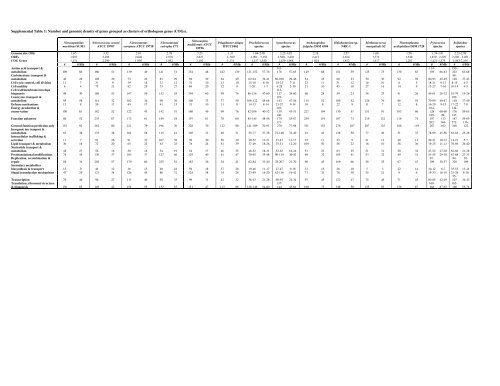
Supplemental Table 1: Number and genomic <strong>de</strong>nsity of genes grouped as clusters of orthologous genes (COGs). <strong>Nitrosopumilus</strong> <strong>maritimus</strong> SCM1 Nitrosococcus oceani ATCC 19707 Nitrosomonas europaea ATCC 19718 Nitrosomonas eutropha C71 Nitrosospira multiformis ATCC 25196 Pe<strong>la</strong>gibacter ubique HTCC1062 Procholoroccus species Synechococcus species Archaeoglobus fulgidus DSM 4304 Halobacterium sp. NRC-1 Methanococcus maripaludis S2 Thermop<strong>la</strong>sma acidophilum DSM 1728 Genome size (Mb) 1.65 3.52 2.81 2.78 3.23 1.31 1.64-2.68 2.22-3.05 2.18 2.57 1.66 1.56 1.74-1.91 2.23-2.99 ORFs 1,997 3,186 2,628 2,578 2,827 1,.389 1,901 - 3,152 2,580 - 3,401 2,471 2,674 1,772 1,548 1,879 - 2,229 2,305 - 3,031 COG Genes 1,131 2,290 1,995 1,952 2,102 1,131 1,157-1,540 1,459-1,968 1,918 1,812 1,417 1,201 1,421-1,575 1,563-2,105 # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb # #/Mb Amino acid transport & 141- 114- 150metabolism 109 66 180 51 139 49 141 51 154 48 182 139 121-152 57-76 178 57-65 149 68 151 59 125 75 130 83 159 66-83 202 63-68 Carbohydrate transport & 88metabolism 46 28 105 30 73 26 81 29 94 29 64 49 62-94 34-41 86-108 29-46 54 25 60 23 50 30 92 59 80-93 45-49 128 37-43 Cell cycle control, cell division 11 7 31 9 39 14 32 12 31 10 13 10 15-18 6-10 18-33 7-11 23 11 31 12 16 10 8 5 18-21 9-12 8-15 4-5 Cell motility 6 4 75 21 82 29 75 27 64 20 12 9 1-20 1-7 4-28 2-10 21 10 45 18 27 16 14 9 13-27 7-16 10-14 4-5 Cell wall/membrane/envelope 113biogenesis 49 30 186 53 167 59 152 55 194 60 99 76 86-136 47-60 137 38-60 60 28 59 23 38 23 41 26 46-61 26-32 51-79 19-26 Coenzyme transport & 122- 102metabolism 95 58 114 32 102 36 99 36 108 33 77 59 106-122 45-66 143 47-56 114 52 109 42 126 76 86 55 70-89 40-47 110 37-49 Defense mechanisms 13 8 59 17 49 17 41 15 31 10 11 8 14-33 8-14 23-37 9-14 18 8 22 9 11 7 12 8 16-29 9-15 17-22 7-8 Energy production & 110- 104- 144conservation 100 61 182 52 122 43 142 51 160 49 99 76 82-106 40-52 139 43-51 227 104 130 51 151 91 103 66 128 60-68 176 59-65 140- 189- 99- 147- Function unknown 86 52 235 67 172 61 149 54 197 61 78 60 85-145 48-56 179 55-67 259 119 187 73 219 132 116 74 197 113 187 59-69 171- 262- 144- 272- 116- General function prediction only Inorganic ion tranport & 152 92 282 80 221 79 196 70 228 70 112 86 141-189 70-91 270 77-98 331 152 276 107 207 125 186 119 287 162 346 122 metabolism Intracellu<strong>la</strong>r trafficking & 63 38 135 38 164 58 115 41 105 32 40 31 55-77 27-38 72-148 32-49 91 42 128 50 77 46 51 33 78-99 45-56 63-85 25-28 secretion 11 7 91 26 91 32 107 38 96 30 38 29 20-52 12-21 27-47 12-17 25 11 23 9 21 13 20 13 20-21 