxon 42:201–238.<br />
———, ———, Ortega A. 1986. Hypogeous fungi from continental<br />
Spain I. Cryptog Mycol 7:201–229.<br />
Norman JE, Egger KN. 1996. Phylogeny <strong>of</strong> <strong>the</strong> genus Plicaria<br />
and its relationship with Peziza inferred from ribosomal<br />
DNA sequence analysis. <strong>Mycologia</strong> 88:986–<br />
995.<br />
———, ———. 1999. <strong>Molecular</strong> <strong>phylogeny</strong> analysis <strong>of</strong> Peziza<br />
and related genera. <strong>Mycologia</strong> 91:820–829.<br />
O’Donnell K, Cigelnik E, Weber NS, Trappe JM. 1997. Phylogenetic<br />
relationships among ascomycetous <strong>truffles</strong><br />
and true and false morels inferred from 18S and 28S<br />
ribosomal DNA sequence analysis. <strong>Mycologia</strong> 89:48–65.
Page RDM. 1996. TREEVIEW: an application to display phylogenetic<br />
trees on personal computers. Comput Appl<br />
Biosci 12:357–358.<br />
Percudani R, Trevisi A, Zambonelli A, Ottonello S. 1999.<br />
<strong>Molecular</strong> <strong>phylogeny</strong> <strong>of</strong> <strong>truffles</strong> Pezizales (Terfeziaceae,<br />
Tuberaceae) derived from nuclear rDNA sequence<br />
analysis. Mol Phylogenet Evol 13:169–180.<br />
Roux C, Sejalon-Delmas N, Martins M, Parguey-Leduc A,<br />
Dargent R, Bécard G. 1999. Phylogenetic relationships<br />
between European and Chinese <strong>truffles</strong> based on parsimony<br />
and distance analysis <strong>of</strong> ITS sequences. FEMS<br />
Microbiol Lett 180:147–155.<br />
Saitou N, Nei M. 1987. The neighbour-joining method: a<br />
new method for reconstructing phylogenetic trees. Mol<br />
Biol Evol 4:406–425.<br />
Spatafora JW. 1995. Ascomal evolution <strong>of</strong> filamentous ascomycetes.<br />
Evidence from molecular data. Can J Bot<br />
73:S811–S815.<br />
Sw<strong>of</strong>ford DL. 1993. PAUP: phylogenetic analysis using parsimony.<br />
Version 3.1.1. Computer Program distributed<br />
DíEZ ET AL: DESERT TRUFFLES PHYLOGENY<br />
259<br />
by <strong>the</strong> Illinois Natural History Survey, Champaign, Illinois,<br />
USA.<br />
Trappe JM. 1971. A synopsis <strong>of</strong> <strong>the</strong> Carbomycetaceae and<br />
Terfeziaceae (Tuberales). Trans Br Mycol Soc 57:85–<br />
92.<br />
———. 1979. The orders, families, and genera <strong>of</strong> hypogeous<br />
Ascomycotina (<strong>truffles</strong> and <strong>the</strong>ir relatives). Mycotaxon<br />
9:297–340.<br />
———. 1992. Use <strong>of</strong> <strong>truffles</strong> and false-<strong>truffles</strong> around <strong>the</strong><br />
world. In: Bencivenga M, Granetti B, eds. Acti del II<br />
Congr Int sul Tartufo. Spoleto, Italy: Comunità Montana<br />
dei Monti Martani e del Serrano. p 19–30.<br />
Van de Peer Y, De Wachter R. 1993. TREECOM: a s<strong>of</strong>tware<br />
package for <strong>the</strong> construction and drawing <strong>of</strong> evolutionary<br />
trees. Comput Appl Biosci 10:177–182.<br />
White TJ, Bruns T, Lee S, Taylor J. 1990. Amplification and<br />
direct sequencing <strong>of</strong> fungal ribosomal RNA genes for<br />
phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ,<br />
White TJ, eds. PCR protocols: a guide to methods and<br />
applications. New York: Academic Press. p 315–322.