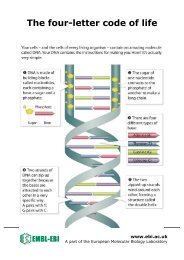
Cell Biology and Biophysics UnitDynamics of membrane traffickingPrevious and current researchMany biological processes at the cellular level are based on complex networks of macromolecularinteractions. These networks have modular organisation, where the modules form dynamic molecularmachines that drive processes such as signalling, cell motility, cytokinesis and vesicle trafficking.Our laboratory’s long-term goal is to contribute to the understanding of the generalprinciples governing the assembly and function of these supramolecular machines.More specifically, we are interested in the formation of cargo-loaded transport vesicles, such asendocytic vesicles. The formation of the endocytic vesicle is driven by a highly dynamic molecularmachinery composed of more than 50 different protein species and several thousand individualprotein molecules. Our main experimental organism is budding yeast, Saccharomyces cerevisiae.We combine powerful yeast genetics with quantitative live-cell imaging methods, with which wehave shown that the endocytic proteins assemble at the endocytic sites in a highly regulated sequenceand form modular machinery that drives vesicle formation. Using mutant yeast strains wehave revealed specific roles for numerous proteins in this process.Marko KaksonenPhD 2002, University ofHelsinki.Postdoctoral research at theUniversity of California,Berkeley.Group leader at <strong>EMBL</strong> since2006.Future projects and goalsIn the future, we will continue to study the membrane trafficking events in budding yeast using live-cell imaging combined with yeast genetics.We will focus on the mechanisms of the assembly of the clathrin-based endocytic machinery and the mechanisms of selective recruitment ofcargo molecules into the endocytic vesicle. We will also extend our work to trafficking events at the Golgi complex. These membrane traffickingevents are highly conserved elemental processes that are involved in multiple biological phenomena ranging from cell polarisation toneural plasticity. As most of the yeast trafficking proteins are widely conserved in eukaryotes, we believe that the mechanisms we unravel inyeast cells will be applicable to eukaryotes in general.A yeast cell expressing fluorescentlylabelledendocytic proteins. The firsttwo images show Sla1 (green) andAbp1 (red) proteins. The last imageshows both channels merged. Thespots at the cell surface revealaccumulation of the proteins atendocytic sites. The proteincomposition of endocytic machinerychanges dynamically during vesicleformation.Selected referencesKaksonen, M., Toret, C.P. & Drubin, D.G. (2006). Harnessing actindynamics for clathrin-mediated endocytosis. Nat. Rev. Mol. CellBiol., 7, 0-1Liu, J., Kaksonen, M., Drubin, D.G. & Oster, G. (2006). Endocyticvesicle scission by lipid phase boundary forces. Proc. Natl. Acad.Sci. USA, 103, 10277-82Kaksonen, M., Toret, C.P. & Drubin, D.G. (2005). A modular designfor the clathrin- and actin-mediated endocytosis machinery. Cell, 12,305-20Kaksonen, M., Sun, Y. & Drubin, D.G. (2003). A pathway forassociation of receptors, adaptors, and actin during endocyticinternalization. Cell, 2, 115, 75-8715
<strong>EMBL</strong> Research at a Glance 2009Michael KnopPhD 1995, University ofStuttgart.Postdoctoral research at theMPI for Biochemistry, Munichand the Beatson Institute forCancer Research, Glasgow.Group leader at the MPI forBiochemistry, Munich.Group leader at <strong>EMBL</strong> since2001.Systems biology of meiosis and mating inbudding yeastPrevious and current researchOur group is interested in the various cellular processes that underlie the sexual cycle of buddingyeast (mating and meiosis). In the past we have addressed the meiosis specific pathways that regulatespore morphogenesis with respect to spindle pole body function, membrane formation andmorphogenesis and cytokinesis (figure 1).We mainly focussed on the processes thatregulate spore morphogenesis in comparisonto cell division by bud formation.Among other things, we concentrated onthe regulation of spindle pole function incontrolling vesicle fusion and in the initiationof spore morphogenesis and on membraneshaping of the spore.Mating is another important aspect of thelife cycle of yeast. How do yeast cells find amating partner? We study the MAP kinasesignal transduction pathway that underliessignal transduction during mating. We established Fluorescence (Cross-) CorrelationSpectroscopy (FCCS) and FLIM (fluorescence lifetime imaging, figure 2) to work withyeast cells. These new quantitative imaging methods enable us to measure protein complexformation and to visualise the activity of the MAP kinases. This yields importantnew insights into the dynamics and the spatial organisation of the signalling process.Future projects and goalsWe continue to use quantitative microscopy approaches and subsequently expand our investigation tothree interconnected MAP kinase signalling pathways by using semi-high throughput screening microscopyto quantify protein concentration, protein-protein interaction and protein localisation of allthe major components involved. We consider both quiescent and signalling conditions. The goal is toenhance our understanding of the spatial and dynamic organisation of the signalling processes. Thiswill help us to derive and further develop quantitative models of the processes that regulate signallingthrough theses pathways.Our work on meiosis has gradually shifted to questions that relate to the role and function of genomerecombination in meiosis. As a model, we use computer simulations of population of yeast-likegenomes that undergo yeast-like life cycles. Here we address the role of meiosis and recombination andthe impact of genome architecture on handling deleterious mutational load. To complement these approaches,we use yeast as a model for experimental evolutionary studies where we address the consequencesof random mutations on fitness, and on the role of meiosis and recombination to purgedeleterious load.Figure 1: Electron micrograph of a forming spore. Thepicture shows a spindle pole body (SPB) that is inprogress of forming a spore membrane.2.2 Lifetime (ns) 2.6Furthermore, we study a novel yeast species with similar live-cycle properties as S. cerevisiae, butwhich has one notable and most interesting difference: this species appears not to recombine its genome during meiosis. We use genome sequencingand experimental approaches to address how this species performs meiosis I and to understand the impact of absent recombinationon the evolution of the genome.Tau avg (ns)Figure 2: High relative Fus3 MAPkinase activity in the matingprojection (shmoo) of pheromonestimulated yeast cells. Fus3 activitywas detected using FLIM (incollaboration with Mark Hink andPhilippe Bastiaens).Selected referencesMaier, P., Rathfelder, N., Maeder, C.I., Colombelli, J., Stelzer, E.H.K.& Knop, M. (2008). The SpoMBe pathway drives membrane bendingnecessary for cytokinesis and spore formation in yeast meiosis.EMBO J., 27, 2363-237Maeder, C.I., Hink, M.A., Kinkhabwala, A., Mayr, R., Bastiaens, P.I. &Knop, M. (2007). Spatial regulation of Fus3 MAP kinase activitythrough a reaction-diffusion mechanism in yeast pheromonesignalling. Nat. Cell Biol., 9, 1319-1326Maier, P., Rathfelder, N., Finkbeiner, M.G., Taxis, C., Mazza, M.,Panse, S.L., Haguenauer-Tsapis, R. & Knop, M. (2007). Cytokinesisin yeast meiosis depends on the regulated removal of Ssp1p fromthe prospore membrane. EMBO J., 26, 183-52Knop, M. (2006). Evolution of the hemiascomycete yeasts: on lifestyles and the importance of inbreeding. Bioessays, 28, 696-70816