Hap Map Project - Bgbunict.it
Hap Map Project - Bgbunict.it
Hap Map Project - Bgbunict.it
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
The International <strong>Hap</strong><strong>Map</strong><br />
<strong>Project</strong><br />
Anno 2009/2010<br />
Dott.ssa Laura R<strong>it</strong>a Duro
Most common diseases, such as diabetes,<br />
cancer, stroke, heart disease, depression, and<br />
asthma, are affected by combinations of multiple<br />
genetic and environmental factors
Genetic and environmental contributions to monogenic and<br />
complex disorders<br />
(A) Monogenic disease. A variant in a single gene is the primary determinant of a<br />
monogenic disease or tra<strong>it</strong>, responsible for most of the disease risk or tra<strong>it</strong> variation<br />
(dark blue sector), w<strong>it</strong>h possible minor contributions of modifier genes (yellow<br />
sectors) or environment (light blue sector).<br />
(B) Complex disease. Many variants of small effect (yellow sectors) contribute to<br />
disease risk or tra<strong>it</strong> variation, along w<strong>it</strong>h many environmental factors (blue sector).
More than a<br />
thousand genes for<br />
rare, highly her<strong>it</strong>able<br />
‘mendelian’ disorders<br />
have been identified,<br />
in which variation in a<br />
single gene is both<br />
necessary and<br />
sufficient to cause<br />
disease.<br />
Complex diseases, in<br />
contrast, have proven<br />
much more<br />
challenging to study,<br />
as they are thought to<br />
be due to the<br />
combined effect of<br />
many different<br />
susceptibil<strong>it</strong>y DNA<br />
variants interacting<br />
w<strong>it</strong>h environmental<br />
factors
Discovering these genetic factors will provide<br />
fundamental new insights into the pathogenesis,<br />
diagnosis and treatment of human disease
Although any two unrelated people are the<br />
same at about 99.9% of their DNA sequences,<br />
the remaining 0.1% is important because <strong>it</strong><br />
contains the genetic variants that influence<br />
how people differ in their risk of disease or<br />
their response to drugs.<br />
Discovering the DNA sequence variants that<br />
contribute to common disease risk offers one<br />
of the best opportun<strong>it</strong>ies for understanding the<br />
complex causes of disease in humans.
Human Genetic Variations<br />
Primarily two types of genetic mutation events create all<br />
forms of variations:<br />
Single base mutation which subst<strong>it</strong>utes<br />
one nucleotide for another<br />
-Single Nucleotide Polymorphisms (SNP)<br />
Insertion or deletion of one or more<br />
nucleotide(s)<br />
-Tandem Repeat Polymorphisms<br />
-Insertion/Deletion Polymorphisms
Tandem Repeat Polymorphisms<br />
Tandem repeats or variable number of tandem repeats (VNTR) are a very<br />
common class of polymorphism, consisting of variable length of sequence<br />
motifs that are repeated in tandem in a variable copy number.<br />
VNTRs are subdivided into two subgroups based on the size of the<br />
tandem repeat un<strong>it</strong>s.<br />
Microsatell<strong>it</strong>es or Short Tandem Repeat (STR)<br />
repeat un<strong>it</strong>: 1-6 (dinucleotide repeat: CACACACACACA)<br />
Minisatell<strong>it</strong>es<br />
repeat un<strong>it</strong>: 10-100
SNPs<br />
S<strong>it</strong>es in the genome where the<br />
DNA sequences of many<br />
individuals differ by a single<br />
base are called single<br />
nucleotide polymorphisms<br />
(SNPs)<br />
For example, some people<br />
may have a chromosome w<strong>it</strong>h<br />
an A at a particular s<strong>it</strong>e where<br />
others have a chromosome<br />
w<strong>it</strong>h a G<br />
Each form is called an allele
Variation Or Mutation ?<br />
Terminology for variation at a single<br />
nucleotide pos<strong>it</strong>ion is defined by allele<br />
frequency
Polymorphism<br />
A sequence variation that occurs at least 1<br />
percent of the time (> 1%)<br />
90% of variations are SNPs<br />
Mutation<br />
If the variation is<br />
present less than<br />
1 percent of the<br />
time (
Trans<strong>it</strong>ions and Transversions<br />
