Nitrosopumilus maritimus genome reveals unique - de la Torre ...
Nitrosopumilus maritimus genome reveals unique - de la Torre ...
Nitrosopumilus maritimus genome reveals unique - de la Torre ...
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
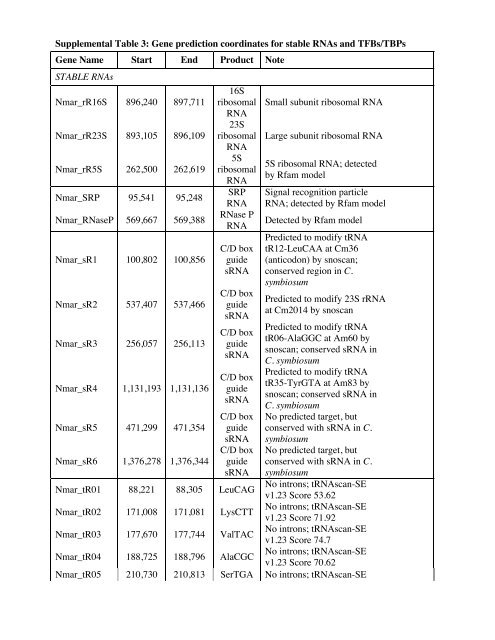
Supplemental Table 3: Gene prediction coordinates for stable RNAs and TFBs/TBPs<br />
Gene Name Start End Product Note<br />
STABLE RNAs<br />
Nmar_rR16S 896,240 897,711<br />
Nmar_rR23S 893,105 896,109<br />
Nmar_rR5S 262,500 262,619<br />
Nmar_SRP 95,541 95,248<br />
Nmar_RNaseP 569,667 569,388<br />
Nmar_sR1 100,802 100,856<br />
Nmar_sR2 537,407 537,466<br />
Nmar_sR3 256,057 256,113<br />
Nmar_sR4 1,131,193 1,131,136<br />
Nmar_sR5 471,299 471,354<br />
Nmar_sR6 1,376,278 1,376,344<br />
16S<br />
ribosomal<br />
RNA<br />
23S<br />
ribosomal<br />
RNA<br />
5S<br />
ribosomal<br />
RNA<br />
SRP<br />
RNA<br />
RNase P<br />
RNA<br />
C/D box<br />
gui<strong>de</strong><br />
sRNA<br />
C/D box<br />
gui<strong>de</strong><br />
sRNA<br />
C/D box<br />
gui<strong>de</strong><br />
sRNA<br />
C/D box<br />
gui<strong>de</strong><br />
sRNA<br />
C/D box<br />
gui<strong>de</strong><br />
sRNA<br />
C/D box<br />
gui<strong>de</strong><br />
sRNA<br />
Nmar_tR01 88,221 88,305 LeuCAG<br />
Nmar_tR02 171,008 171,081 LysCTT<br />
Nmar_tR03 177,670 177,744 ValTAC<br />
Small subunit ribosomal RNA<br />
Large subunit ribosomal RNA<br />
5S ribosomal RNA; <strong>de</strong>tected<br />
by Rfam mo<strong>de</strong>l<br />
Signal recognition particle<br />
RNA; <strong>de</strong>tected by Rfam mo<strong>de</strong>l<br />
Detected by Rfam mo<strong>de</strong>l<br />
Predicted to modify tRNA<br />
tR12-LeuCAA at Cm36<br />
(anticodon) by snoscan;<br />
conserved region in C.<br />
symbiosum<br />
Predicted to modify 23S rRNA<br />
at Cm2014 by snoscan<br />
Predicted to modify tRNA<br />
tR06-A<strong>la</strong>GGC at Am60 by<br />
snoscan; conserved sRNA in<br />
C. symbiosum<br />
Predicted to modify tRNA<br />
tR35-TyrGTA at Am83 by<br />
snoscan; conserved sRNA in<br />
C. symbiosum<br />
No predicted target, but<br />
conserved with sRNA in C.<br />
symbiosum<br />
No predicted target, but<br />
conserved with sRNA in C.<br />
symbiosum<br />
No introns; tRNAscan-SE<br />
v1.23 Score 53.62<br />
No introns; tRNAscan-SE<br />
v1.23 Score 71.92<br />
No introns; tRNAscan-SE<br />
v1.23 Score 74.7<br />
Nmar_tR04 188,725 188,796 A<strong>la</strong>CGC<br />
No introns; tRNAscan-SE<br />
v1.23 Score 70.62<br />
Nmar_tR05 210,730 210,813 SerTGA No introns; tRNAscan-SE