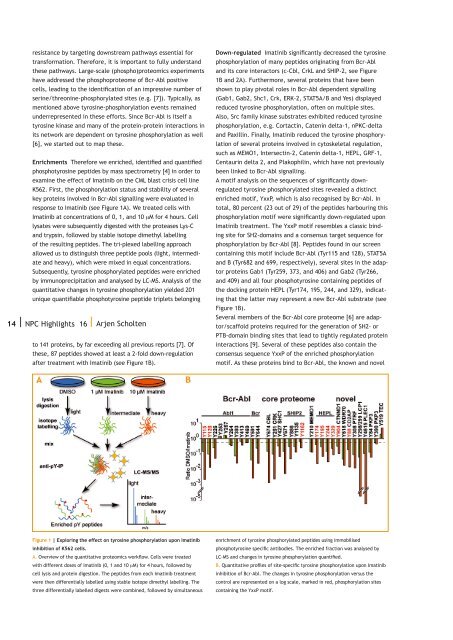
esistance by targeting downstream pathways essential fortransformation. Therefore, it is important to fully understandthese pathways. Large-scale (phospho)proteomics experimentshave addressed the phosphoproteome of Bcr-Abl positivecells, leading to the identification of an impressive number ofserine/threonine-phosphorylated sites (e.g. [7]). Typically, asmentioned above tyrosine-phosphorylation events remainedunderrepresented in these efforts. Since Bcr-Abl is itself atyrosine kinase and many of the protein-protein interactions inits network are dependent on tyrosine phosphorylation as well[6], we started out to map these.Enrichments Therefore we enriched, identified and quantifiedphosphotyrosine peptides by mass spectrometry [4] in order toexamine the effect of Imatinib on the CML blast crisis cell lineK562. First, the phosphorylation status and stability of severalkey proteins involved in Bcr-Abl signalling were evaluated inresponse to Imatinib (see Figure 1A). We treated cells withImatinib at concentrations of 0, 1, and 10 µM for 4 hours. Celllysates were subsequently digested with the proteases Lys-Cand trypsin, followed by stable isotope dimethyl labellingof the resulting peptides. The tri-plexed labelling approachallowed us to distinguish three peptide pools (light, intermediateand heavy), which were mixed in equal concentrations.Subsequently, tyrosine phosphorylated peptides were enrichedby immunoprecipitation and analysed by LC-MS. Analysis of thequantitative changes in tyrosine phosphorylation yielded 201unique quantifiable phosphotyrosine peptide triplets belonging14 | <strong>NPC</strong> Highlights 16 | Arjen Scholtento 141 proteins, by far exceeding all previous reports [7]. Ofthese, 87 peptides showed at least a 2-fold down-regulationafter treatment with Imatinib (see Figure 1B).Down-regulated Imatinib significantly decreased the tyrosinephosphorylation of many peptides originating from Bcr-Abland its core interactors (c-Cbl, CrkL and SHIP-2, see Figure1B and 2A). Furthermore, several proteins that have beenshown to play pivotal roles in Bcr-Abl dependent signalling(Gab1, Gab2, Shc1, Crk, ERK-2, STAT5A/B and Yes) displayedreduced tyrosine phosphorylation, often on multiple sites.Also, Src family kinase substrates exhibited reduced tyrosinephosphorylation, e.g. Cortactin, Catenin delta-1, nPKC-deltaand Paxillin. Finally, Imatinib reduced the tyrosine phosphorylationof several proteins involved in cytoskeletal regulation,such as MEMO1, Intersectin-2, Catenin delta-1, HEPL, GRF-1,Centaurin delta 2, and Plakophilin, which have not previouslybeen linked to Bcr-Abl signalling.A motif analysis on the sequences of significantly downregulatedtyrosine phosphorylated sites revealed a distinctenriched motif, YxxP, which is also recognised by Bcr-Abl. Intotal, 80 percent (23 out of 29) of the peptides harbouring thisphosphorylation motif were significantly down-regulated uponImatinib treatment. The YxxP motif resembles a classic bindingsite for SH2-domains and a consensus target sequence forphosphorylation by Bcr-Abl [8]. Peptides found in our screencontaining this motif include Bcr-Abl (Tyr115 and 128), STAT5Aand B (Tyr682 and 699, respectively), several sites in the adaptorproteins Gab1 (Tyr259, 373, and 406) and Gab2 (Tyr266,and 409) and all four phosphotyrosine containing peptides ofthe docking protein HEPL (Tyr174, 195, 244, and 329), indicatingthat the latter may