Principles of cell signaling - UT Southwestern
Principles of cell signaling - UT Southwestern
Principles of cell signaling - UT Southwestern
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Ser Thr<br />
IRAK-M<br />
TRIF/<br />
TICAM-1<br />
SOCS1<br />
/JAB<br />
ys63TRAF6<br />
0<br />
CAPK<br />
CAPK<br />
IRAK1<br />
Lys<br />
TBK-1<br />
P<br />
P<br />
TRAF2 TRAF1<br />
A20<br />
P Tyr STAT6<br />
K<br />
-2<br />
TNF<br />
TICAM-1<br />
P<br />
Ser<br />
IRAK1<br />
Thr<br />
Lys<br />
Ser<br />
IRAK1<br />
Thr Lys<br />
IKK<br />
Ser176<br />
Ser181 IKK<br />
IKK<br />
IKK<br />
P<br />
Ser176<br />
Ser181 IKK<br />
P<br />
IKK<br />
GM-CSFR<br />
GM-CSFR<br />
Uev1A<br />
Ubc13<br />
P Tyr STAT5<br />
PI3K<br />
MEKK<br />
Tyr STAT5<br />
GM-CSF<br />
TLR4<br />
PP2A<br />
TAB2<br />
P<br />
P<br />
P<br />
Thr184<br />
Ser192 TAK1 Thr187 P<br />
P P<br />
TBK-1<br />
P Ser TAB1<br />
Thr<br />
P<br />
P<br />
Tyr580<br />
P<br />
TRIF/<br />
TICAM-1<br />
TRAM<br />
MD-2<br />
P P<br />
Tyr STAT5<br />
Tyr<br />
Ser386<br />
Ser385<br />
IRF-3<br />
Ser Thr IRAK1<br />
Lys<br />
Ub<br />
P<br />
P<br />
P<br />
P Tyr STAT3<br />
Tyr STAT3<br />
Thr38 Ser473<br />
Akt/PKB<br />
P<br />
P<br />
Thr38 Ser473<br />
Akt/PKB<br />
Ser Thr IRAK1 Lys<br />
Ub<br />
PKA<br />
PKA<br />
TRAM<br />
ASK<br />
Lys21<br />
Lys21<br />
I B<br />
Ser32<br />
Lys22<br />
Ser36<br />
NF-<br />
B<br />
p65+p50<br />
Ser276<br />
Ser529<br />
P<br />
Lys21<br />
I B<br />
Ser32<br />
Lys22<br />
Ser36<br />
P<br />
PKA<br />
NF-<br />
B<br />
p65+p50<br />
Ser276<br />
Ser529<br />
P<br />
Tyr701<br />
TAB2<br />
Tyr701<br />
Ser727 STAT1<br />
IKK<br />
Thr184 Ser192 TAK1 Thr187<br />
P<br />
P<br />
Ser<br />
TAB2<br />
TAB1<br />
Ser32<br />
Ser36<br />
IKK<br />
Ser Thr IRAK1<br />
Lys<br />
Ub<br />
Thr184<br />
Ser192<br />
TAK1<br />
Thr187 P<br />
SEK1/MKK4<br />
P<br />
Tyr701<br />
Ser727<br />
STAT1<br />
P<br />
P<br />
Tyr STAT2<br />
IRF-9<br />
Thr<br />
Ser386<br />
P<br />
P<br />
CK II<br />
Tyr JAK1<br />
IL-10R<br />
Tyr<br />
Tyr JAK1<br />
Ser<br />
IL-10R<br />
TAB1<br />
NF- B<br />
p65+p50<br />
Ser276 Ser529<br />
IL-10R<br />
Tyr<br />
NF- B<br />
p65+p50<br />
Ser276 Ser529<br />
P<br />
P<br />
NF- B<br />
p65+p50<br />
Ser276 Ser529<br />
P<br />
NF-<br />
B<br />
PKA<br />
p65+p50<br />
Ser276<br />
Ser529<br />
IRF-9<br />
P<br />
P<br />
Ser<br />
Thr<br />
glycerol 3P<br />
TAB2<br />
TAB1<br />
P<br />
IL-10R<br />
Tyr<br />
Tyk2<br />
Tyr<br />
IL-10R<br />
Thr184<br />
Ser192<br />
TAK1<br />
Thr187<br />
P<br />
Ser<br />
TAB2<br />
TAB1<br />
Thr<br />
Ser Thr IRAK1<br />
Lys<br />
Ub<br />
Thr184<br />
Ser192<br />
TAK1<br />
Thr187<br />
Thr<br />
SEK2/MKK7<br />
SEK1/MKK4<br />
Tyk2 Tyr<br />
Tyr STAT5<br />
p50<br />
p60<br />
P<br />
Thr183 ERK1<br />
Tyr185ERK2<br />
P<br />
TG<br />
PKA<br />
malic enzyme<br />
P<br />
