H. SahbiConsider a chunk {µ ip }i=1 N <strong>and</strong> fix {µ jk, k ̸= p} N j=1 ,theobjective function (7) is linear in terms of {µ ip } <strong>and</strong> canbe solved, with respect to this chunk, using linear programming.Only one equality constraint ∑ i µ ip = 1 is takeninto account in the linear programming problem as the otherequality constraints in (6) are independent from the chunk{µ ip }.We can further reduce the size of the chunk {µ ip }i=1 N toonly two parameters. Given i 1 , i 2 ∈ {1,...,N}, we canrewrite (7) as:min 2 ( µ i1 p c i1 p + µ i2 p c i2 p)+ bµ i1 p, µ i2 ps.t µ i1 p + µ i2 p = 1 − ∑µ ipi ̸= i 1 ,i 20 ≤ µ i1 p ≤ 10 ≤ µ i2 p ≤ 1,here b is a constant independent from the chunk {µ i1 p,µ i2 p}(see 7). When c i1 p = c i2 p the above objective function <strong>and</strong>the equality constraints are linearly dependent, so we have aninfinite set of optimal solutions; any one which satisfies theconstraints in (9) can be considered. Let us assume c i1 p ̸=c i2 p <strong>and</strong> denote d = 1 − ∑ i ̸= i 1 ,i 2µ ip , since µ i2 p = d −µ i1 p, the above linear programming problem can be writtenas:minµ i1 p (c i1 p − c i2 p)µ i1 ps.t max(d − 1, 0) ≤ µ i1 p ≤ min(d, 1)(9)(10)Depending on the sign of c i1 p − c i2 p, the solution of (10) issimply taken as one of the bounds of the inequality constraintin (10). At this stage, the parameters {µ 1p ,...,µ Np } whichsolve (7) are found by solving iteratively the trivial minimizationproblem (10) <strong>for</strong> different chunks, as shown inAlgorithm1,Algorithm1(p,µ)Do the following steps ITERMAX1 iterationsSelect r<strong>and</strong>omly (i 1 , i 2 ) ∈{1,...,N} 2 .(c i1 p, c i2 p) ←− (8).if (c i1 p − c i2 p < 0) µ i1 p ←− min(d, 1) (see 10)else µ i1 p ←− max(d − 1, 0)µ i2 p ←− d − µ i1 p (see 9)endDoreturn (µ i1 p,...,µ iN p)<strong>and</strong> the whole QP (6) is solved by looping several timesthrough different cluster indices as shown in Algorithm2.For a convex <strong>for</strong>m of the QP (6) <strong>and</strong> <strong>for</strong> a large value of ITER-MAX1 <strong>and</strong> ITERMAX2, each subproblem will be convex<strong>and</strong> the convergence of the decomposition algorithms (1,2) isguaranteed as also discussed <strong>for</strong> support vector machine training(Platt 1999). In practice ITERMAX1 <strong>and</strong> ITERMAX2are set to C <strong>and</strong> N 2 respectively.Algorithm2Set {µ} to r<strong>and</strong>om values.Do the following steps ITERMAX2 iterationsFix p in {1,...,C}, update µ 1p ,…,µ Np using:(µ 1p ,..., µ Np ) ←− Algorithm1(p,µ)endDo4.2 AgglomerationInitially, the algorithm described above considers an overestimatednumber of clusters (denoted C). This will resultinto several overlapping cluster GMMs which might be detectedusing a similarity measure <strong>for</strong> instance the KullbackLeibler divergence (Bishop 1995).Given two cluster GMMs p(x|y k ), p(x|y l ),<strong>for</strong>afixedɛ>0 we declare p(x|y k ), p(x|y l ) as similar if:∫X∥ p(x|yk ) − p(x|y l ) ∥ ∥ 2 dx < ɛ(11)As p(x|y k ) =〈ω k ,Φ σ (x)〉 <strong>and</strong> p(x|y l ) =〈ω l ,Φ σ (x)〉, wecan simply rewrite (11) as:∫X〈ω k − ω l ,Φ σ (x)〉 2 dx < ɛ(12)We can set the memberships in ω k , ω l such that ‖ω k ‖=‖ω l ‖=1. Now, from (12) it is clear that two overlappingGMMs correspond to two highly correlated hyperplanes inthe mapping space, i.e., 〈ω k ,ω l 〉 ≥ τ (<strong>for</strong> a fixed thresholdτ ∈[0, 1]).Regularization: notice that,∂〈 ω k ,ω l 〉∂σ≥ 0, (13)so the correlation is an increasing function of the scale σ .Assuming ‖w k ‖=‖w l ‖=1, it results that ∀ τ ∈[0, 1], ∃ σsuch that 〈ω k ,ω l 〉 ≥ τ.Let us define the following adjacency matrix:A k,l ={ 1 if〈ωk ,ω l 〉 ≥ τ0 otherwise(14)This matrix characterizes a graph where a node is a hyperplane<strong>and</strong> an arc corresponds to two highly correlated hyperplanes(i.e., 〈ω k ,ω l 〉 ≥ τ). Now, the actual number of clusters isfound as the number of connected components in this graph.For a fixed threshold τ, the scale parameter σ acts as a regularizer.When σ is overestimated, the graph A ={A k,l } willbe connected resulting into only one big cluster whereas a123
