Free-energy calculations - Theoretical Biophysics Group
Free-energy calculations - Theoretical Biophysics Group
Free-energy calculations - Theoretical Biophysics Group
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
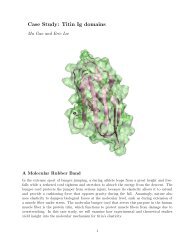
∆ Abinding<br />
A<br />
protein + ligand A<br />
0<br />
∆ Amutation<br />
protein + ligand B<br />
B<br />
∆ Abinding<br />
<strong>Free</strong>-<strong>energy</strong> perturbation <strong>calculations</strong> “Alchemical transformations”<br />
protein ... ligand A<br />
1<br />
∆ Amutation<br />
protein ... ligand B<br />
29 PEARLMAN, CHARIFSON (2001); CHIPOT ET AL. (2005)<br />
In many instances, determining relative binding free energies<br />
for a series of ligands is preferred over a single<br />
binding free <strong>energy</strong>.<br />
This is the case, for example, potential inhibitors of a target<br />
protein are sought in the context of computer–aided<br />
drug design. 29<br />
This can be handled by repeating the “absolute” transformations<br />
for each ligand of interest.<br />
There is an alternate pathway, likely to be more efficient:<br />
Mutation of ligand A into ligand B in both the bound and<br />
the free states, following a different thermodynamic cycle.<br />
Chris Chipot <strong>Free</strong>-<strong>energy</strong> <strong>calculations</strong>