Tyr<br />
P<br />
Tyr542<br />
Tyr580 SHP-2<br />
Tyr580<br />
P<br />
lipid droplet<br />
SOCS3<br />
SHP-1<br />
Tyr JAK1<br />
Tyr759<br />
Tyr915<br />
IL-6R<br />
Tyr767<br />
Tyr905<br />
Tyr814<br />
Tyr JAK1<br />
Tyr JAK1<br />
Tyr STAT3 P Tyr STAT3<br />
Thr<br />
Thr<br />
Ub<br />
Lys21<br />
Ub<br />
IL-1ra<br />
malate<br />
Tyr580<br />
Tyr542 SHP-2<br />
P Tyr STAT5<br />
JNK<br />
JNK<br />
Tyr<br />
Tyr<br />
P<br />
NF- B<br />
p65+p50<br />
Ser276 Ser529<br />
Ser276 Ser529<br />
P<br />
P<br />
P<br />
P<br />
Ser32<br />
Ser36<br />
P<br />
P<br />
pyruvate<br />
carrier<br />
SEK2/MKK7<br />
PAFR<br />
TNF<br />
TG<br />
y<br />
Tyr<br />
P<br />
malate<br />
dehydrogenase<br />
I B<br />
GM-CSF<br />
IL-6<br />
p50<br />
IL-1<br />
oxaloacetate<br />
PKA<br />
NADH+H+<br />
Tyk2<br />
Tyr<br />
Tyr759<br />
Tyr915<br />
Tyr767<br />
IL-6R<br />
Tyr905<br />
Tyr814<br />
Tyk2<br />
Tyr<br />
Tyr759 Tyr915<br />
IL-6R<br />
Tyr767 Tyr905<br />
Tyr814<br />
Ub<br />
P<br />
Lys21<br />
I B<br />
Ser32<br />
Lys22<br />
Ser36<br />
Ub<br />
P<br />
PKA<br />
NF-<br />
B<br />
p65+p50<br />
Ser276<br />
Ser529<br />
IFN-<br />
IFN-<br />
IFNpyruvate<br />
pyruvate acetyl CoA<br />
A20<br />
P<br />
P<br />
Ser385<br />
Ser386<br />
P<br />
Ser386<br />
P<br />
NF- B<br />
p65+p50<br />
Ser276 Ser529<br />
Ser276 Ser529<br />
P<br />
P<br />
P<br />
P<br />
IL-6R<br />
gp130<br />
P<br />
P<br />
P<br />
Tyr759 Tyr915 Tyr759 Tyr915<br />
Tyr767 IL-6R IL-6R<br />
Tyr905<br />
Tyr767<br />
Tyr905<br />
P<br />
Tyr814<br />
P<br />
Tyr814<br />
P<br />
P<br />
P<br />
Tyr JAK1<br />
Tyk2<br />
Tyr<br />
P<br />
P<br />
SOCS3<br />
IL-1ra<br />
pyruvate<br />
carboxylase<br />
P<br />
Lys21<br />
I B<br />
Ser32<br />
Lys22<br />
Ser36<br />
P<br />
NF-<br />
B<br />
p65+p50<br />
Ser276<br />
Ser529<br />
IFN<br />
P<br />
P<br />
Ser385<br />
Ser386<br />
Ser386<br />
P<br />
P<br />
citrate<br />
liase<br />
PDH kinase<br />
PDH kinase<br />
P<br />
P<br />
pyruvate<br />
dehydrogenase<br />
dehydrogenase<br />
y<br />
Tyr JAK1<br />
IFN<br />
IFN<br />
Tyr440<br />
Tyr JAK1<br />
Tyk2 Tyr<br />
p38MAPK<br />
IFN-<br />
SCF<br />
R1<br />
IFN<br />
Tyr440<br />
R1<br />
Tyr701<br />
Ser727 STAT1<br />
P P<br />
Tyr STAT3<br />
Tyr<br />
P P<br />
Tyr Tyr STAT3<br />
P P<br />
Tyr Tyr STAT5<br />
TrCP<br />
UbcH5<br />
IFN<br />
acetyl CoA<br />
citrate<br />
Tyr1007<br />
R2<br />
Tyr<br />
P P<br />
Tyr Tyr<br />
STAT3<br />
SHP-1<br />
IFN<br />
R2<br />
Tyr<br />
JAK2<br />
Tyr1007<br />
PIAS3<br />
JAK2Tyr1007<br />
P<br />
P<br />
SOCS3<br />
P<br />
Tyr701<br />
Ser727 STAT1<br />
P<br />
P Tyr701 Tyr701<br />
P<br />
Ser727 STAT1<br />
Ser727<br />
P<br />
CoASH<br />
carnitine<br />
P<br />
P Tyr701<br />
Ser727 Tyr701 STAT1<br />
P<br />
P<br />
P Tyr701 Tyr701<br />
Ser727 STAT1<br />
P Ser727<br />
P<br />
carnitine<br />