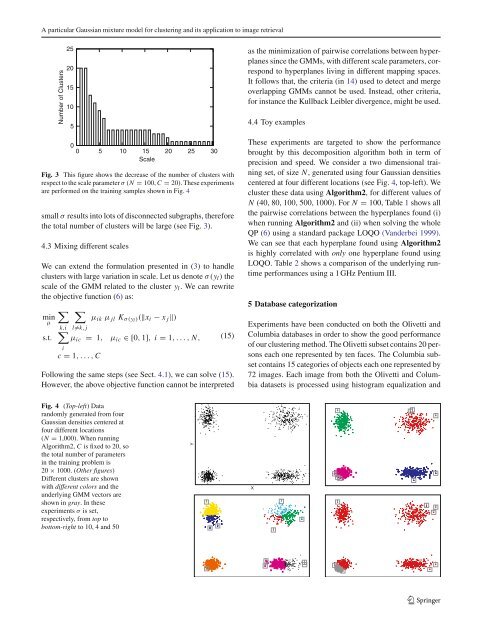
A <strong>particular</strong> <strong>Gaussian</strong> <strong>mixture</strong> <strong>model</strong> <strong>for</strong> <strong>clustering</strong> <strong>and</strong> <strong>its</strong> application to image retrievalNumber of Clusters25201510500 5 10 15 20 25 30ScaleFig. 3 This figure shows the decrease of the number of clusters withrespect to the scale parameter σ (N = 100, C = 20). These experimentsare per<strong>for</strong>med on the training samples shown in Fig. 4small σ results into lots of disconnected subgraphs, there<strong>for</strong>ethe total number of clusters will be large (see Fig. 3).4.3 Mixing different scalesWe can extend the <strong>for</strong>mulation presented in (3) to h<strong>and</strong>leclusters with large variation in scale. Let us denote σ(y l ) thescale of the GMM related to the cluster y l . We can rewritethe objective function (6)as:minµs.t.∑k,i∑l̸=k, jµ ik µ jl K σ(yl )(‖x i − x j ‖)∑µ ic = 1, µ ic ∈[0, 1], i = 1,...,N,ic = 1,...,C(15)Following the same steps (see Sect. 4.1), we can solve (15).However, the above objective function cannot be interpretedas the minimization of pairwise correlations between hyperplanessince the GMMs, with different scale parameters, correspondto hyperplanes living in different mapping spaces.It follows that, the criteria (in 14) used to detect <strong>and</strong> mergeoverlapping GMMs cannot be used. Instead, other criteria,<strong>for</strong> instance the Kullback Leibler divergence, might be used.4.4 Toy examplesThese experiments are targeted to show the per<strong>for</strong>mancebrought by this decomposition algorithm both in term ofprecision <strong>and</strong> speed. We consider a two dimensional trainingset, of size N, generated using four <strong>Gaussian</strong> densitiescentered at four different locations (see Fig. 4, top-left). Wecluster these data using Algorithm2, <strong>for</strong> different values ofN (40, 80, 100, 500, 1000). For N = 100, Table 1 shows allthe pairwise correlations between the hyperplanes found (i)when running Algorithm2 <strong>and</strong> (ii) when solving the wholeQP (6) using a st<strong>and</strong>ard package LOQO (V<strong>and</strong>erbei 1999).We can see that each hyperplane found using Algorithm2is highly correlated with only one hyperplane found usingLOQO. Table 2 shows a comparison of the underlying runtimeper<strong>for</strong>mances using a 1 GHz Pentium III.5 Database categorizationExperiments have been conducted on both the Olivetti <strong>and</strong>Columbia databases in order to show the good per<strong>for</strong>manceof our <strong>clustering</strong> method. The Olivetti subset contains 20 personseach one represented by ten faces. The Columbia subsetcontains 15 categories of objects each one represented by72 images. Each image from both the Olivetti <strong>and</strong> Columbiadatasets is processed using histogram equalization <strong>and</strong>Fig. 4 (Top-left) Datar<strong>and</strong>omly generated from four<strong>Gaussian</strong> densities centered atfour different locations(N = 1,000). When runningAlgorithm2, C is fixed to 20, sothe total number of parametersin the training problem is20 × 1000. (Other figures)Different clusters are shownwith different colors <strong>and</strong> theunderlying GMM vectors areshown in gray. Intheseexperiments σ is set,respectively, from top tobottom-right to 10, 4 <strong>and</strong> 50YX123