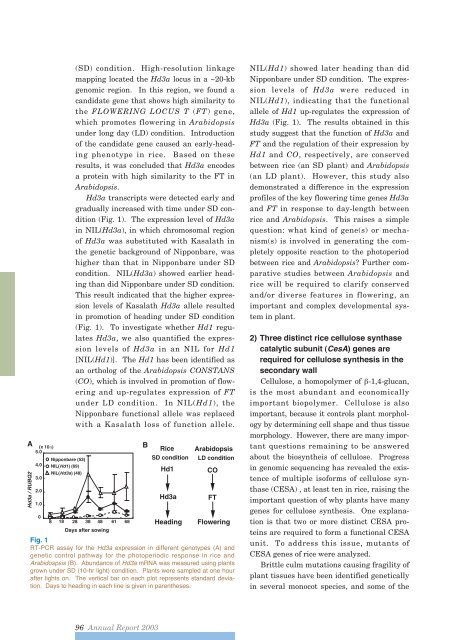
AHd3a / RUBQ25.04.03.02.01.0(x 10-3)0Nipponbare (53)NIL(Hd1) (69)NIL(Hd3a) (48)(SD) condition. High-resolution linkagemapping located the Hd3a locus in a ~20-kbgenomic region. In this region, we found acandidate gene that shows high similarity tothe FLOWERING LOCUS T (FT) gene,which promotes flowering in Arabidopsisunder long day (LD) condition. Introductionof the candidate gene caused an early-headingphenotype in rice. Based on theseresults, it was concluded that Hd3a encodesa protein with high similarity to the FT inArabidopsis.Hd3a transcripts were detected early andgradually increased with time under SD condition(Fig. 1). The expression level of Hd3ain NIL(Hd3a), in which chromosomal regionof Hd3a was substituted with Kasalath inthe genetic background of Nipponbare, washigher than that in Nipponbare under SDcondition. NIL(Hd3a) showed earlier headingthan did Nipponbare under SD condition.This result indicated that the higher expressionlevels of Kasalath Hd3a allele resultedin promotion of heading under SD condition(Fig. 1). To investigate whether Hd1 regulatesHd3a, we also quantified the expressionlevels of Hd3a in an NIL for Hd1[NIL(Hd1)]. The Hd1 has been identified asan ortholog of the Arabidopsis CONSTANS(CO), which is involved in promotion of floweringand up-regulates expression of FTunder LD condition. In NIL(Hd1), theNipponbare functional allele was replacedwith a Kasalath loss of function allele.8 18 28 38 48 61 68Days after sowingFig. 1RT-PCR assay for the Hd3a expression in different genotypes (A) andgenetic control pathway for the photoperiodic response in rice andArabidospsis (B). Abundance of Hd3a mRNA was measured using plantsgrown under SD (10-hr light) condition. Plants were sampled at one hourafter lights on. The vertical bar on each plot represents standard deviation.Days to heading in each line is given in parentheses.BRiceSD conditionHd1Hd3aHeadingArabidopsisLD conditionCOFTFloweringNIL(Hd1) showed later heading than didNipponbare under SD condition. The expressionlevels of Hd3a were reduced inNIL(Hd1), indicating that the functionalallele of Hd1 up-regulates the expression ofHd3a (Fig. 1). The results obtained in thisstudy suggest that the function of Hd3a andFT and the regulation of their expression byHd1 and CO, respectively, are conservedbetween rice (an SD plant) and Arabidopsis(an LD plant). However, this study alsodemonstrated a difference in the expressionprofiles of the key flowering time genes Hd3aand FT in response to day-length betweenrice and Arabidopsis. This raises a simplequestion: what kind of gene(s) or mechanism(s)is involved in generating the completelyopposite reaction to the photoperiodbetween rice and Arabidopsis? Further comparativestudies between Arabidopsis andrice will be required to clarify conservedand/or diverse features in flowering, animportant and complex developmental systemin plant.2) Three distinct rice cellulose synthasecatalytic subunit (CesA) genes arerequired for cellulose synthesis in thesecondary wallCellulose, a homopolymer of -1,4-glucan,is the most abundant and economicallyimportant biopolymer. Cellulose is alsoimportant, because it controls plant morphologyby determining cell shape and thus tissuemorphology. However, there are many importantquestions remaining to be answeredabout the biosyntheis of cellulose. Progressin genomic sequencing has revealed the existenceof multiple isoforms of cellulose synthase(CESA) , at least ten in rice, raising theimportant question of why plants have manygenes for cellulose synthesis. One explanationis that two or more distinct CESA proteinsare required to form a functional CESAunit. To address this issue, mutants ofCESA genes of rice were analyzed.Brittle culm mutations causing fragility ofplant tissues have been identified geneticallyin several monocot species, and some of the96 <strong>Annual</strong> <strong>Report</strong> <strong>2003</strong>