SNPs include single base subst<strong>it</strong>utions such as:<br />
Trans<strong>it</strong>ions:<br />
change of one purine (A,G) for a purine,<br />
or a pyrimidine (C,T) for a pyrimidine<br />
A G G A C T T C<br />
Transversions:<br />
change of a purine (A,G) for a pyrimidine (C,T),<br />
or viceversa<br />
A C A T G C G T C A C G T A T G
In principle, SNPs could be bi-, tri-, or tetra-allelic<br />
polymorphisms<br />
However, in humans, tri-allelic and tetra-allelic<br />
However, in humans, tri-allelic and tetra-allelic<br />
SNPs are rare almost to the point of<br />
non-existence, and so SNPs are sometimes<br />
simply referred to as bi-allelic markers
Non-coding SNPs:<br />
5’ and 3’ UTRs<br />
Introns<br />
Intergenic Spaces<br />
Non-synonymous Coding<br />
SNPs:<br />
when single base subst<strong>it</strong>utions<br />
cause a change in the resultant<br />
amino acid<br />
Synonymous Coding SNPs:<br />
when single base subst<strong>it</strong>utions do<br />
not cause a change in the<br />
resultant amino acid
Non-coding SNPs<br />
Example: Regulatory SNPs (rSNPs)<br />
Two allelic variants of the same gene are transcribed in different<br />
amounts as a consequence of an adjacent polymorphism. In this<br />
example, allele G, located upstream of the gene, has a higher<br />
transcript level than does allele T.
Coding SNPs<br />
Example: Synonymous, mutation does not change<br />
amino acid.
Coding SNPs<br />
Example: Non-synonymous, mutation change<br />
amino acid.
SNPs<br />
It has been estimated that, in the world’s human population,<br />
about 10 million s<strong>it</strong>es (that is, one variant per 300 bases on<br />
average) vary such that both alleles are observed at a<br />
frequency of > 1%, and that these 10 million common SNPs<br />
const<strong>it</strong>ute 90% of the variation in the population.<br />
The remaining 10% is due to a vast array of variants that are<br />
each rare in the population.<br />
The presence of particular SNP alleles in an individual is<br />
determined by testing (‘genotyping’) a genomic DNA sample.<br />
NATURE |VOL 426 | 18/25 DECEMBER 2003
A particular combination of alleles along a<br />
chromosome is termed a haplotype<br />
<strong>Hap</strong>lotype is a set of SNPs on a single chromatid<br />
that are statistically associated
The coinher<strong>it</strong>ance of SNP alleles on these haplotypes<br />
leads to associations between these alleles in the<br />
population<br />
(known as linkage disequilibrium, LD)
Linkage disequilibrium<br />
<br />
S<strong>it</strong>uation in which some combinations of alleles or genetic<br />
markers occur more or less frequently in a population than<br />
would be expected from a random formation of haplotypes<br />
from alleles based on their frequencies.<br />
Non-random associations between polymorphisms at<br />
different loci are measured by the degree of linkage<br />
disequilibrium (LD).
The LD between many neighboring SNPs generally persists because meiotic recombination<br />
does not occur at random, but is concentrated in recombination hot spots.<br />
Adjacent SNPs that lack a hot spot between them are likely to be in strong LD.<br />
r 2 = 1: two SNPs that are perfectly correlated (allele A of SNP1 is always observed w<strong>it</strong>h<br />
allele C of SNP2, and viceversa)<br />
r 2 = 0: allele A of SNP1 providing no information at all about which allele of SNP4 is<br />
present.<br />
Complete independence of these 6 SNPs would predict the possibil<strong>it</strong>y of 64 different<br />
haplotypes (because n biallelic SNPs could generate 2 n haplotypes), but in real<strong>it</strong>y just 4<br />
haplotypes comprise 90% of observed chromosomes, indicating that LD is present.<br />
Because of the strong associations<br />
among the SNPs in most chromosomal<br />
regions, only a few carefully chosen<br />
SNPs (known as tag SNPs) need to be<br />
typed to predict the likely variants at the<br />
rest of the SNPs in each region<br />
SNP1, SNP2, and SNP3 are strongly correlated, and SNP4, SNP5, and SNP6<br />
are strongly correlated, so that any of SNP1–SNP3 (or SNP4–SNP6) could<br />
serve as tags for the other 2 SNPs in each group.
