CPGR Workshop â Tutorials Tutorial 1 â Evaluating loci to be ...
CPGR Workshop â Tutorials Tutorial 1 â Evaluating loci to be ...
CPGR Workshop â Tutorials Tutorial 1 â Evaluating loci to be ...
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
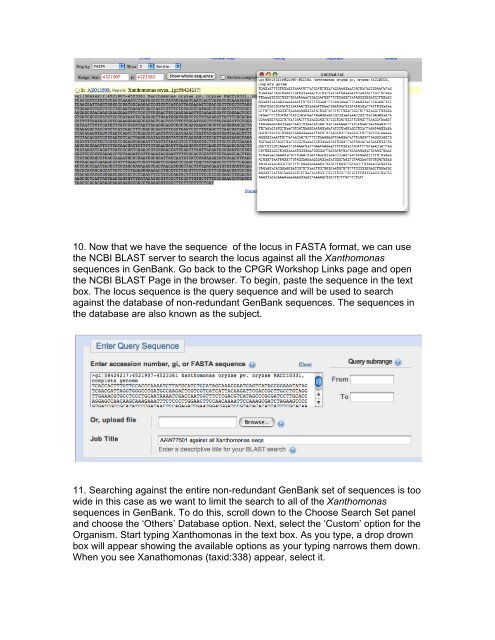
10. Now that we have the sequence of the locus in FASTA format, we can use<br />
the NCBI BLAST server <strong>to</strong> search the locus against all the Xanthomonas<br />
sequences in GenBank. Go back <strong>to</strong> the <strong>CPGR</strong> <strong>Workshop</strong> Links page and open<br />
the NCBI BLAST Page in the browser. To <strong>be</strong>gin, paste the sequence in the text<br />
box. The locus sequence is the query sequence and will <strong>be</strong> used <strong>to</strong> search<br />
against the database of non-redundant GenBank sequences. The sequences in<br />
the database are also known as the subject.<br />
11. Searching against the entire non-redundant GenBank set of sequences is <strong>to</strong>o<br />
wide in this case as we want <strong>to</strong> limit the search <strong>to</strong> all of the Xanthomonas<br />
sequences in GenBank. To do this, scroll down <strong>to</strong> the Choose Search Set panel<br />
and choose the ‘Others’ Database option. Next, select the ‘Cus<strong>to</strong>m’ option for the<br />
Organism. Start typing Xanthomonas in the text box. As you type, a drop drown<br />
box will appear showing the available options as your typing narrows them down.<br />
When you see Xanathomonas (taxid:338) appear, select it.