mutants have been studied in relation to cellulosesynthesis, but they have not beencharacterized at a molecular level. Weshowed that the genes responsible for threedistinct brittle mutations of rice (Fig. 2),induced by the insertion of the endogenousretrotransposon Tos17, correspond to cellulosesynthase catalytic subunit (CesA) genes,OsCesA4, OsCesA7 and OsCesA9. ThreeCesA genes were expressed in seedlings,culms, premature panicles and roots, but notin mature leaves, and the expression profileswere almost identical among the three genes.Cellulose contents were dramaticallydecreased (8.9% to 25.5% of the wild typelevel) in the culms of null-mutants of thethree genes, indicating these genes are notfunctionally redundant. Consistent withthese results, cell walls in the cortical fibercells were shown to be thinner in all themutants than in wild-type plants. Theseresults suggest that in rice at least three distinctCesA genes are required to form a functionalCESA unit.ABC1 2 3 4 5 62 3W M W ML4 5W M W MC LW M W M6W M W MCLCL3) Rice ß-tubulin genes are up-regulatedin response to gibberllinsGibberellins (GAs) play a significant rolein various processes during plant development,including seed germination, plantgrowth and flower initiation. To elucidatethe effect of GA on these processes, the GAbiosynthesis pathway and GA signaling pathwayhave been widely studied in many plantsusing genetic, molecular and biochemicalapproaches. To screen downstream genesrelated to shoot elongation in GA signaling inrice (Oryza sativa L. cv Nipponbare), a cDNAmicroarray containing 9,000 ESTs washybridized to the RNA isolated from the leafsheath including apical meristem of riceseedling treated with GA for 24 hr. Among9,000 genes analyzed, 16 genes showed theenhanced expression, while 6 genes wererepressed by GA treatment as compared tothe control. Northern blot analyses usingrice EST clones in secondary screeningshowed that -tubulin gene (OsTUB) is significantlyup-regulated by GA treatment.W M W MCLFig. 2Phenotypes of the five rice brittle culm mutants.A, Plants of the wild-type (1) and the mutants (2-6). B, Brittlenessof culm (C) and mature leaf (L) from wild-type plant (W) and themutant (M) as demonstrated by the damage caused by stressingbetween fingers. Numbers indicates mutant lines as describeabove. All culms were prepared from the 2nd internodes.Homology searching within the EST databaseidentified at least 8 OsTUB isotypes.Specific probes to 3’-UTR of OsTUB isotypesshowed differential and tissue-specificexpression and were significantly inducedby GA, but were repressed by light andabscisic acid. Seven OsTUB genes showedhigh expression in the leaf sheath includingapical meristem, which is very active inshoot elongation, signifying their role inshoot elongation through cell division andcell elongation. In contrast, OsTUB8 wasconsistent expressed only in anthers and hada different branch in the dendrogram (Fig.3), consistent with the AtTUA1 and AtTUB9<strong>Annual</strong> <strong>Report</strong> <strong>2003</strong> 97
- Page 3 and 4:
Message from the PresidentIn cooper
- Page 5 and 6:
¡X-ray crystallographic studies of
- Page 7 and 8:
Topics of Research in This YearComp
- Page 9 and 10:
Elucidation of genomic structure ar
- Page 11 and 12:
the total library) were mapped onto
- Page 13 and 14:
observed among the ribosomal protei
- Page 15 and 16:
molecular analyses and morphologica
- Page 17 and 18:
with 5 IU equine chorionic gonadotr
- Page 19 and 20:
serious problem in rice production