10-12 14-21 5-7 Lipid transport & metabolism Nucleoti<strong>de</strong> transport & 30 18 72 20 65 23 63 23 78 24 51 39 37-49 18-26 35-51 12-20 109 50 56 22 16 10 56 36 19-25 11-13 70-89 26-40 metabolism 45 27 56 16 50 18 54 19 54 17 46 35 48-52 18-31 52-63 18-24 51 23 63 25 51 31 50 32 47-52 27-28 62-66 21-28 Posttrans<strong>la</strong>tional modifications 74 45 130 37 103 37 127 46 129 40 61 47 70-95 35-48 90-119 36-42 69 32 105 41 53 32 49 31 51-55 29-30 67-88 27-33 Replication, recombination & 67- 84- 38repair Secondary metabolites 56 34 202 57 170 60 225 81 182 56 54 41 62-84 31-40 78-207 31-70 98 45 169 66 58 35 67 43 109 38-57 326 109 biosynthesis & transport 15 9 44 12 36 13 49 18 59 18 37 28 19-46 11-17 27-47 9-18 33 15 26 10 5 3 22 14 10-12 6-7 35-53 13-24 Singal transduction mechanisms 47 29 121 34 126 45 86 31 124 38 34 26 23-49 14-20 42-116 19-42 71 33 76 30 35 21 9 6 19-33 10-19 23-28 95- 8-10 Transcription 76 46 96 27 113 40 92 33 99 31 42 32 36-63 21-26 60-93 24-34 97 45 122 47 75 45 71 45 80-85 42-49 107 34-43 Trans<strong>la</strong>tion, ribosomal structure 137- 160- 165- & biogenesis 136 83 149 42 154 55 152 55 151 47 113 86 128-146 54-80 144 45-64 160 73 148 58 155 93 136 87 166 87-92 166 55-74 Pyrococcus species Sulfolobus species
Supplemental Table 2: Predicted coordinates for genes implicated in novel archaeal biochemistry for ammonia oxidation and respiratory chain. NCBI locus tag Protein # Strand Left bp Right bp Annotation Best Hit Analog in Ammonia oxidizing Bacteria Nmar_0588 ABX12484 F 523,890 525,197 ammonium transporter AmtB Nmar_1698 ABX13594 R 1,559,816 1,558,251 ammonium transporter AmtB Nmar_1500 ABX13396 R 1,367,890 1,367,240 putative ammonia monooxygenase, subunit A AMO (AmoA) Nmar_1501 ABX13397 R 1,368,387 1,368,025 hypothetical protein Nmar_1502 ABX13398 R 1,369,079 1,368,507 putative ammonia monooxygenase, subunit C AMO (AmoC) Nmar_1503 ABX13399 F 1,369,326 1,369,895 putative ammonia monooxygenase, subunit B AMO (AmoB) Nmar_0815 ABX12711 R 718,761 720,380 soluble perip<strong>la</strong>smic BCP (complete upstream CxxxxxC domain, complete CxxHxxM domain [C]) PetE, p<strong>la</strong>stocyanin cytochrome c552 Nmar_1102 ABX12998 F soluble perip<strong>la</strong>smic BCP (complete upstream CxxxxxC domain, complete 1,004,439 1,005,395 CxxHxxM domain [C]) PetE, p<strong>la</strong>stocyanin cytochrome c552 Nmar_1443 ABX13339 R soluble perip<strong>la</strong>smic BCP (complete upstream CxxxxxC domain, complete 1,314,030 1,313,572 CxxHxxM domain [C]) PetE, p<strong>la</strong>stocyanin cytochrome c552 Nmar_1637 ABX13533 R soluble perip<strong>la</strong>smic BCP (complete upstream CxxxxxC domain, complete 1,494,547 1,492,676 CxxHxxM domain [C]) PetE, p<strong>la</strong>stocyanin cytochrome c552 Nmar_0004 ABX11904 F 3,463 4,359 soluble perip<strong>la</strong>smic BCP (modified upstream CxxxxxC domain, modified CxxHxxM domain [C]) Halocyanin, P39442 cytochrome c552 Nmar_1307 ABX13203 F soluble perip<strong>la</strong>smic BCP (modified upstream CxxxxxC domain, modified 1,195,212 1,195,736 CxxHxxM domain [C]) Halocyanin, P39442 cytochrome c552 Nmar_0918 ABX12814 R 802,019 802,861 perip<strong>la</strong>smic membrane BCP (1 TMS [C], modified upstream CxxxxxC domain, complete CxxHxxM domain [M]) p<strong>la</strong>stocyanin anchored cytochrome c552 Nmar_1678 ABX13574 R perip<strong>la</strong>smic membrane BCP (1 TMS [C], modified upstream CxxxxxC domain, 1,538,377 1,537,367 complete CxxHxxM domain [N]) p<strong>la</strong>stocyanin anchored cytochrome c552 Nmar_1273 ABX13169 R perip<strong>la</strong>smic membrane BCP (1 TMS [C], complete upstream CxxxxxC domain, 1,169,255 1,168,392 modified CxxHxxM domain [C]) p<strong>la</strong>stocyanin anchored cytochrome c552 Nmar_1161 ABX13057 R 1063718 1062348 perip<strong>la</strong>smic membrane BCP (1 TMS [N], modified upstream CxxxxxC domain, modified CxxHxxM domain [M]) p<strong>la</strong>stocyanin anchored cytochrome c552 Nmar_1226 ABX13122 F perip<strong>la</strong>smic membrane BCP (4 TMS [N], modified upstream CxxxxxC domain, 2 1,126,185 1,127,261 complete CxxHxxM domains [C]) p<strong>la</strong>stocyanin cM552 Nmar_1142 ABX13038 F cytop<strong>la</strong>smic membrane BCP (1 TMS [N], modified upstream CxxxxxC domain, 1,045,689 1,046,150 complete CxxHxxM domain [C]) Nmar_1542 "cytop<strong>la</strong>smic BCP" Nmar_0276 ABX12172 F 243,850 244,182 CI, NADH-ubiquinone/p<strong>la</strong>stoquinone oxidoreductase chain 3 Nmar_0277 ABX12173 F 244,223 244,747 CI, NADH-quinone oxidoreductase, B subunit Nmar_0278 ABX12174 F 244,747 245,349 CI, NADH <strong>de</strong>hydrogenase (ubiquinone) 30 kDa subunit Nmar_0279 ABX12175 F 245,352 246,488 CI, NADH <strong>de</strong>hydrogenase (quinone) Nmar_0280 ABX12176 F 246,489 247,787 CI, NADH <strong>de</strong>hydrogenase (quinone) Nmar_0281 ABX12177 F 247,787 248,284 CI, [4Fe-4S] ferredoxin iron-sulfur binding domain protein Complex I Nmar_0282 ABX12178 F 248,277 248,789 CI, NADH-ubiquinone/p<strong>la</strong>stoquinone oxidoreductase chain 6 Nmar_0283 ABX12179 F 248,770 249,075 CI, NADH-ubiquinone oxidoreductase chain 4L Nmar_0284 ABX12180 F 249,075 250,628 CI, proton-translocating NADH-quinone oxidoreductase, chain M Nmar_0285 ABX12181 F 250,630 252,708 CI, proton-translocating NADH-quinone oxidoreductase, chain L Nmar_0286 ABX12182 F 252,721 254,205 CI, proton-translocating NADH-quinone oxidoreductase, chain N Nmar_1542 ABX13438 