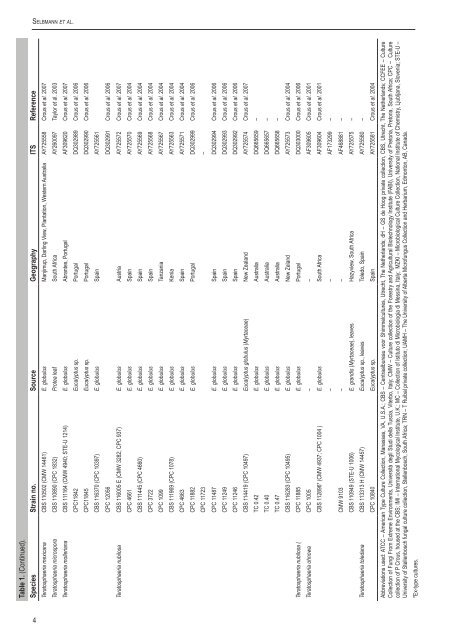
Se l b m a n n e t a l.Table 1. (Continued).Species Strain no. Source Geography ITS ReferenceTeratosphaeria mexicana <strong>CBS</strong> 110502 (CMW 14461) E. globulus Manjimup, Darling View, Plantation, Western Australia AY725558 Crous et al. 2007Teratosphaeria microspora <strong>CBS</strong> 110890 (CPC 1832) Protea leaf South Africa AY260097 Taylor et al. 2003Teratosphaeria molleriana <strong>CBS</strong> 111164 (CMW 4940; STE-U 1214) E. globulus Abrantes, Portugal AF309620 Crous et al. 2007CPC11842 Eucalyptus sp. Portugal DQ302989 Crous et al. 2006CPC11845 Eucalyptus sp. Portugal DQ302990 Crous et al. 2006<strong>CBS</strong> 116370 (CPC 10397) E. globulus Spain AY725561CPC 12056 DQ302991 Crous et al. 2006Teratosphaeria nubilosa <strong>CBS</strong> 116005 E (CMW 3282; CPC 937) E. globulus Austria AY725572 Crous et al. 2007CPC 4661 E. globulus Spain AY725570 Crous et al. 2004<strong>CBS</strong> 111445 (CPC 4660) E. globulus Spain AY725569 Crous et al. 2004CPC 3722 E. globulus Spain AY725568 Crous et al. 2004CPC 1099 E. globulus Tanzania AY725567 Crous et al. 2004<strong>CBS</strong> 111969 (CPC 1078) E. globulus Kenia AY725563 Crous et al. 2004CPC 4663 E. globulus Spain AY725571 Crous et al. 2004CPC 11882 E. globulus Portugal DQ302999 Crous et al. 2006CPC 11723 –CPC 11487 E. globulus Spain DQ302994 Crous et al. 2006CPC 11249 E. globulus Spain DQ302993 Crous et al. 2006CPC 11246 E. globulus Spain DQ302992 Crous et al. 2006<strong>CBS</strong> 114419 (CPC 10497) Eucalyptus globulus (Myrtaceae) New Zealand AY725574 Crous et al. 2007TC 0.42 E. globulus Australia DQ665659 –TC 0.40 E. globulus Australia DQ665657 –TC 0.47 E. globulus Australia DQ665658 –<strong>CBS</strong> 116283 (CPC 10495) E. globulus New Zeland AY725573 Crous et al. 2004Teratosphaeria nubilosa ( CPC 11885 E. globulus Portugal DQ303000 Crous et al. 2006Teratosphaeria ohnowa CPC 1005 – – AF309605 Crous et al. 2001<strong>CBS</strong> 112896* (CMW 4937; CPC 1004 ) E. globulus South Africa AF309604 Crous et al. 2001– – AF173299 –CMW 9103 – – AF468881 –<strong>CBS</strong> 110949 (STE-U 1006) E. grandis (Myrtaceae), leaves Hazyview, South Africa AY725575 –Teratosphaeria toledana <strong>CBS</strong> 113313 H (CMW 14457) Eucalyptus sp., leaves Toledo, Spain AY725580 –CPC 10840 Eucalyptus sp. Spain AY725581 Crous et al. 2004Abbreviations used: ATCC – American Type Culture Collection, Manassas, VA, U.S.A.; <strong>CBS</strong> – Centraalbureau voor Shimmelcultures, Utrecht, The Netherlands; dH – GS de Hoog private collection, <strong>CBS</strong>, Utrecht, The Netherlands; CCFEE – CultureCollection of Fungi From Extreme Environments, Università degli Studi della Tuscia, Viterbo, Italy; CMW – Culture collection of the Forestry and Agricultural Biotechnology Institute (FABI), University of Pretoria, Pretoria, South Africa; CPC – Culturecollection of P Crous, housed at the <strong>CBS</strong>; IMI – International Mycological Institute, U.K.; MC – Collection of Istituto di Microbiologia di Messina, Italy; MZKI – Microbiological Culture Collection, National Institute of Chemistry, Ljubljana, Slovenia; STE-U –University of Stellenbosch <strong>fungal</strong> culture collection, Stellenbosch, South Africa; TRN – T Ruibal private collection; UAMH – The University of Alberta Microfungus Collection and Herbarium, Edmonton, AB, Canada.*Ex-type cultures.4