- Page 21 and 22:
exogenous genes into a hymenopteran
- Page 23 and 24:
Isolation and characterization of B
- Page 25 and 26:
oids could be regenerated from endo
- Page 27 and 28:
Quantitative trait locus analyses o
- Page 29 and 30:
ion, induces hypercholesterolemia,
- Page 31 and 32:
is an important source for amino ac
- Page 33 and 34:
In this study, 1.4 and 1.6 kb-long
- Page 35 and 36:
aqueous solution using cyanuric chl
- Page 37 and 38:
Developing a waste selection device
- Page 39 and 40:
decreased every five days by 5˚C,
- Page 41 and 42:
ticides. If the viral dose that kil
- Page 43 and 44:
In addition, the number of crown ro
- Page 45 and 46:
Molecular structure of the GARP fam
- Page 47 and 48:
X-ray crystallographic studies ofSt
- Page 49 and 50:
Rapid and high resolution QTL analy
- Page 51 and 52: Plant regeneration system through m
- Page 53 and 54: Table 1 Transformation Efficiency a
- Page 55 and 56: from the tissues of mutants. The In
- Page 57 and 58: the regenerated plants were transpl
- Page 59 and 60: plants and fungi, and RNA interfere
- Page 61 and 62: eeding, classification of microbes
- Page 63 and 64: efforts are made to obtain single m
- Page 65 and 66: was carried out by shotgun sequenci
- Page 67 and 68: epetitive sequence specific to Oryz
- Page 69 and 70: 75 µM of ABA at 25ºC. More than 8
- Page 71 and 72: Change of the molecular weightforms
- Page 73 and 74: A unique landrace group recognizedb
- Page 75 and 76: eaction (PCR) primer pairs to detec
- Page 77 and 78: This was the first successful case
- Page 79 and 80: to disappear by degradation or dilu
- Page 81 and 82: tained 1933 independent genes and b
- Page 83 and 84: that antigen-specific regulatory T
- Page 85 and 86: neural activity patterns evoked by
- Page 87 and 88: considerable effect on the suscepti
- Page 89 and 90: Insect Genetics and Evolution Depar
- Page 91 and 92: activity, which was found to detoxi
- Page 93 and 94: Insect Biomaterial and Technology D
- Page 95 and 96: peptides obtained from sericin a re
- Page 97 and 98: DNA markers for Nid-1, a resistance
- Page 99 and 100: or carotenoid pigment. When sericin
- Page 101: Plant Science DivisionThe Plant Sci
- Page 105 and 106: have verified that candidate genes
- Page 107 and 108: and C, connected by a distorted typ
- Page 109 and 110: mammalian cells, respectively, inhi
- Page 111 and 112: undle sheath cells (BSCs), the land
- Page 113 and 114: (OsNHX1), barley (HvNHX1) and baril
- Page 115 and 116: nation rates. However, in our attem
- Page 117 and 118: Yeast two-hybrid assay showed inter
- Page 119 and 120: and a few NADPH-cytochrome P450 oxi
- Page 121 and 122: the R0(53%).Foreign genes (35S + np
- Page 123 and 124: gamma ray from 44 TBq 60 Co was use
- Page 125 and 126: List of PublicationOriginal Papers1
- Page 127 and 128: 35 Fujisaki S, Mizoguchi Y, Takahas
- Page 129 and 130: 71 Imai K, Khandoker MY, Yonai M, T
- Page 131 and 132: 107 Kiuchi S, Inage Y, Hiraiwa H, U
- Page 133 and 134: 147 Miyamoto Y, Sakumoto R, Sakabe
- Page 135 and 136: 184 Ochi A, Hossain KS, Magoshi J,
- Page 137 and 138: 219 Takahashi M, Nagai T, Okamura N
- Page 139 and 140: 258 Watanabe H, Nakashima K, Saito
- Page 141 and 142: Author Department Paper.No.Yasushi
- Page 143 and 144: Author Department Paper.No.Sadao Wa
- Page 145 and 146: Monograph1 Magoshi J, Nakamura S (2
- Page 147 and 148: on Lepidopteran Genomics”.NIAS-CO
- Page 149 and 150: tion of the high-quality draft sequ
- Page 151 and 152: Executive Members andResearch Staff
- Page 153 and 154:
Insect Growth Regulation Laboratory
- Page 155 and 156:
Sericultural Science LaboratoryMole
- Page 157 and 158:
Members of NIAS EvaluationComittee(
- Page 159 and 160:
Annual Report 2003 153