Combinatorial and High-Throughput Screening of Materials ...
Combinatorial and High-Throughput Screening of Materials ...
Combinatorial and High-Throughput Screening of Materials ...
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
ACS <strong>Combinatorial</strong> Science<br />
REVIEW<br />
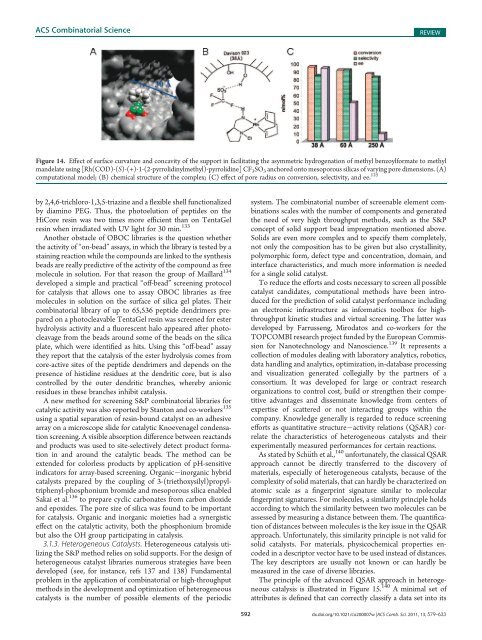
Figure 14. Effect <strong>of</strong> surface curvature <strong>and</strong> concavity <strong>of</strong> the support in facilitating the asymmetric hydrogenation <strong>of</strong> methyl benzoylformate to methyl<br />
m<strong>and</strong>elate using [Rh(COD)-(S)-(+)-1-(2-pyrrolidinylmethyl)-pyrrolidine] CF 3 SO 3 anchored onto mesoporous silicas <strong>of</strong> varying pore dimensions. (A)<br />
computational model; (B) chemical structure <strong>of</strong> the complex; (C) effect <strong>of</strong> pore radius on conversion, selectivity, <strong>and</strong> ee. 125<br />
by 2,4,6-trichloro-1,3,5-triazine <strong>and</strong> a flexible shell functionalized<br />
by diamino PEG. Thus, the photoelution <strong>of</strong> peptides on the<br />
HiCore resin was two times more efficient than on TentaGel<br />
resin when irradiated with UV light for 30 min. 133<br />
Another obstacle <strong>of</strong> OBOC libraries is the question whether<br />
the activity <strong>of</strong> “on-bead” assays, in which the library is tested by a<br />
staining reaction while the compounds are linked to the synthesis<br />
beads are really predictive <strong>of</strong> the activity <strong>of</strong> the compound as free<br />
molecule in solution. For that reason the group <strong>of</strong> Maillard 134<br />
developed a simple <strong>and</strong> practical “<strong>of</strong>f-bead” screening protocol<br />
for catalysis that allows one to assay OBOC libraries as free<br />
molecules in solution on the surface <strong>of</strong> silica gel plates. Their<br />
combinatorial library <strong>of</strong> up to 65,536 peptide dendrimers prepared<br />
on a photocleavable TentaGel resin was screened for ester<br />
hydrolysis activity <strong>and</strong> a fluorescent halo appeared after photocleavage<br />
from the beads around some <strong>of</strong> the beads on the silica<br />
plate, which were identified as hits. Using this “<strong>of</strong>f-bead” assay<br />
they report that the catalysis <strong>of</strong> the ester hydrolysis comes from<br />
core-active sites <strong>of</strong> the peptide dendrimers <strong>and</strong> depends on the<br />
presence <strong>of</strong> histidine residues at the dendritic core, but is also<br />
controlled by the outer dendritic branches, whereby anionic<br />
residues in these branches inhibit catalysis.<br />
A new method for screening S&P combinatorial libraries for<br />
catalytic activity was also reported by Stanton <strong>and</strong> co-workers 135<br />
using a spatial separation <strong>of</strong> resin-bound catalyst on an adhesive<br />
array on a microscope slide for catalytic Knoevenagel condensation<br />
screening. A visible absorption difference between react<strong>and</strong>s<br />
<strong>and</strong> products was used to site-selectively detect product formation<br />
in <strong>and</strong> around the catalytic beads. The method can be<br />
extended for colorless products by application <strong>of</strong> pH-sensitive<br />
indicators for array-based screening. Organic inorganic hybrid<br />
catalysts prepared by the coupling <strong>of</strong> 3-(triethoxysilyl)propyltriphenyl-phosphonium<br />
bromide <strong>and</strong> mesoporous silica enabled<br />
Sakai et al. 136 to prepare cyclic carbonates from carbon dioxide<br />
<strong>and</strong> epoxides. The pore size <strong>of</strong> silica was found to be important<br />
for catalysis. Organic <strong>and</strong> inorganic moieties had a synergistic<br />
effect on the catalytic activity, both the phosphonium bromide<br />
but also the OH group participating in catalysis.<br />
3.1.3. Heterogeneous Catalysts. Heterogeneous catalysis utilizing<br />
the S&P method relies on solid supports. For the design <strong>of</strong><br />
