WORKSHOP BOOK Fluorescent in Situ Hybridisation - FISH - on Human Sperm
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
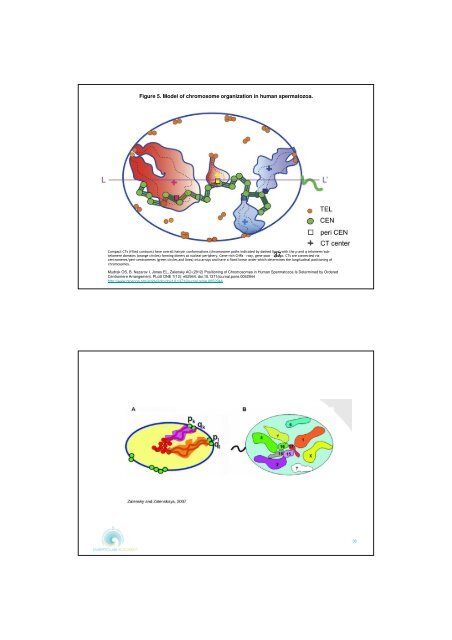
Figure 5. Model of chromosome organizati<strong>on</strong> <str<strong>on</strong>g>in</str<strong>on</strong>g> human spermatozoa.<br />
Compact CTs (filled c<strong>on</strong>tours) have overall hairp<str<strong>on</strong>g>in</str<strong>on</strong>g> c<strong>on</strong>formati<strong>on</strong>s (chromosome paths <str<strong>on</strong>g>in</str<strong>on</strong>g>dicated by dashed l<str<strong>on</strong>g>in</str<strong>on</strong>g>es) with the p and q telomere/subtelomere<br />
doma<str<strong>on</strong>g>in</str<strong>on</strong>g>s (orange circles) form<str<strong>on</strong>g>in</str<strong>on</strong>g>g dimers at nuclear periphery. Gene-rich CHRs – rosy, gene-poor – 37 <str<strong>on</strong>g>in</str<strong>on</strong>g>digo. CTs are c<strong>on</strong>nected via<br />
centromeres/peri-centromeres (green circles and l<str<strong>on</strong>g>in</str<strong>on</strong>g>es) <str<strong>on</strong>g>in</str<strong>on</strong>g>to arrays and have a fixed l<str<strong>on</strong>g>in</str<strong>on</strong>g>ear order which determ<str<strong>on</strong>g>in</str<strong>on</strong>g>es the l<strong>on</strong>gitud<str<strong>on</strong>g>in</str<strong>on</strong>g>al positi<strong>on</strong><str<strong>on</strong>g>in</str<strong>on</strong>g>g of<br />
chromosomes.<br />
Mudrak OS, B. Nazarov I, J<strong>on</strong>es EL, Zalensky AO (2012) Positi<strong>on</strong><str<strong>on</strong>g>in</str<strong>on</strong>g>g of Chromosomes <str<strong>on</strong>g>in</str<strong>on</strong>g> <strong>Human</strong> <strong>Sperm</strong>atozoa Is Determ<str<strong>on</strong>g>in</str<strong>on</strong>g>ed by Ordered<br />
Centromere Arrangement. PLoS ONE 7(12): e52944. doi:10.1371/journal.p<strong>on</strong>e.0052944<br />
http://www.plos<strong>on</strong>e.org/article/<str<strong>on</strong>g>in</str<strong>on</strong>g>fo:doi/10.1371/journal.p<strong>on</strong>e.0052944<br />
38