Many empirical studies have shown highly significant levels of LD, and<br />
often strong associations between nearby SNPs, in the human genome.<br />
Because the likelihood of recombination between two SNPs increases<br />
w<strong>it</strong>h the distance between them, on average such associations between<br />
SNPs decline w<strong>it</strong>h distance.<br />
Average linkage disequilibrium, |D|, vs.<br />
distance between SNPs for 2597<br />
genes in which accurate distances<br />
were available.<br />
Lower values indicate a stronger effect<br />
of recombination and recurrent<br />
mutation.<br />
LD decreases w<strong>it</strong>h distance.<br />
B.A. Salisbury et al. Mutation Research 2003
The strong<br />
associations between<br />
SNPs in a region have<br />
a practical value<br />
Genotyping only a few, carefully chosen SNPs in the region will provide enough<br />
information to predict much of the information about the remainder of the common<br />
SNPs in that region. As a result, only a few of these ‘tag’ SNPs are required to<br />
identify each of the common haplotypes in a region.<br />
On the basis of empirical studies, <strong>it</strong> has been estimated that most of<br />
the information about genetic variation represented by the 10 million<br />
common SNPs in the population could be provided by genotyping<br />
200.000 to 1.000.000 tag SNPs across the genome<br />
These observations are the conceptual and empirical foundation for<br />
developing a haplotype map of the human genome, the ‘<strong>Hap</strong><strong>Map</strong>’.
The International <strong>Hap</strong><strong>Map</strong> <strong>Project</strong> is a partnership of scientists<br />
and funding agencies from Canada, China, Japan, Nigeria, the<br />
Un<strong>it</strong>ed Kingdom and the Un<strong>it</strong>ed States to develop a public resource<br />
that will help researchers find genes associated w<strong>it</strong>h human<br />
disease and response to pharmaceuticals.<br />
An in<strong>it</strong>ial meeting to discuss the scientific and ethical issues associated<br />
w<strong>it</strong>h developing a human haplotype map was held in Washington in 2001.<br />
The International <strong>Hap</strong><strong>Map</strong> <strong>Project</strong> was then formally in<strong>it</strong>iated in 2002.
The goal of the International <strong>Hap</strong><strong>Map</strong> <strong>Project</strong> is to develop a<br />
haplotype map of the human genome, the <strong>Hap</strong><strong>Map</strong>,<br />
which will describe the common patterns of human DNA<br />
sequence variation.<br />
The <strong>Hap</strong><strong>Map</strong> is expected to be a key resource for researchers<br />
to use to find genes affecting health, disease, and responses<br />
to drugs and environmental factors.<br />
The information produced by the <strong>Project</strong> is freely available<br />
(www.hapmap.org)<br />
NATURE |VOL 426 | 18/25 DECEMBER 2003
The <strong>Hap</strong><strong>Map</strong> was designed to determine the frequencies and<br />
patterns of association among roughly 3 million common<br />
SNPs in four populations, for use in genetic association<br />
studies<br />
The <strong>Hap</strong><strong>Map</strong> project focuses only on common SNPs, those<br />
where each allele occurs in at least 1% of the population
The project studied a total of 270 DNA samples:<br />
<br />
90 samples from a US Utah population w<strong>it</strong>h<br />
Northern and Western European ancestry<br />
(samples collected in 1980 by the Centre<br />
d’Etude du Polymorphisme Humain (CEPH)<br />
and used for other human genetic maps)<br />
new samples collected from 90 Yoruba<br />
people in Ibadan, Nigeria<br />
<br />
45 unrelated Japanese in Tokyo, Japan<br />
<br />
45 unrelated Han Chinese in Beijing, China
The International <strong>Hap</strong><strong>Map</strong> Consortium decided to include several<br />
populations from different ancestral geographic locations to ensure that the<br />
<strong>Hap</strong><strong>Map</strong> would include most of the common variation and some of the less<br />
common variation in different populations.<br />
NATURE |VOL 426 | 18/25 DECEMBER 2003
Human Genome <strong>Project</strong><br />
vs<br />
International <strong>Hap</strong><strong>Map</strong> <strong>Project</strong><br />
In <strong>it</strong>s scope and potential consequences, the International <strong>Hap</strong><strong>Map</strong> <strong>Project</strong><br />
has much in common w<strong>it</strong>h the Human Genome <strong>Project</strong>, which sequenced the<br />
human genome.<br />
Both projects have been scientifically amb<strong>it</strong>ious and technologically<br />
demanding, have involved intense international collaboration, have been<br />
dedicated to the rapid release of data into the public domain, and promise to<br />
have profound implications for our understanding of human biology and<br />
human health.<br />
Whereas the sequencing project covered the entire genome, including the<br />
99.9% of the genome where we are all the same, the <strong>Hap</strong><strong>Map</strong> will<br />
characterize the common patterns w<strong>it</strong>hin the 0.1% where we differ from each<br />
other.
