reSolution_LNT_No1_en - Leica Microsystems
reSolution_LNT_No1_en - Leica Microsystems
reSolution_LNT_No1_en - Leica Microsystems
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
BIOLOGY<br />
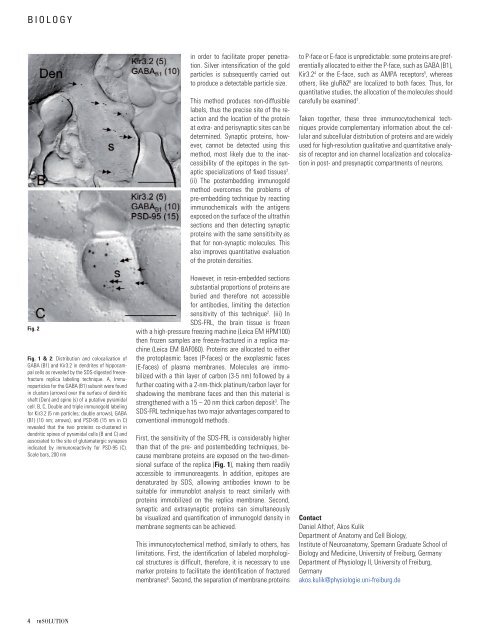
Fig. 2<br />
Fig. 1 & 2: Distribution and colocalization of<br />
GABA (B1) and Kir3.2 in d<strong>en</strong>drites of hippocampal<br />
cells as revealed by the SDS-digested freezefracture<br />
replica labeling technique. A, Immunoparticles<br />
for the GABA (B1) subunit were found<br />
in clusters (arrows) over the surface of d<strong>en</strong>dritic<br />
shaft (D<strong>en</strong>) and spine (s) of a putative pyramidal<br />
cell. B, C, Double and triple immunogold labeling<br />
for Kir3.2 (5 nm particles; double arrows), GABA<br />
(B1) (10 nm; arrows), and PSD-95 (15 nm in C)<br />
revealed that the two proteins co-clustered in<br />
d<strong>en</strong>dritic spines of pyramidal cells (B and C) and<br />
associated to the site of glutamatergic synapses<br />
indicated by immunoreactivity for PSD-95 (C).<br />
Scale bars, 200 nm<br />
4 reSOLUTION<br />
in order to facilitate proper p<strong>en</strong>etration.<br />
Silver int<strong>en</strong>sifi cation of the gold<br />
particles is subsequ<strong>en</strong>tly carried out<br />
to produce a detectable particle size.<br />
This method produces non-diffusible<br />
labels, thus the precise site of the reaction<br />
and the location of the protein<br />
at extra- and perisynaptic sites can be<br />
determined. Synaptic proteins, however,<br />
cannot be detected using this<br />
method, most likely due to the inaccessibility<br />
of the epitopes in the synaptic<br />
specializations of fi xed tissues 2 .<br />
(ii) The postembedding immunogold<br />
method overcomes the problems of<br />
pre-embedding technique by reacting<br />
immunochemicals with the antig<strong>en</strong>s<br />
exposed on the surface of the ultrathin<br />
sections and th<strong>en</strong> detecting synaptic<br />
proteins with the same s<strong>en</strong>sititvity as<br />
that for non-synaptic molecules. This<br />
also improves quantitative evaluation<br />
of the protein d<strong>en</strong>sities.<br />
However, in resin-embedded sections<br />
substantial proportions of proteins are<br />
buried and therefore not accessible<br />
for antibodies, limiting the detection<br />
s<strong>en</strong>sitivity of this technique 2 . (iii) In<br />
SDS-FRL, the brain tissue is froz<strong>en</strong><br />
with a high-pressure freezing machine (<strong>Leica</strong> EM HPM100)<br />
th<strong>en</strong> froz<strong>en</strong> samples are freeze-fractured in a replica machine<br />
(<strong>Leica</strong> EM BAF060). Proteins are allocated to either<br />
the protoplasmic faces (P-faces) or the exoplasmic faces<br />
(E-faces) of plasma membranes. Molecules are immobilized<br />
with a thin layer of carbon (3-5 nm) followed by a<br />
further coating with a 2-nm-thick platinum/carbon layer for<br />
shadowing the membrane faces and th<strong>en</strong> this material is<br />
str<strong>en</strong>gth<strong>en</strong>ed with a 15 – 20 nm thick carbon deposit 3 . The<br />
SDS-FRL technique has two major advantages compared to<br />
conv<strong>en</strong>tional immunogold methods.<br />
First, the s<strong>en</strong>sitivity of the SDS-FRL is considerably higher<br />
than that of the pre- and postembedding techniques, because<br />
membrane proteins are exposed on the two-dim<strong>en</strong>sional<br />
surface of the replica (Fig. 1), making them readily<br />
accessible to immunoreag<strong>en</strong>ts. In addition, epitopes are<br />
d<strong>en</strong>aturated by SDS, allowing antibodies known to be<br />
suitable for immunoblot analysis to react similarly with<br />
proteins immobilized on the replica membrane. Second,<br />
synaptic and extrasynaptic proteins can simultaneously<br />
be visualized and quantifi cation of immunogold d<strong>en</strong>sity in<br />
membrane segm<strong>en</strong>ts can be achieved.<br />
This immunocytochemical method, similarly to others, has<br />
limitations. First, the id<strong>en</strong>tifi cation of labeled morphological<br />
structures is diffi cult, therefore, it is necessary to use<br />
marker proteins to facilitate the id<strong>en</strong>tifi cation of fractured<br />
membranes 6 . Second, the separation of membrane proteins<br />
to P-face or E-face is unpredictable: some proteins are prefer<strong>en</strong>tially<br />
allocated to either the P-face, such as GABA (B1),<br />
Kir3.2 4 or the E-face, such as AMPA receptors 5 , whereas<br />
others, like gluRδ2 6 are localized to both faces. Thus, for<br />
quantitative studies, the allocation of the molecules should<br />
carefully be examined 7 .<br />
Tak<strong>en</strong> together, these three immunocytochemical techniques<br />
provide complem<strong>en</strong>tary information about the cellular<br />
and subcellular distribution of proteins and are widely<br />
used for high-resolution qualitative and quantitative analysis<br />
of receptor and ion channel localization and colocalization<br />
in post- and presynaptic compartm<strong>en</strong>ts of neurons.<br />
Contact<br />
Daniel Althof, Akos Kulik<br />
Departm<strong>en</strong>t of Anatomy and Cell Biology,<br />
Institute of Neuroanatomy, Spemann Graduate School of<br />
Biology and Medicine, University of Freiburg, Germany<br />
Departm<strong>en</strong>t of Physiology II, University of Freiburg,<br />
Germany<br />
akos.kulik@physiologie.uni-freiburg.de