represent a new Bcr-Abl substrate (seeFigure 1B).Several members of the Bcr-Abl core proteome [6] are adaptor/scaffoldproteins required for the generation of SH2- orPTB-domain binding sites that lead to tightly regulated proteininteractions [9]. Several of these peptides also contain theconsensus sequence YxxP of the enriched phosphorylationmotif. As these proteins bind to Bcr-Abl, the known and novelFigure 1 | Exploring the effect on tyrosine phosphorylation upon Imatinibinhibition of K562 cells.A. Overview of the quantitative proteomics workflow. Cells were treatedwith different doses of Imatinib (0, 1 and 10 µM) for 4 hours, followed bycell lysis and protein digestion. The peptides from each Imatinib treatmentwere then differentially labelled using stable isotope dimethyl labelling. Thethree differentially labelled digests were combined, followed by simultaneousenrichment of tyrosine phosphorylated peptides using immobilisedphosphotyrosine specific antibodies. The enriched fraction was analysed byLC-MS and changes in tyrosine phosphorylation quantified.B. Quantitative profiles of site-specific tyrosine phosphorylation upon Imatinibinhibition of Bcr-Abl. The changes in tyrosine phosphorylation versus thecontrol are represented on a log scale, marked in red, phosphorylation sitescontaining the YxxP motif.
Figure 2 | Alterations in the Gab2 interactome after Imatinib treatment.A. Imatinib treatment of K562 cells leads to destruction of the core Bcr-Abl complex (green, adapted from Brehme et al. [6]). A small selectionof peripheral interactors are depicted in blue. GAB2 (yellow) is used asflag-tagged entry point for interaction analysis upon Imatinib treatment.LC-MS/MS revealed release of GAB2 from the Bcr-Abl core proteome, exceptGRB2 (bold red). Dashed lines indicate as yet to be explored lost/intactinteractions within the Bcr-Abl Core proteome upon Imatinib treatment.B. Mutation of all regulated GAB2 tyrosine phosphorylated residues tophenylalanine reveals abolishment of cell periphery localisation of both GAB2and likely also the Bcr-Abl Core interactome.phosphorylation sites detected here may also be putative Bcr-Abl substrates.Gab2 interactome One member of the Bcr-Abl core proteome,the small adaptor protein GRB2, plays a key role inBcr-Abl signalling through binding to phosphorylated Tyr177on Bcr-Abl. This is required for activation of the Raf-MEK-ERKpathway in leukemic cells [10]. GRB2 also interacts with thedocking proteins Gab1 and Gab2 via GRB2’s SH3 domain andthe proline rich regions in both Gab proteins. The observedtyrosine phosphorylations of both Gab1 and Gab2 were founddown-regulated by Imatinib. We therefore quantitativelyevaluated the changes in the Gab2 interactome in responseto Imatinib treatment. Parallel FLAG-immuno-precipitationsfrom K562 cells transfected with either an empty FLAG-vectoror a FLAG-Gab2 encoding construct and treated with orwithout 10 µM Imatinib were performed. This affinity basedapproach revealed a severe alteration in the Gab2 interactomeafter Imatinib treatment (see Figure 2A). Whereas GRB2remained tightly bound, interactions between Gab2 and othercomponents of the Bcr-Abl core-interactome (e.g. Bcr-Abl,SHIP-2, and Shc1) were severely disrupted, suggesting that theGab2 interactome is dependent on the tyrosine phosphorylationstate of this protein. Subsequent immunoprecipitation ofendogenous Gab2 and Western blot detection of interactorsconfirms these results.The intracellular