New Approaches to in silico Design of Epitope-Based Vaccines
New Approaches to in silico Design of Epitope-Based Vaccines
New Approaches to in silico Design of Epitope-Based Vaccines
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
104 APPENDIX B. EPITOPE DISCOVERY<br />
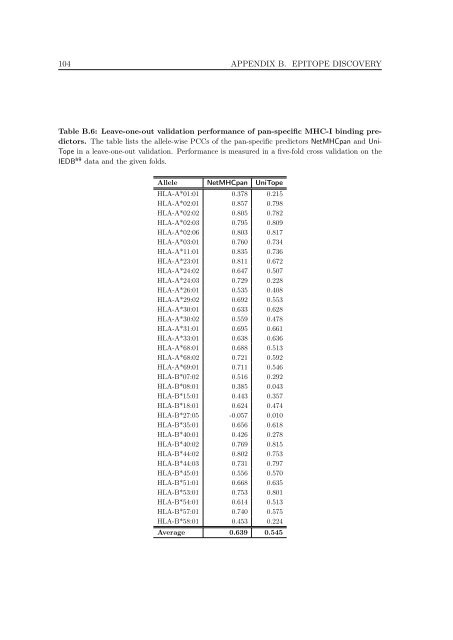
Table B.6: Leave-one-out validation performance <strong>of</strong> pan-specific MHC-I b<strong>in</strong>d<strong>in</strong>g predic<strong>to</strong>rs.<br />
The table lists the allele-wise PCCs <strong>of</strong> the pan-specific predic<strong>to</strong>rs NetMHCpan and Uni-<br />
Tope <strong>in</strong> a leave-one-out validation. Performance is measured <strong>in</strong> a five-fold cross validation on the<br />
IEDB h9 data and the given folds.<br />
Allele NetMHCpan UniTope<br />
HLA-A*01:01 0.378 0.215<br />
HLA-A*02:01 0.857 0.798<br />
HLA-A*02:02 0.805 0.782<br />
HLA-A*02:03 0.795 0.809<br />
HLA-A*02:06 0.803 0.817<br />
HLA-A*03:01 0.760 0.734<br />
HLA-A*11:01 0.835 0.736<br />
HLA-A*23:01 0.811 0.672<br />
HLA-A*24:02 0.647 0.507<br />
HLA-A*24:03 0.729 0.228<br />
HLA-A*26:01 0.535 0.408<br />
HLA-A*29:02 0.692 0.553<br />
HLA-A*30:01 0.633 0.628<br />
HLA-A*30:02 0.559 0.478<br />
HLA-A*31:01 0.695 0.661<br />
HLA-A*33:01 0.638 0.636<br />
HLA-A*68:01 0.688 0.513<br />
HLA-A*68:02 0.721 0.592<br />
HLA-A*69:01 0.711 0.546<br />
HLA-B*07:02 0.516 0.292<br />
HLA-B*08:01 0.385 0.043<br />
HLA-B*15:01 0.443 0.357<br />
HLA-B*18:01 0.624 0.474<br />
HLA-B*27:05 -0.057 0.010<br />
HLA-B*35:01 0.656 0.618<br />
HLA-B*40:01 0.426 0.278<br />
HLA-B*40:02 0.769 0.815<br />
HLA-B*44:02 0.802 0.753<br />
HLA-B*44:03 0.731 0.797<br />
HLA-B*45:01 0.556 0.570<br />
HLA-B*51:01 0.668 0.635<br />
HLA-B*53:01 0.753 0.801<br />
HLA-B*54:01 0.614 0.513<br />
HLA-B*57:01 0.740 0.575<br />
HLA-B*58:01 0.453 0.224<br />
Average 0.639 0.545