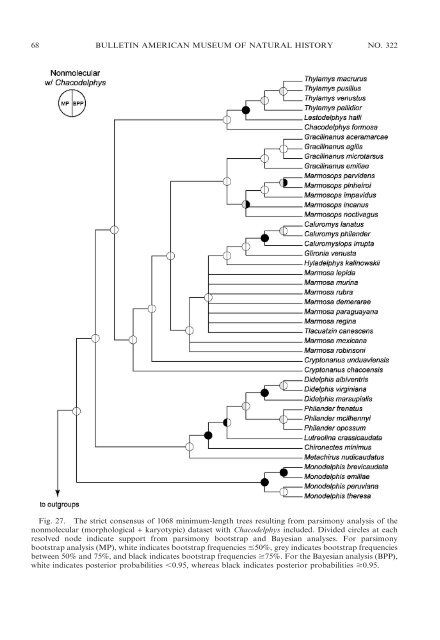
68 BULLETIN AMERICAN MUSEUM OF NATURAL HISTORY NO. 322 Fig. 27. The strict consensus <strong>of</strong> 1068 minimum-length trees resulting from parsimony analysis <strong>of</strong> the nonmolecular (morphological + karyotypic) dataset with Chacodelphys included. Divided circles at each resolved node indicate support from parsimony bootstrap <strong>and</strong> Bayesian analyses. For parsimony bootstrap analysis (MP), white indicates bootstrap frequencies #50%, grey indicates bootstrap frequencies between 50% <strong>and</strong> 75%, <strong>and</strong> black indicates bootstrap frequencies $75%. For the Bayesian analysis (BPP), white indicates posterior probabilities ,0.95, whereas black indicates posterior probabilities $0.95.
2009 VOSS AND JANSA: DIDELPHID MARSUPIALS 69 ly <strong>of</strong> Marmosops (only moderately supported by parsimony). However, Bayesian support is weak for several nodes with moderate parsimony bootstrap support, including the genus Thylamys, the genus Caluromys, <strong>and</strong> the genus Phil<strong>and</strong>er. Most <strong>of</strong> the obvious differences between these results <strong>and</strong> our previously published analyses <strong>of</strong> <strong>didelphid</strong> morphological character data are attributable to rooting. Whereas we previously (Voss <strong>and</strong> Jansa, 2003; Voss et al., 2004a; Jansa <strong>and</strong> Voss, 2005) rooted our estimates <strong>of</strong> didelphine <strong>relationships</strong> using Caluromys, Caluromysiops, <strong>and</strong> Glironia as outgroups, these analyses are rooted using non<strong>didelphid</strong> marsupial outgroups. Considered as an undirected network, the ingroup topology <strong>of</strong> figure 27 is almost identical to that obtained by Jansa <strong>and</strong> Voss (2005: fig. 1C) for the same set <strong>of</strong> <strong>didelphid</strong> terminal taxa. Single-gene Analyses Maximum-likelihood topologies with superimposed summaries <strong>of</strong> nodal statistics from three analyses (MP, ML, <strong>and</strong> Bayesian) <strong>of</strong> each single-gene dataset (figs. 28–32) illustrate similarities <strong>and</strong> differences in clade support among the five loci we sequenced. Because our analytic results for IRBP <strong>and</strong> DMP1 have already been published (Jansa <strong>and</strong> Voss, 2005), only summary comments are warranted here. Briefly, the monophyly <strong>of</strong> all genera represented by two or more terminal taxa is strongly supported by both <strong>of</strong> these genes (figs. 28, 29) with the exception <strong>of</strong> Gracilinanus (weakly to moderately supported by IRBP, not recovered as monophyletic by DMP1), Thylamys (only weakly supported by Bayesian analysis <strong>of</strong> DMP1), <strong>and</strong> Didelphis (inconsistently recovered as monophyletic by analyses <strong>of</strong> IRBP). On the assumption that the DMP1 tree (lacking outgroup terminals) is appropriately rooted within a node or two <strong>of</strong> Glironia, several patterns <strong>of</strong> intergeneric <strong>relationships</strong> are also supported by both genes, including the nested clades (Metachirus (Chironectes (Lutreolina (Didelphis + Phil<strong>and</strong>er)))); a group that includes Lestodelphys <strong>and</strong> Thylamys; a group that includes Cryptonanus, Gracilinanus, <strong>and</strong> Lestodelphys + Thylamys; a group that includes the latter four genera plus Marmosops; <strong>and</strong> a group that includes Marmosa, Monodelphis, <strong>and</strong> Tlacuatzin. As reported elsewhere (Jansa <strong>and</strong> Voss, 2005), there are no examples <strong>of</strong> hard incongruence to be found in comparing analytic results between IRBP <strong>and</strong> DMP1 because all conflicting nodes (e.g., those resolving the position <strong>of</strong> Hyladelphys) have weak support from one or both genes. Analyses <strong>of</strong> first <strong>and</strong> second codon positions <strong>of</strong> RAG1 (fig. 30) support the monophyly <strong>of</strong> most polytypic <strong>didelphid</strong> genera (except Caluromys, Didelphis, <strong>and</strong>Phil<strong>and</strong>er), but some genera recovered as monophyletic groups (e.g., Thylamys, Marmosops, <strong>and</strong> Monodelphis) do not receive consistently strong support. Many <strong>of</strong> the same intergeneric <strong>relationships</strong> supported by IRBP <strong>and</strong> DMP1 were also recovered, notably including the branching patterns among Metachirus, Chironectes, Lutreolina, <strong>and</strong> Phil<strong>and</strong>er + Didelphis. Additionally, Marmosops was recovered as the sister group to a clade that includes Cryptonanus, Gracilinanus, <strong>and</strong> Thylamys. Unfortunately, the uncertain position <strong>of</strong> the root (probably an artifact <strong>of</strong> incomplete outgroup sequences; see above) is reflected in the lack <strong>of</strong> strong support for any basal <strong>relationships</strong> in this topology. Analyses <strong>of</strong> sequence data from BRCA1 produced the most compelling single-gene estimate <strong>of</strong> <strong>didelphid</strong> <strong>relationships</strong> recovered to date, with strikingly consistent high nodal support values <strong>and</strong> an almost completely resolved consensus topology (fig. 31). Among other salient features, every polytypic opossum genus with the exception <strong>of</strong> Didelphis was recovered with strong support, as were almost all <strong>of</strong> the higher-level clades recovered with strong support by any analysis <strong>of</strong> other single-gene datasets (the only exceptions are groups that include Lestodelphys <strong>and</strong> Caluromysiops, taxa from which BRCA1 sequences are not available). Three <strong>of</strong> these merit particular attention. One is ‘‘clade H’’ <strong>of</strong> Jansa <strong>and</strong> Voss (2000), which unites the Thylamys cluster with Metachirus <strong>and</strong> the large opossums with 22 chromosomes (Chironectes, Lutreolina, Phil<strong>and</strong>er, <strong>and</strong> Didelphis). Another is the subfamily Didelphinae <strong>of</strong> traditional usage (excluding Hyladelphys, Caluromys, <strong>and</strong>Glironia), <strong>and</strong>