Dr o u g h t m e e t s a c i d2003/2004, and from rock samples collected in Mount Aconcagua,Andes, Argentina, and Monte Rosa in the Alps, Italy, as reportedin Table 1. The samples were collected using a sterile chisel,sterlilised in the field before each sampling, and preserved in sterilebags at –20 °C. To remove potential contaminants, rocks collectedin other environments than Antarctica were washed 15 min insterile physiological solution added with 0.1 % Tween 20 (Sigma- Aldrich, Munich, Germany) and rinsed 4 times with physiologicalsolution to remove any trace of the detergent. Isolations from rockswere performed by powdering the samples and seeding fragmentsin triplicate Petri dishes filled with 2 % malt extract agar (MEA,AppliChem, GmbH) and dichloran-rose bengal agar (DRBC, OxoidLtd., Basingstoke, Hampshire, U.K.). Media were supplementedwith chloramphenicol 100 ppm to prevent bacterial growth. Fungifrom lichens were isolated as follows: small fragments of thalli weresterilised by washing with H 2O 2(8 %) for 5 min, washed with steriledeionised water to remove any trace of H 2O 2and finally seededon MEA and DRBC. Plates were incubated at 5 and 15 °C andinspected every 15 d until no new colonies appeared. Colonieswere transferred to MEA slants and incubated at 15 °C. Strainsanalysed for comparison are listed in Table 1; they were taken fromreference collections of the Culture Collection of Fungi from ExtremeEnvironments (CCFEE, Viterbo, Italy) and the Centraalbureau voorSchimmelcultures (<strong>CBS</strong>, Utrecht, The Netherlands), including alarge set of strains sampled by P.W. Crous from plants with leatheryleaves in a semi-arid climate.MorphologyHyphal maturation and conidiogenesis were studied using bothlight and scanning electron microscope (SEM). Slide cultureswere seeded onto MEA, incubated for 10 wk and mounted in lacticacid. Samples for SEM observations were prepared according tomethods described by Onofri et al. (1980).DNA extraction and sequencingDNA was extracted from mycelial fragments taken from 6-mos-oldMEA slants grown at 10 °C, using Nucleospin Plant kit (Macherey-Nagel, Düren, Germany) following the protocol optimised forfungi. PCR reactions were performed using BioMix (BioLineGmbH, Luckenwalde, Germany). In each 25 µL reaction tube 5pmol of each primer and 40 ng on template DNA were added. Theamplification was carried out using MiniCycler (MJ Research,Waltham, Massachusetts, U.S.A.) equipped with a heated lid.The first denaturation step at 95 °C for 3 min was followed by:denaturation at 95 °C for 2 s, annealing at 55 °C for 30 s, extensionat 72 °C for 30 s. The last three steps were repeated 35 times,with a last extension 72 °C for 5 min. The products were purifiedusing Nucleospin Extract kit (Macherey-Nagel, Düren, Germany).Primers NS1, NS2, NS3, NS4, NS5, NS8, ITS1, ITS4 (White etal. 1990), SR10R (Bruns et al. 1992), ITS5, and ITS4a (Larenaet al. 1999) were employed to amplify SSU and ITS rDNAportions. Sequencing reactions were performed according to thedideoxynucleotide method (Sanger et al. 1977) using the TF BigDye Terminator 1,1 RR kit (Applied Biosystems). Fragments wereanalysed using an ABI 310 Genetic Analyser (Applied Biosystems).Sequence assembly was done using the software Chromas (v. 1.451996–1998, Conor McCarthy School of Health Science, GriffithUniversity, Southport, Queensland, Australia).www.studiesinmycology.orgAlignment and tree reconstructionSSU sequences were aligned with ARB beta-package (v. 22-08-2003, Ludwig et al. 2004; www.mikro.biologie.tu-muenchen.de/pub/ARB). The SSU alignment spanned positions 141–2512,which corresponds to 1515 bp with reference to Saccharomycescerevisiae. Trees based on SSU sequences were reconstructedwith neighbour-joining in ARB.ITS sequences were aligned iteratively with Ward’s averaging(Van Ooyen 2002) in a research data base of black yeasts presentat <strong>CBS</strong> using the BioNumerics package (Applied Maths, Kortrijk,Belgium). Due to gaps necessary for alignment, the ITS1 domainspanned 187 positions (real lengths 147–154 bp), the 5.8S gene156 positions and the ITS2 domain 184 positions (real lengths143–155 bp). The alignments were based on the positions 28–481,the initial and the final parts were cut off to compare fragmentswith the same length. Alignments were exported and the best-fitsubstitution model was determined using Modeltest MrAic.pl 1.4.3(Nylander 2004, program distributed by the author) estimatedusing Ph y m l (Guindon & Gascuel 2003) through hierarchicallikelihood ratio tests. MrAic calculates the Akaike InformationCriterion (AIC), corrected Akaike Information Criterion (AICc)and Bayesian Information Criterion (BIC); Akaike weights fornucleotide substitution model and model uncertainty. All 56 modelsimplemented in Modeltest were evaluated. Phylogenetic trees werereconstructed by Maximum Likelihood, using Tr e e f i n d e r (Jobbet al. 2004) and the resulting tree was displayed using Tr e e v i e wv. 1.6.6 (Page 1996). The robustness of the phylogenetic inferencewas estimated using the bootstrap method (Felsenstein 1985) with100 pseudoreplicates generated and analysed with Tr e e f i n d e r.As alignment over the entire complex was highly ambiguous, analgorithm for tree reconstruction without alignment, the DNA-walkDivergence method (DNAWD, Licinio & Caligiorne 2004; Caligiorneet al. 2005) was used, involving the entire spacer region. DNAwalksare defined by incrementing walk steps for each nucleotide inthe sequence (for example a positive step for purines, and negativefor pyrimidines). It makes simultaneous comparisons of the threedimensionalwalks (representing three composition skews): AG-TC, AC-TG, and AT-CG for each pair of sequences. One sequenceslides against the other until the minimum squared walk difference isfound, corresponding to a global