The project had become practical by the confluence of the following:<br />
the availabil<strong>it</strong>y of the human genome sequence;<br />
databases of common SNPs (subsequently enriched by this<br />
project) from which genotyping assays could be designed;<br />
insights into human LD;<br />
development of inexpensive, accurate technologies for highthroughput<br />
SNP genotyping;<br />
web-based tools for storing and sharing data.<br />
The International <strong>Hap</strong><strong>Map</strong> Consortium NATURE October 2005
<strong>Hap</strong><strong>Map</strong> <strong>Project</strong> comprises two phases<br />
The complete data obtained<br />
in Phase I were published<br />
on October 2005.<br />
The analysis of the Phase II<br />
dataset was published in<br />
October 2007.
The Phase I <strong>Hap</strong><strong>Map</strong><br />
Phase I of the <strong>Hap</strong><strong>Map</strong> <strong>Project</strong> set as a<br />
goal genotyping at least one common<br />
SNP every 5 kb across the genome in<br />
each of 269 DNA samples.<br />
For the sake of practical<strong>it</strong>y, and motivated<br />
by the allele frequency distribution of<br />
variants in the human genome, a minor<br />
allele frequency (MAF) of 0.05 or greater<br />
was targeted for study.<br />
Minor Allele Frequency (MAF) : The frequency at which the less abundant<br />
(or minor) allele of a SNP is present in a population. The MAF for a SNP to<br />
be considered common is usually above 1%.
The project required a dense map of SNPs, ideally containing<br />
information about validation and frequency of each candidate SNP.<br />
When the project started, the public SNP<br />
database (dbSNP) contained 2.6 million<br />
candidate SNPs, few of which were<br />
annotated w<strong>it</strong>h the required information.<br />
The <strong>Hap</strong><strong>Map</strong> <strong>Project</strong> contributed about 6<br />
million new SNPs to dbSNP. At October<br />
2005 dbSNP contains 9.2 million<br />
candidate human SNPs.
To study patterns of genetic variation were selected ten 500-kb regions<br />
from the ENCODE (Encyclopedia of DNA Elements) <strong>Project</strong>.<br />
These ten regions were chosen to approximate the genome-wide<br />
average for G+C content, recombination rate, percentage of sequence<br />
conserved relative to mouse sequence, and gene dens<strong>it</strong>y.<br />
Each 500-kb region was sequenced in 48 individuals, and all SNPs in<br />
these regions (discovered or in dbSNP) were genotyped in the complete<br />
set of 269 DNA samples.
Using the data provided by <strong>Hap</strong><strong>Map</strong>, a team of scientists at Harvard Medical<br />
School and the Broad Inst<strong>it</strong>ute has discovered a new genetic variant<br />
associated w<strong>it</strong>h age-related macular degeneration (AMD), the leading cause<br />
of blindness in people over 60 years of age, as well as confirming previously<br />
reported variants<br />
Nature Genetics - 38, 1055 - 1059 (2006)
They estimate that genotypes related to just five variants in three different genes can<br />
explain 50% of the risk of developing AMD<br />
The new genetic common variant identified was found in a non-coding region of the<br />
Complement Factor H (CFH) gene, other variants of which were recently shown to<br />
be associated w<strong>it</strong>h the risk of developing AMD.<br />
In add<strong>it</strong>ion to CFH on chromosome 1<br />
the complement factor B (BF) gene on chromosome 6<br />
complement component 2 (C2) gene on chromosome 6<br />
a common variant (A69S) is in hypothetical gene LOC387715 on chromosome 10.<br />
Interestingly, these three genes do not appear to interact directly, but instead<br />
contribute to the risk of AMD independently.
