Ensembl Compara - CNB - Protein Design Group
Ensembl Compara - CNB - Protein Design Group
Ensembl Compara - CNB - Protein Design Group
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
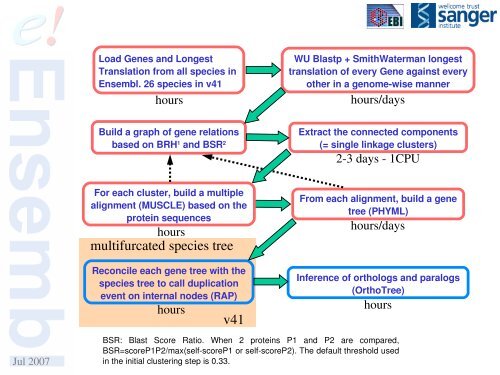
Load Genes and LongestTranslation from all species in<strong>Ensembl</strong>. 26 species in v41hoursWU Blastp + SmithWaterman longesttranslation of every Gene against everyother in a genomewise mannerhours/daysBuild a graph of gene relationsbased on BRH 1 and BSR 2Extract the connected components(= single linkage clusters)23 days 1CPUFor each cluster, build a multiplealignment (MUSCLE) based on theprotein sequenceshoursmultifurcated species treeReconcile each gene tree with thespecies tree to call duplicationevent on internal nodes (RAP)hoursv41From each alignment, build a genetree (PHYML)hours/daysInference of orthologs and paralogs(OrthoTree)hoursJul 2007BSR: Blast Score Ratio. When 2 proteins P1 and P2 are compared,BSR=scoreP1P2/max(selfscoreP1 or selfscoreP2). The default threshold usedin the initial clustering step is 0.33.