Representing multiple sequence alignments Although most of the time you will represent a multiple sequence alignment as a matrix of aligned residues, in some cases it is c<strong>on</strong>venient to use alternative representati<strong>on</strong>s (as shortcuts or for generati<strong>on</strong> of nice figures). Two of these alternative representati<strong>on</strong>s are the c<strong>on</strong>sensus sequences, and sequence logos. Alternative more complex representati<strong>on</strong>s will be presented in the next secti<strong>on</strong>s (Working with profiles). C<strong>on</strong>sensus sequence C<strong>on</strong>sensus sequence refers to the most comm<strong>on</strong> nucleotide or amino acid at a particular positi<strong>on</strong> after multiple sequences are aligned. A c<strong>on</strong>sensus sequence is a way of representing the results of a multiple sequence alignment, where related sequences are compared to each other, and similar functi<strong>on</strong>al sequence motifs are found. The c<strong>on</strong>sensus sequence shows the residues that are most abundant in the alignment at each positi<strong>on</strong>. Example: WW domain (from SMART database) represented as c<strong>on</strong>sensus sequences c<strong>on</strong>sidering 80, 65 and 50%. O54971/1-33 CONSENSUS/80% CONSENSUS/65% CONSENSUS/50% PLPPGWEKRTDSN-GRVYFVN---HNTRITQWEDPRS .h..sW..hhs.p.sh.aahs.....stpopWptPt. slsssWppthsss.GphYYhs...ppocpopWpcPp. sLPsGWccttsss.G+sYYaN...ppT+copW-cPss The grouping of amino acids to classes and class abbreviati<strong>on</strong> (the key) used within c<strong>on</strong>sensus sequences are shown below. Class Key Residues alcohol o S,T aliphatic l I,L,V any . A,C,D,E,F,G,H,I,K,L,M,N,P,Q,R,S,T,V,W,Y aromatic a F,H,W,Y charged c D,E,H,K,R hydrophobic h A,C,F,G,H,I,K,L,M,R,T,V,W,Y negative - D,E polar p C,D,E,H,K,N,Q,R,S,T positive + H,K,R small s A,C,D,G,N,P,S,T,V tiny u A,G,S turnlike t A,C,D,E,G,H,K,N,Q,R,S,T <strong>Sequence</strong> logos <strong>Sequence</strong> logos are a graphical way for presenting multiple alignments. The sequence logo shows how well residues are c<strong>on</strong>served at each positi<strong>on</strong>: the fewer the number of residues, the higher the letters will be, because the better the c<strong>on</strong>servati<strong>on</strong> is at that positi<strong>on</strong>. Different residues at the same positi<strong>on</strong> are 20
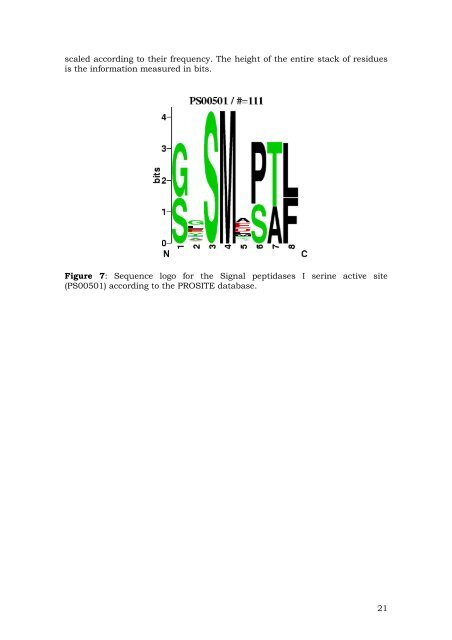
scaled according to their frequency. The height of the entire stack of residues is the informati<strong>on</strong> measured in bits. Figure 7: <strong>Sequence</strong> logo for the Signal peptidases I serine active site (PS00501) according to the PROSITE database. 21