Phase II <strong>Hap</strong><strong>Map</strong> characterizes over 3.1 million human<br />
SNPs genotyped in 270 individuals from four<br />
geographically diverse populations
Genotyping in phase II was attempted for about 4.4 million<br />
distinct SNPs, of which roughly 1.3 million e<strong>it</strong>her could<br />
not be<br />
typed,<br />
were not<br />
polymorphic<br />
in any<br />
of the<br />
populations, or did not pass genotyping qual<strong>it</strong>y control<br />
filters.<br />
<br />
Certain regions of the genome were recognized as being<br />
challenging to study, such as centromeres, telomeres,<br />
gaps in genome sequence, and segmental duplications,<br />
regions declared to be not <strong>Hap</strong><strong>Map</strong>able.
The resulting <strong>Hap</strong><strong>Map</strong> has an SNP dens<strong>it</strong>y of approximately<br />
one per kilobase and is estimated to contain approximately<br />
25–35% of all the 9–10 million common SNPs in the<br />
assembled human genome
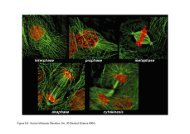
Variation in SNP dens<strong>it</strong>y w<strong>it</strong>hin the Phase II <strong>Hap</strong><strong>Map</strong><br />
Phase I<br />
Phase II<br />
Example of the fine-scale structure of SNP dens<strong>it</strong>y for a 100-kb region on chromosome<br />
17 showing polymorphic Phase I SNPs in the consensus data set (red triangles) and<br />
polymorphic Phase II SNPs in the consensus data set (blue triangles)<br />
The Phase II <strong>Hap</strong><strong>Map</strong> differs from the Phase I <strong>Hap</strong><strong>Map</strong> also in minor allele frequency<br />
(MAF) distribution. SNPs added in Phase II have lower MAF. Phase II <strong>Hap</strong><strong>Map</strong><br />
includes a better representation of rare variation than the Phase I <strong>Hap</strong><strong>Map</strong>
Advances in technology for high-throughput SNP genotyping<br />
Advances in genotyping technology have vastly increased the number<br />
of variants that can be typed and decreased the per-sample costs<br />
These advances have made possible the<br />
dense genotyping needed to capture the<br />
major<strong>it</strong>y of SNP variation w<strong>it</strong>hin an individual<br />
at a sufficiently low cost to allow the large<br />
sample sizes needed for comparison of<br />
individuals w<strong>it</strong>h and w<strong>it</strong>hout disease
Studies in add<strong>it</strong>ional populations have shown that the tag<br />
SNPs chosen using the <strong>Hap</strong><strong>Map</strong> are generally transferable<br />
across other populations, but there are some lim<strong>it</strong>ations.<br />
So add<strong>it</strong>ional samples from the populations used to develop<br />
the <strong>Hap</strong><strong>Map</strong> as well as from seven more populations have<br />
recently been genotyped across the genome.<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
Luhya from Webuye, Kenya<br />
Maasai from Kenya<br />
Tuscans from Italy<br />
Indian-Americans (Gujarati) from Houston, TX<br />
Han Chinese from Denver<br />
Mexican-Americans from Los Angeles<br />
Americans of African Descent from the SW USA
It is now clear that the <strong>Hap</strong><strong>Map</strong> can be<br />
a useful resource for the design and<br />
analysis of disease association studies<br />
in populations across the world
APPLICATION OF THE HAPMAP TO<br />
COMMON DISEASE
The technological advances directly stimulated or<br />
indirectly facil<strong>it</strong>ated by the <strong>Hap</strong><strong>Map</strong> have had a<br />
profound impact on the study of the genetics of<br />
common diseases
The history of high-dens<strong>it</strong>y GWA<br />
scanning to date has<br />
demonstrated the striking<br />
success of this approach in<br />
finding genetic variants<br />
associated w<strong>it</strong>h disease.<br />
Variants or regions associated<br />
w<strong>it</strong>h nearly 40 complex diseases<br />
have been identified in diverse<br />
population samples.