localisation of Gab2 and the closely relatedGab1 protein were evaluated by confocal microscopy. Whilewild-type GFP-Gab2 smoothly stained the circumference ofthe cell (see Figure 2B), the phospho-dead mutant where allImatinib sensitive phosphotyrosine residues were mutated tophenylalanine showed a severe alteration in localisation withspeckled staining of the cell periphery in addition to cytosolicstaining. Thus, tyrosine phosphorylation of Gab1 and Gab2 andthe subsequent ability to promote the formation of a complexwith Bcr-Abl and other core members appears to be requiredfor appropriate membrane localisation. These observationsstrongly advocate an involvement of Gab2 in the assembly ofthe Bcr-Abl signalling network at the cell periphery and theability of the core proteome to transform cells.summaryReferences1 Hunter, T. (1998) The Croonian Lecture 1997. The phosphorylationof proteins on tyrosine: its role in cell growth anddisease. Philos Trans R Soc Lond B Biol Sci 353, 583-605.2 Salomon, A. R. et al. (2003) Profiling of tyrosine phosphorylationpathways in human cells using mass spectrometry.Proc Natl Acad Sci U S A 100, 443-448.3 Rush, J. et al. (2005) Immunoaffinity profiling of tyrosinephosphorylation in cancer cells. Nat Biotechnol 23, 94-101.4 Boersema, P. J. et al. (2010) In-depth qualitative andquantitative profiling of tyrosine phosphorylation using acombination of phosphopeptide immunoaffinity purificationand stable isotope dimethyl labeling. Mol Cell <strong>Proteomics</strong>9, 84-99.5 Rikova, K. et al. (2007) Global survey of phosphotyrosinesignaling identifies oncogenic kinases in lung cancer. Cell131, 1190-1203.6 Brehme, M. et al. (2009).Charting the molecular networkof the drug target Bcr-Abl. Proc Natl Acad Sci U S A 106,7414-7419.7 Pan, C. et al. (2009) Global effects of kinase inhibitors onsignaling networks revealed by quantitative phosphoproteomics.Mol Cell <strong>Proteomics</strong> 8, 2796-2808.8 Amanchy, R. et al. (2007) A curated compendium of phosphorylationmotifs. Nature biotechnology 25, 285-286.9 Pawson, T. (2004) Specificity in signal transduction: fromphosphotyrosine-SH2 domain interactions to complex cellularsystems. Cell 116, 191-203.10 Pendergast, A. M. et al. (1993) BCR-ABL-induced oncogenesisis mediated by direct interaction with the SH2 domainof the GRB-2 adaptor protein. Cell 75, 175-185.ContactArjen Scholten, PhDBiomolecular Mass Spectrometry & <strong>Proteomics</strong> groupUtrecht UniversityPadualaan 83584 CH Utrecht, the <strong>Netherlands</strong>T +31 30 253 6793a.scholten@uu.nl| 15To probe the effect of Imatinib on tyrosine kinase Bcr-Abl positive cells, we applied a targeted quantitativemass spectrometry based proteomics approach. We usedphosphotyrosine specific immuno-capturing to systematicallyscreen for Imatinib sensitive tyrosine phosphorylation eventsin K562 cells. Our data reveal that several core Bcr-Ablinteractors are putative direct substrates, but also newputative targets are discovered. In following up a novelImatinib inhibited target, the adaptor protein Gab2 wasfound to regulate localisation of the Bcr-Abl core complex tothe cell periphery in a phosphorylation dependent manner,enabling us to add crucial new dimensions of detail to the Bcr-Abl signalling network. This work provides insights into howprotein phosphorylation influences protein networks and showswhich widespread effects a kinase inhibitor may have on thenetwork biology of this leukemic cell.