alignment. This is then taken as ameasure of their distance since statistically independent mutationsand indels increase the mean square walk differences linearly. Theresulting distance matrices are then fed into the Kitsch program ofthe Phylip package (v. 3.572c, Felsenstein 1996).Cultural preferencesCultural characteristics and growth rates were recorded onPotato-Dextrose Agar (PDA), MEA, Czapek Dox Agar (CzA) andOatmeal Agar (OA). Strain <strong>CBS</strong> 119434 was incubated at 10 °C,strains <strong>CBS</strong> 122538, 122539 and 122540 at 20 °C and strains<strong>CBS</strong> 899.87, <strong>CBS</strong> 335.97, dH 12881, dH 11526 and dH 13081 at25 °C. The diameter of the colonies was recorded monthly. Testswere performed in triplicate.Temperature preferencesTemperature preferences for the strains <strong>CBS</strong> 119434, 122538,122539 and 122540 were tested by incubating them on MEA,5
- Page 1: Studies in Mycology 61 (2008)Black
- Page 4 and 5: Studies in MycologyThe Studies in M
- Page 7 and 8: PREFACEThe terms "black fungi" or "
- Page 9 and 10: available online at www.studiesinmy
- Page 11: Dr o u g h t m e e t s a c i dTable
- Page 15 and 16: Dr o u g h t m e e t s a c i din Pe
- Page 17 and 18: Dr o u g h t m e e t s a c i d99100
- Page 19 and 20: Dr o u g h t m e e t s a c i dAnamo
- Page 21 and 22: Dr o u g h t m e e t s a c i dFig.
- Page 23 and 24: Dr o u g h t m e e t s a c i dFig.
- Page 25 and 26: Dr o u g h t m e e t s a c i dconve
- Page 27 and 28: Dr o u g h t m e e t s a c i dal. 2
- Page 29: available online at www.studiesinmy
- Page 32 and 33: Za l a r e t a l.Table 1. List of a
- Page 34 and 35: Za l a r e t a l.Table 1. (Continue
- Page 36 and 37: Za l a r e t a l.Fig. 2. Consensus
- Page 38 and 39: Za l a r e t a l.Fig. 4. Macromorph
- Page 40 and 41: Za l a r e t a l.Fig. 5. Aureobasid
- Page 42 and 43: Za l a r e t a l.Fig. 7. Aureobasid
- Page 44 and 45: Za l a r e t a l.the type species o
- Page 46 and 47: Za l a r e t a l.Vishniac HS, Onofr
- Page 48 and 49: Yu r l o va e t a l.Table 1. Strain
- Page 50 and 51: Yu r l o va e t a l.112ОС2Fig. 1.
- Page 52 and 53: Yu r l o va e t a l.RESULTSFourteen
- Page 54 and 55: Yu r l o va e t a l.100908070605040
- Page 56 and 57: Yu r l o va e t a l.AcetateMelanin(
- Page 58 and 59: available online at www.studiesinmy
- Page 60 and 61: Fat t y-a c i d-m o d if y i n g e
- Page 62 and 63:
Fat t y-a c i d-m o d if y i n g e
- Page 64 and 65:
Fat t y-a c i d-m o d if y i n g e
- Page 66 and 67:
Fat t y-a c i d-m o d if y i n g e
- Page 68 and 69:
Va u p o t i č e t a l.cerevisiae)
- Page 70 and 71:
Va u p o t i č e t a l.Fig. 3. Mor
- Page 72 and 73:
Va u p o t i č e t a l.HMG-CoA red
- Page 74 and 75:
Pl e m e n i ta s e t a l.entire en
- Page 76 and 77:
Pl e m e n i ta s e t a l.glycerol
- Page 78 and 79:
Pl e m e n i ta s e t a l.Fig. 1. T
- Page 80 and 81:
Pl e m e n i ta s e t a l.diversity
- Page 82 and 83:
available online at www.studiesinmy
- Page 84 and 85:
Ti n e a n i g r aFig. 1. Map showi
- Page 86 and 87:
Ti n e a n i g r aand Polynesia (Ue
- Page 88 and 89:
available online at www.studiesinmy
- Page 90 and 91:
Bl a c k f u n g i in l i ch e n st
- Page 92 and 93:
Bl a c k f u n g i in l i ch e n sD
- Page 94 and 95:
Bl a c k f u n g i in l i ch e n sF
- Page 96 and 97:
available online at www.studiesinmy
- Page 98 and 99:
Ce l l u l a r r e s p o n s e s t
- Page 100 and 101:
Ce l l u l a r r e s p o n s e s t
- Page 102 and 103:
Ce l l u l a r r e s p o n s e s t
- Page 104 and 105:
On o f r i e t a l.Table 1. Test pa
- Page 106 and 107:
On o f r i e t a l.Analyses of resp
- Page 108 and 109:
On o f r i e t a l.Fig. 6. Percenta
- Page 110 and 111:
On o f r i e t a l.Fig. 11. Percent
- Page 112 and 113:
On o f r i e t a l.2005). The abili
- Page 114 and 115:
available online at www.studiesinmy
- Page 116 and 117:
A r o c k-i n h a b i t i n g a n c
- Page 118 and 119:
A r o c k-i n h a b i t i n g a n c
- Page 120 and 121:
A r o c k-i n h a b i t i n g a n c
- Page 122 and 123:
A r o c k-i n h a b i t i n g a n c
- Page 124 and 125:
De n g e t a l.Table 1. Strains use
- Page 126 and 127:
De n g e t a l.Table 3. Primer sequ
- Page 128 and 129:
De n g e t a l.10098CBS 157.67 Exop
- Page 130 and 131:
De n g e t a l.Table 5. Number of m
- Page 132 and 133:
available online at www.studiesinmy
- Page 134 and 135:
Co n i o s p o r i u m o n s k i nT
- Page 136 and 137:
Co n i o s p o r i u m o n s k i nC
- Page 138 and 139:
available online at www.studiesinmy
- Page 140 and 141:
e n v i r o nm e n ta l is o l at i
- Page 142 and 143:
e n v i r o nm e n ta l is o l at i
- Page 144 and 145:
e n v i r o nm e n ta l is o l at i
- Page 146 and 147:
available online at www.studiesinmy
- Page 148 and 149:
Origin o f Ex o p h i a l a d e r m
- Page 150 and 151:
Origin o f Ex o p h i a l a d e r m
- Page 152 and 153:
Origin o f Ex o p h i a l a d e r m
- Page 154 and 155:
Origin o f Ex o p h i a l a d e r m
- Page 156 and 157:
Origin o f Ex o p h i a l a d e r m
- Page 158 and 159:
Sat o w e t a l.Table 1. Compositio
- Page 160 and 161:
Sat o w e t a l.Assimilation of min
- Page 162 and 163:
Sat o w e t a l.pollutants in biore
- Page 164 and 165:
available online at www.studiesinmy
- Page 166 and 167:
s u b s t r at e s p e c i f i c i
- Page 168 and 169:
s u b s t r at e s p e c i f i c i
- Page 170 and 171:
s u b s t r at e s p e c i f i c i
- Page 172 and 173:
s u b s t r at e s p e c i f i c i
- Page 174 and 175:
available online at www.studiesinmy
- Page 176 and 177:
Bi o d i v e rs i t y o f t h e g e
- Page 178 and 179:
Bi o d i v e rs i t y o f t h e g e
- Page 180 and 181:
Bi o d i v e rs i t y o f t h e g e
- Page 182 and 183:
Bi o d i v e rs i t y o f t h e g e
- Page 184 and 185:
Bi o d i v e rs i t y o f t h e g e
- Page 186 and 187:
Bi o d i v e rs i t y o f t h e g e
- Page 188 and 189:
Bi o d i v e rs i t y o f t h e g e
- Page 190 and 191:
Bi o d i v e rs i t y o f t h e g e
- Page 192 and 193:
homogentisate 165, 171, 173Hortaea
- Page 194 and 195:
CBS Biodiversity SeriesNo. 6: Alter
- Page 196 and 197:
Studies in Mycology (ISSN 0166-0616
- Page 198:
Satow MM, Attili-Angelis D, Hoog GS