Major Autism Gene Found w<strong>it</strong>h Help of <strong>Hap</strong><strong>Map</strong><br />
Using data from the <strong>Hap</strong><strong>Map</strong>, along w<strong>it</strong>h DNA samples collected from many<br />
families who have affected children, researchers have discovered a genetic<br />
variation linked to autism, one of the most her<strong>it</strong>able mental health<br />
cond<strong>it</strong>ions.<br />
They found a variation in the sequence of a gene - the “MET receptor<br />
tyrosine kinase gene” - that is associated w<strong>it</strong>h autism. This gene is involved<br />
in brain development, immune function, and digestive system repair.<br />
The MET promoter variant rs1858830 allele "C" is strongly associated w<strong>it</strong>h<br />
ASD and results in reduced gene transcription. MET protein levels were<br />
significantly decreased in ASD cases compared w<strong>it</strong>h control subjects.<br />
People who have the variation are more than twice as likely as others to<br />
have “autism spectrum disorders”<br />
Campbell DB et al, Ann Neurol. 2007
A genome-wide association study identifies novel risk loci for<br />
type 2 diabetes<br />
Type 2 diabetes mell<strong>it</strong>us results from the interaction of environmental factors<br />
w<strong>it</strong>h a combination of genetic variants.<br />
A systematic search for these variants was recently made possible by the<br />
development of high-dens<strong>it</strong>y arrays that perm<strong>it</strong> the genotyping of hundreds of<br />
thousands of polymorphisms.<br />
Researchers tested 392,935 SNPs in a French case–control cohort.<br />
Markers w<strong>it</strong>h the most significant difference in genotype frequencies between<br />
cases of type 2 diabetes and controls were fast-tracked for testing in a second<br />
cohort.<br />
This identified four loci containing variants that confer type 2 diabetes risk, in<br />
add<strong>it</strong>ion to confirming the known association w<strong>it</strong>h the TCF7L2 gene.<br />
These loci include a non-synonymous polymorphism in the zinc transporter<br />
SLC30A8, which is expressed exclusively in insulin-producing β-cells, and two<br />
linkage disequilibrium blocks that contain genes potentially involved in β-cell<br />
development or function (IDE–KIF11–HHEX and EXT2–ALX4).<br />
Sladek R et al. Nature 445, 881-885 (2007)
Future of the <strong>Hap</strong><strong>Map</strong> <strong>Project</strong><br />
Currently, add<strong>it</strong>ional samples from the populations used to develop<br />
the in<strong>it</strong>ial <strong>Hap</strong><strong>Map</strong>, as well as samples from seven add<strong>it</strong>ional<br />
populations will be sequenced and genotyped extensively to extend<br />
the <strong>Hap</strong><strong>Map</strong>, providing information on rarer variants and helping to<br />
enable genome-wide association studies in add<strong>it</strong>ional populations.<br />
There are also ongoing efforts by many groups to characterize<br />
add<strong>it</strong>ional forms of genetic variation, such as structural variation, and<br />
molecular phenotypes in the <strong>Hap</strong><strong>Map</strong> samples. Finally, in the future,<br />
whole-genome sequencing will provide a natural convergence of<br />
technologies to type both SNP and structural variation.<br />
Nevertheless, until that point the <strong>Hap</strong><strong>Map</strong> <strong>Project</strong> data will provide an<br />
invaluable resource for understanding the structure of human genetic<br />
variation and <strong>it</strong>s link to phenotype.
