The Staphylococcus aureus secretome - TI Pharma
The Staphylococcus aureus secretome - TI Pharma
The Staphylococcus aureus secretome - TI Pharma
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
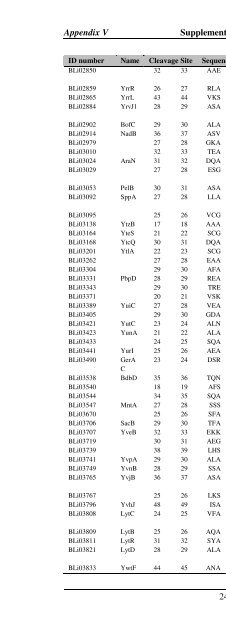
Appendix V Supplemental Table Va<br />
ID number Name Cleavage Site Sequence Function<br />
BLi02850 32 33 AAE DS putative cell wall-associated protease<br />
precursor(EC 3.4.21.-)<br />
BLi02859 YrrR 26 27 RLA EI unknown; similar to penicillin-binding protein<br />
BLi02865 YrrL 43 44 VKS AL unknown; similar to folate metabolism<br />
BLi02884 YrvJ1 28 29 ASA AI unknown; similar to N-acetylmuramoyl-Lalanine<br />
amidase<br />
BLi02902 BofC 29 30 ALA EK forespore regulator of the sigma-K checkpoint<br />
BLi02914 NadB 36 37 ASV KD L-aspartate oxidase<br />
BLi02979 27 28 GKA EF hypothetical protein<br />
BLi03010 32 33 TEA SE unknown<br />
BLi03024 AraN 31 32 DQA DG L-arabinose transport (sugar-binding protein)<br />
BLi03029 27 28 ESG KA close homolog to AbnA arabinan-endo 1,5-α-Larabinase<br />
BLi03053 PelB 30 31 ASA AN pectate lyase (EC 4.2.2.10)<br />
BLi03092 SppA 27 28 LLA VF putative signal peptide peptidase required for<br />
efficient processing of pre-proteins (EC 3.4.21.-)<br />
BLi03095 25 26 VCG YY unknown<br />
BLi03138 YtzB 17 18 AAA VV unknown<br />
BLi03164 YteS 21 22 SCG KD unknown<br />
BLi03168 YtcQ 30 31 DQA SS unknown; similar to lipoprotein<br />
BLi03201 YtlA 22 23 SCG GQ unknown<br />
BLi03262 27 28 EAA SQ unknown<br />
BLi03304 29 30 AFA AS putative sugar hydrolase<br />
BLi03331 PbpD 28 29 REA QN penicillin-binding protein 4<br />
BLi03343 29 30 TRE QT unknown<br />
BLi03371 20 21 VSK AG putative lipase precursor<br />
BLi03389 YuiC 27 28 VEA QD unknown; similar to proteins<br />
BLi03405 29 30 GDA EF hypothetical protein<br />
BLi03421 YutC 23 24 ALN DT unknown; similar to proteins<br />
BLi03423 YunA 21 22 ALA KE unknown<br />
BLi03433 24 25 SQA AD unknown<br />
BLi03441 YurI 25 26 AEA FQ unknown; similar to ribonuclease<br />
BLi03490 GerA<br />
C<br />
23 24 DSR QI germination response to L-alanine<br />
BLi03538 BdbD 35 36 TQN AS thiol-disulfide oxidoreductase<br />
BLi03540 18 19 AFS AG putative ABC transporter sugar binding protein<br />
BLi03544 34 35 SQA DE putative sugar hydrolase<br />
BLi03547 MntA 27 28 SSS EE manganese ABC transporter (membrane protein)<br />
BLi03670 25 26 SFA KD unknown<br />
BLi03706 SacB 29 30 TFA KE levansucrase (EC 2.4.1.10)<br />
BLi03707 YveB 32 33 EKK GE unknown; similar to levanase<br />
BLi03719 30 31 AEG PQ putative ribonuclease (EC 3.1.27.-)<br />
BLi03739 38 39 LHS SH unknown<br />
BLi03741 YvpA 29 30 ALA AE unknown; similar to pectate lyase<br />
BLi03749 YvnB 28 29 SSA SG unknown<br />
BLi03765 YvjB 36 37 ASA AE unknown; similar to carboxy-terminal<br />
processing protease<br />
BLi03767 25 26 LKS NH putative cell wall-binding protein<br />
BLi03796 YvhJ 48 49 ISA AD unknown; similar to transcriptional regulator<br />
BLi03808 LytC 24 25 VFA AN N-acetylmuramoyl-L-alanine amidase<br />
(EC3.5.1.28)<br />
BLi03809 LytB 25 26 AQA AD modifier protein of major autolysin LytC<br />
BLi03811 LytR 31 32 SYA YY attenuator role for lytABC and lytR expression<br />
BLi03821 LytD 28 29 ALA AY N-acetylglucosaminidase (major autolysin)<br />
(EC3.2.1.96)<br />
BLi03833 YwtF 44 45 ANA SK unknown; similar to transcriptional regulator<br />
246