R CIII, cytop<strong>la</strong>smic membrane BCP, 1 TMS (N), modified upstream CxxxxxC 1,404,537 1,404,073 domain, complete CxxHxxM domain (C) Nmar_1142 membrane-bound cytochrome c1 Nmar_1543 ABX13439 R 1,406,185 1,404,590 CIII, Cytochrome b/b6 domain Complex III cyt. B Nmar_1544 ABX13440 R 1,406,774 1,406,169 CIII, Rieske [2Fe-2S] domain protein Complex III [Fe-S] Nmar_0182 ABX12078 F 166,066 166,326 CIV, hypothetical protein HCO subunit IV Nmar_0183 ABX12079 F 166,329 166,760 CIV, heme-copper oxidase subunit II HCO subunit II Nmar_0184 ABX12080 F 166,798 168,324 CIV, heme-copper oxidase subunit I HCO subunit I Nmar_0185 ABX12081 F 168,337 169,284 CIV, perip<strong>la</strong>smic membrane BCP, 1 TMS (N), complete upstream CxxxxxC domain, complete CxxHxxM domain (C) Nmar_1354 cytochrome c subunit III Nmar_1259 ABX13155 R 1,155,995 1,154,583 NirK, soluble perip<strong>la</strong>smic MCO (Cu-oxidase_1; Cu-oxidase_2; Cu-oxidase_3), Nwi_2648; Nham_3281; NE0924; Neut_1403; Noc_0089 3dMCO, NirK Nmar_1661 ABX13557 R 1,519,832 1,519,440 Transcriptional regu<strong>la</strong>tor, MarR family (winged helix-turn-helix HxlR type) HTH -regu<strong>la</strong>tor MarR Nmar_1662 ABX13558 R 1,521,322 1,520,144 Metal cation transporter (ZIP Zinc transporter; cl00437) Zip Nmar_1663 ABX13559 R soluble perip<strong>la</strong>smic 2dMCO (Cu-oxidase_2; Cu-oxidase_3), simi<strong>la</strong>r to NcgA 1,522,373 1,521,312 (BCO) Nwi_2651; Nham_3284; NE0927; Neut_1406 2dMCO, NcgA (BCO) Nmar_1664 ABX13560 R 1,522,974 1,522,483 metal (Mn, Fe) <strong>de</strong>pen<strong>de</strong>nt repressor protein, DtxR family Nmar_1132 HTH -regu<strong>la</strong>tor DtxR Nmar_1665 ABX13561 R soluble perip<strong>la</strong>smic BCP (modified upstream CxxxxxC domain, modified 1,523,638 1,523,135 CxxHxxM domain [C]) Halocyanin, P39442 monoheme cytochrome c Nmar_1666 ABX13562 R beta-propeller structure oxidase (Kelch repeat-containing protein 1,524,788 1,523,754 [CDD:121499]) Nmar_1133 beta propeller structure Nmar_1667 ABX13563 R 1,526,542 1,525,130 Nmar_1136 ABX13032 R 1,041,807 1,040,200 NirK, soluble perip<strong>la</strong>smic 3d MCO (Cu-oxidase_1; Cu-oxidase_2; Cuoxidase_3), NsrR motif in upstream region. soluble perip<strong>la</strong>smic 3dMCO (Cu-oxidase_1; Cu-oxidase_2; Cu-oxidase_3), simi<strong>la</strong>r to NcgA Nwi_2648; Nham_3281; NE0924; Neut_1403; Noc_0089 Nwi_2651; Nham_3284; NE0927; Neut_1406 3dMCO, Nirk 3dMCO- modified NcgA (BCO) Nmar_1135 ABX13031 R 1,040,071 1,038,395 putative <strong>de</strong>hydrogenase with Rossmann-fold NAD(P) + -binding domain [cl09931] Rossmann-fold - DH Nmar_1134 ABX13030 R 1,038,244 1,037,921 hypothetical protein Nmar_1133 ABX13029 R beta-propeller structure oxidase (Kelch repeat-containing protein 1,037,891 1,036,866 [CDD:121499]) Nmar_1666 beta propeller structure Nmar_1132 ABX13028 R 1,036,419 1,035,937 metal (Mn, Fe) <strong>de</strong>pen<strong>de</strong>nt repressor protein, DtxR family Nmar_1664 HTH -regu<strong>la</strong>tor DtxR Nmar_1131 ABX13027 R soluble perip<strong>la</strong>smic 2d MCO (Cu-oxidase_2; Cu-oxidase_3), simi<strong>la</strong>r to NcgA 1,035,788 1,034,733 (BCO) Nwi_2651; Nham_3284; NE0927; Neut_1406 2dMCO NcgA (BCO) Nmar_1130 ABX13026 R 1,034,743 1,033,565 Metal cation transporter (ZIP Zinc transporter; cl00437) Zip Nmar_1129 ABX13025 R cytop<strong>la</strong>smic membrane BCP (3 TMS [1N, 2C], complete upstream CxxxxxC 1,031,846 1,033,210 domain, complete CxxHxxM domain [N) "cytop<strong>la</strong>smic BCP" Nmar_1128 ABX13024 R perip<strong>la</strong>smic binding protein (transport of ferric si<strong>de</strong>rophores and metal ions such 1,031,477 1,030,587 as Mn2+, Fe3+, Cu2+ and/or Zn2+) cation BP Nmar_1352 ABX13248 R 1,238,660 1,237,860 Transcriptional regu<strong>la</strong>tor, ArsR family (winged helix-turn-helix type, COG4742) HTH -regu<strong>la</strong>tor Nmar_1353 ABX13249 R 1,239,270 1,238,728 Uncharacterized conserved protein [COG3945; Function unknown] Nmar_1354 ABX13250 R Fusion Protein: soluble perip<strong>la</strong>smic 2d MCO (Cu-oxidase_2; Cu-oxidase_3) - 1,240,615 1,239,281 BCP (modified upstream CxxxxxC domain & CxxHxxM domain [C]) Nmar_1131; Nmar_0185 2dMCO - BCP bi-functional MCO Nmar_1355 ABX13251 R 1,242,507 1,241,821 hypothetical protein
- Page 1 and 2: Nitrosopumilus maritimus genome rev
- Page 3 and 4: ammonia oxidation. There are two hy
- Page 5 and 6: Fig. 3. Comparison of N. maritimus
- Page 7 and 8: Supporting Information Walker et al
- Page 9 and 10: Fig. S2. Archaeal ammonia monooxyge
- Page 11: A. 0.1 B. 100 90 52 Other Supportin
- Page 15 and 16: Supplemental Table 3: Gene predicti
- Page 17 and 18: Nmar_tR26 1,630,610 1,630,537 ProGG
- Page 19 and 20: Supplemental Table 4: Gene predicti
- Page 21 and 22: Nmar_0258 CENSYa_1705 82 227,624 22
- Page 23 and 24: Nmar_0493 CENSYa_1354 56 443,851 44
- Page 25 and 26: Nmar_0748 CENSYa_0915 84 675,576 67
- Page 27 and 28: Nmar_1022 CENSYa_1172 31 901,921 90
- Page 29 and 30: Nmar_1303 CENSYa_0131 86 1,193,261
- Page 31 and 32: Nmar_1564 CENSYa_2017 44 1,432,780
- Page 33 and 34: Nmar_1783 CENSYa_1861 52 1,629,775
- Page 35 and 36: Molybdopterin-guanine dinucleotide
- Page 37 and 38: Nmar0935 DeepAntEC39 f 815,847 816,
- Page 39 and 40: Nmar0101 EQ085329 r 91,543 93,711 A
- Page 41 and 42: Nmar0436 EQ085799 f 386,930 387,319
- Page 43 and 44: Nmar0982 EP710054 r 859,116 859,526
- Page 45 and 46: Nmar1297 EQ086682 f 1,187,165 1,188