2011 Annual Report - Center for Integrated Nanotechnologies - Los ...
2011 Annual Report - Center for Integrated Nanotechnologies - Los ...
2011 Annual Report - Center for Integrated Nanotechnologies - Los ...
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Theory & Simulation of Nanoscale Phenomena Thrust<br />
Electronic Fingerprints of DNA Bases on Graphene<br />
Accomplishment: In this letter,<br />
the use of graphene is proposed<br />
as a deposition substrate <strong>for</strong><br />
macromolecular sequencing.<br />
Both the Dirac linear dispersion<br />
of electronic states in graphene<br />
and its robust mechanical properties<br />
serve to make it a superior<br />
substrate compared to metals.<br />
The authors have calculated the<br />
tunneling conductance and local<br />
density of states (LDOS) <strong>for</strong><br />
the specific case of single DNA<br />
bases at various orientations<br />
on the surface and have shown<br />
that the different nucleotides<br />
have significantly different LDOS<br />
peaks (fingerprints), allowing<br />
differentiation via local tunneling<br />
conductance. The calculated<br />
nitrogen atom LDOS <strong>for</strong> A, T,<br />
C, and G bases show distinct<br />
peaks that vary depending on<br />
configuration. However, a planar<br />
orientation of DNA bases with<br />
respect to the graphene surface<br />
is the most stable, since it favors<br />
π−π stacking between the base<br />
and aromatic carbons in graphene.<br />
It is found that guanine<br />
provides the largest peak at a<br />
negative bias at −1.3 eV. Peaks<br />
<strong>for</strong> the other bases at higher energies<br />
have features at positive and negative scanning tunneling<br />
microscopy (STM) bias and provide fingerprints that allow unique<br />
identification. The calculated LDOS fingerprints can help guide<br />
scanning tunneling spectroscopy (STS) as an approach to identify<br />
DNA bases, and likely other biomolecules on graphene surfaces.<br />
Challenge: The ability to sequence DNA quickly and af<strong>for</strong>dably is<br />
one of the great challenges in this era. The determination of the<br />
precise sequence of the four nucleotides [adenine (A), cytosine<br />
(C), guanine (G), and thymine (T)] in DNA molecules is an important<br />
goal <strong>for</strong> both fundamental research interests as well as large<br />
number of applications in biomedical research, biotechnology,<br />
drug delivery, and biomaterial growth.<br />
Significance: This work develops a detailed understanding of the<br />
interactions and adsorption between graphene and biomolecules<br />
using an approach goes beyond the structural analysis of such<br />
hybrid systems. In this letter, the possibilities of using STM and<br />
STS <strong>for</strong> the determination and sequencing of DNA on substrate<br />
helps provide an opportunity <strong>for</strong> electronic identifications of all<br />
bases via the tunneling conductance from a local probe.<br />
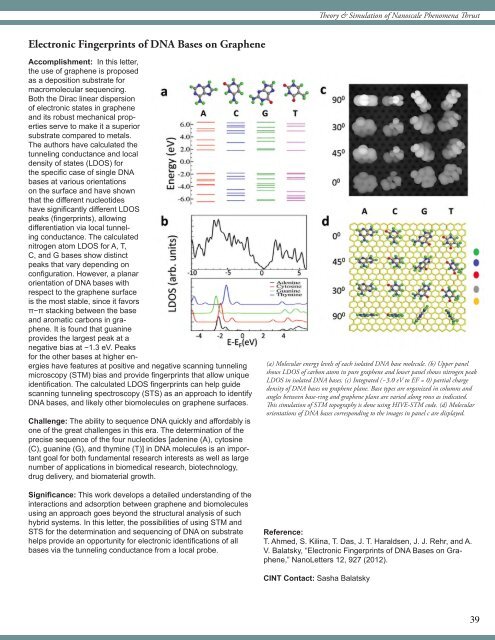
(a) Molecular energy levels of each isolated DNA base molecule. (b) Upper panel<br />
shows LDOS of carbon atom in pure graphene and lower panel shows nitrogen peak<br />
LDOS in isolated DNA bases. (c) <strong>Integrated</strong> (−3.0 eV to EF = 0) partial charge<br />
density of DNA bases on graphene plane. Base types are organized in columns and<br />
angles between base-ring and graphene plane are varied along rows as indicated.<br />
This simulation of STM topography is done using HIVE-STM code. (d) Molecular<br />
orientations of DNA bases corresponding to the images in panel c are displayed.<br />
Reference:<br />
T. Ahmed, S. Kilina, T. Das, J. T. Haraldsen, J. J. Rehr, and A.<br />
V. Balatsky, “Electronic Fingerprints of DNA Bases on Graphene,”<br />
NanoLetters 12, 927 (2012).<br />
CINT Contact: Sasha Balatsky<br />
39