heterogeneous catalyst libraries numerous strategies have been<br />
developed (see, for instance, refs 137 <strong>and</strong> 138) Fundamental<br />
problem in the application <strong>of</strong> combinatorial or high-throughput<br />
methods in the development <strong>and</strong> optimization <strong>of</strong> heterogeneous<br />
catalysts is the number <strong>of</strong> possible elements <strong>of</strong> the periodic<br />
system. The combinatorial number <strong>of</strong> screenable element combinations<br />
scales with the number <strong>of</strong> components <strong>and</strong> generated<br />
the need <strong>of</strong> very high throughput methods, such as the S&P<br />
concept <strong>of</strong> solid support bead impregnation mentioned above.<br />
Solids are even more complex <strong>and</strong> to specify them completely,<br />
not only the composition has to be given but also crystallinity,<br />
polymorphic form, defect type <strong>and</strong> concentration, domain, <strong>and</strong><br />
interface characteristics, <strong>and</strong> much more information is needed<br />
for a single solid catalyst.<br />
To reduce the efforts <strong>and</strong> costs necessary to screen all possible<br />
catalyst c<strong>and</strong>idates, computational methods have been introduced<br />
for the prediction <strong>of</strong> solid catalyst performance including<br />
an electronic infrastructure as informatics toolbox for highthroughput<br />
kinetic studies <strong>and</strong> virtual screening. The latter was<br />
developed by Farrusseng, Mirodatos <strong>and</strong> co-workers for the<br />
TOPCOMBI research project funded by the European Commission<br />
for Nanotechnology <strong>and</strong> Nanoscience. 139 It represents a<br />
collection <strong>of</strong> modules dealing with laboratory analytics, robotics,<br />
data h<strong>and</strong>ling <strong>and</strong> analytics, optimization, in-database processing<br />
<strong>and</strong> visualization generated collegially by the partners <strong>of</strong> a<br />
consortium. It was developed for large or contract research<br />
organizations to control cost, build or strengthen their competitive<br />
advantages <strong>and</strong> disseminate knowledge from centers <strong>of</strong><br />
expertise <strong>of</strong> scattered or not interacting groups within the<br />
company. Knowledge generally is regarded to reduce screening<br />
efforts as quantitative structure activity relations (QSAR) correlate<br />
the characteristics <strong>of</strong> heterogeneous catalysts <strong>and</strong> their<br />
experimentally measured performances for certain reactions.<br />
As stated by Sch€uth et al., 140 unfortunately, the classical QSAR<br />
approach cannot be directly transferred to the discovery <strong>of</strong><br />
materials, especially <strong>of</strong> heterogeneous catalysts, because <strong>of</strong> the<br />
complexity <strong>of</strong> solid materials, that can hardly be characterized on<br />
atomic scale as a fingerprint signature similar to molecular<br />
fingerprint signatures. For molecules, a similarity principle holds<br />
according to which the similarity between two molecules can be<br />
assessed by measuring a distance between them. The quantification<br />
<strong>of</strong> distances between molecules is the key issue in the QSAR<br />
approach. Unfortunately, this similarity principle is not valid for<br />
solid catalysts. For materials, physicochemical properties encoded<br />
in a descriptor vector have to be used instead <strong>of</strong> distances.<br />
The key descriptors are usually not known or can hardly be<br />
measured in the case <strong>of</strong> diverse libraries.<br />
The principle <strong>of</strong> the advanced QSAR approach in heterogeneous<br />
catalysis is illustrated in Figure 15. 140 A minimal set <strong>of</strong><br />
attributes is defined that can correctly classify a data set into its<br />
592 dx.doi.org/10.1021/co200007w |ACS Comb. Sci. 2011, 13, 579–633