Beyond SNPs:<br />
Copy Number Variants and Other<br />
Structural Variation
Current generation high-throughput<br />
genotyping platforms are<br />
extraordinarily efficient at genotyping<br />
SNPs, but they are less effective at<br />
genotyping structural variants, such<br />
as insertions, deletions, inversions,<br />
and copy number variants
Although not as common as SNPs, these variants also occur<br />
commonly in the human genome<br />
The distribution of copy number variation in the human genome among 270 <strong>Hap</strong><strong>Map</strong> samples
A Copy number variants (CNV) is<br />
a segment of DNA in which copynumber<br />
differences have been<br />
found by comparison of two or more<br />
genomes.<br />
CNV in which stretches of genomic<br />
sequence of roughly 1 kb to 3 Mb in<br />
size are deleted or are duplicated in<br />
varying numbers, have gained<br />
increasing attention because of their<br />
apparent ubiqu<strong>it</strong>y and potential<br />
dosage effect on gene expression.
In 2004, the interrogation of genomic variabil<strong>it</strong>y by array<br />
hybridization methods clearly demonstrated the existence of copy<br />
number variants.<br />
Intense analysis of this type of genomic variabil<strong>it</strong>y followed, and<br />
the current conservative estimate from studies in a few hundred<br />
individuals is that at least 10% of the genome is subject to copy<br />
number variation
Although a typical SNP affects only one single nucleotide<br />
pair, their genomic abundance (over 10 million) makes<br />
them the most frequent source of polymorphic changes<br />
By contrast, CNVs are far less numerous but can affect<br />
from one kilobase to several megabases of DNA per<br />
event, adding up to a significant fraction of the genome
It is now recognized that the genomes of any two<br />
individuals in the human population differ more at the<br />
structural level than at the nucleotide sequence level<br />
NATURE GENETICS SUPPLEMENT | VOLUME 39 | JULY 2007
Much of what was previously known about the role of CNVs in disease<br />
comes from a rich l<strong>it</strong>erature on ‘genomic disorders’.<br />
<br />
Genomic disorders are defined as a diverse group of genetic diseases that<br />
are each caused by an alteration in DNA copy number.<br />
These mutations can be relatively large, microscopically visible<br />
imbalances, such as in Prader-Willi syndrome, or they may be much<br />
smaller, requiring higher resolution detection methods, such as in Williams<br />
Syndrome.<br />
<br />
Genomic disorders are typically sporadic in nature because the CNV in<br />
most cases is a de novo mutation w<strong>it</strong>h nearly complete penetrance, and<br />
because the affected individuals have severe developmental problems and<br />
are unlikely to have offspring.<br />
However, there are notable examples of mendelian disease tra<strong>it</strong>s<br />
associated w<strong>it</strong>h CNVs. For example, duplications of the gene for peripheral<br />
myelin protein 22 (PMP22) cause the dominant neuropathy Charcot-Marie<br />
Tooth disease type 1A, and deletions of the α-globin gene cluster cause<br />
the recessive anemia α-thalassemia.
Bibliografia<br />
The International <strong>Hap</strong><strong>Map</strong> Consortium. The International <strong>Hap</strong><strong>Map</strong> <strong>Project</strong>.<br />
NATURE. 426: 18/25, December 2003.<br />
Deloukas P, Bentley D. The <strong>Hap</strong><strong>Map</strong> project and <strong>it</strong>s application to genetic<br />
studies of drug response. The Pharmacogenomics Journal. 4, 88–90 (2004).<br />
The International <strong>Hap</strong><strong>Map</strong> Consortium. A haplotype map of the human<br />
genome. NATURE. 437: 27, October 2005.<br />
Manolio TA, Brooks LD, Collins FS. A <strong>Hap</strong><strong>Map</strong> harvest of insights into the<br />
genetics of common disease. The Journal of Clinical Investigation. 118: 5,<br />
May 2008.<br />
The International <strong>Hap</strong><strong>Map</strong> Consortium. A second generation human<br />
haplotype map of over 3.1 million SNPs. NATURE. 449: 18, October 2007.<br />
<br />
Maller J, George S, Purcell S, Fagerness J, Altshuler D, Daly MJ, Seddon JM.<br />
Common variation in three genes, including a noncoding variant in CFH,<br />
strongly influences risk of age-related macular degeneration. Nat Genet. 38:9<br />
(1055-9), Sep 2006.








![Glucidi-Lipidi-010210 [modalità compatibilità] - Bgbunict.it](https://img.yumpu.com/41100335/1/190x134/glucidi-lipidi-010210-modalita-compatibilita-bgbunictit.jpg?quality=85)