acyl-CoA<br />
y<br />
P<br />
PIAS3<br />
P<br />
IL-4R<br />
Tyr<br />
IFN<br />
R1<br />
IFN<br />
R2<br />
Tyr440<br />
Tyr<br />
P<br />
P<br />
JAK2<br />
Tyr JAK1<br />
Tyr1007<br />
P<br />
SOCS1<br />
/JAB<br />
P<br />
P<br />
Thr38 Ser473<br />
Akt/PKB<br />
Thr38 Ser473<br />
Akt/PKB<br />
P<br />
IRF-9<br />
CoASH<br />
Tyr1007<br />
y<br />
P<br />
P<br />
acyl-CoA<br />
CoASH<br />
CPT I<br />
IL-4R<br />
Tyr580<br />
P<br />
P<br />
P<br />
P<br />
common<br />
Tyr<br />
chain<br />
JAK3Tyr<br />
SOCS1<br />
/JAB<br />
SOCS1<br />
/JAB<br />
SOCS1/JAB<br />
Tyr701<br />
Ser727<br />
STAT1<br />
Tyr STAT2<br />
IRF-9<br />
P<br />
Tyr STAT6<br />
IRF-2<br />
acylcarnitine<br />
acylcarnitine<br />
P<br />
J 3 y<br />
P<br />
IL-4R<br />
Tyr<br />
P Tyr STAT6<br />
Tyr<br />
TyrSTAT6<br />
MKP<br />
CACT<br />
CPT II<br />
P<br />
Tyr580<br />
Tyr542 SHP-2 SHP-1<br />
P<br />
SHIP<br />
PI3K<br />
PI3K<br />
PIAS1<br />
IRF-2<br />
IRF-2<br />
P P<br />
Tyr Tyr STAT6<br />
P P<br />
Tyr Tyr STAT6<br />
ATP<br />
ADP<br />
Tyr Fyn<br />
IRF-1<br />
y<br />
IL-4R<br />
Tyr<br />
Tyr JAK1<br />
P Tyr Fyn<br />
Tyr JAK1<br />
Tyr701<br />
Tyr701<br />
Ser727 STAT1<br />
Ser727<br />
P P Tyr701<br />
Ser727<br />
Tyr701<br />
STAT1<br />
P<br />
Ser727<br />
P<br />
PIAS1<br />
IRF-1<br />
IRF-1<br />
malonyl CoA<br />
fatty acid<br />
ATP<br />
synthetase<br />
IL-4<br />
JAK3Tyr<br />
IL-4<br />
y<br />
common<br />
Tyr<br />
chain<br />
JAK3Tyr<br />
Tyr<br />
IRS<br />
Ser312(307:R)<br />
p53<br />
NOSII/iNOS<br />
NOSII/iNOS<br />
common<br />
Tyr<br />
chain<br />
P<br />
SOCS3<br />
IRF-7<br />
Ser484<br />
Ser485 IRF-7<br />
Tyr<br />
Gab2<br />
P Tyr Gab2<br />
P<br />
Tyr<br />
IRS<br />
Ser312(307:R)<br />
P<br />
P Ser21 P<br />
Ser32 Ser374<br />
c-Fos<br />
Ser42 Ser113<br />
P Ser70 P<br />
P<br />
P<br />
P<br />
Ser63 Ser73<br />
c-Jun<br />
P<br />
P Ser21 P<br />
Ser32 Ser374<br />
c-Fos<br />
Ser42 Ser113<br />
P Ser70 P<br />
P<br />
P<br />
Tyr<br />
P<br />
P<br />
Ser63 Ser73<br />
c-Jun<br />
CPT1<br />
acyl CoA<br />
synthetase<br />
Tyr<br />
IRS<br />
Ser312(307:R)<br />
P<br />
P<br />
Ser484<br />
IRF-7<br />
Ser485<br />
P<br />
P<br />
Ser484<br />
IRF-7<br />
Ser485<br />
P<br />
calpain<br />
Tyr706<br />
Tyr544 Tyr974Tyr921<br />
M-CSFR<br />
Tyr559 Tyr807<br />
Tyr721<br />
P Tyr Gab2<br />
AP-1<br />
c-Fos+c-Jun<br />
Tyr580 Tyr542SHP-2<br />
P<br />
P<br />
Thr<br />
Tyr<br />
JNK<br />
Ser133<br />
CREB<br />
Tyr580<br />
nucleus<br />
acetyl CoA<br />
carboxylase<br />
NADPH<br />
Grb2<br />
acetyl CoA carboxylase<br />
xanthine<br />
oxidase<br />
Tyr771<br />
PLC<br />
Tyr783 Tyr1254<br />
PP2B<br />
P<br />
Thr183 ERK1<br />
Tyr185<br />
ERK2<br />
P<br />
P<br />
P<br />
P<br />
P<br />
Ser63 Ser73<br />
Ser63 Ser73<br />
c-Jun<br />
P Tyr Gab2<br />
P<br />
Tyr580 Tyr542SHP-2<br />
P<br />
P<br />
P<br />
Ser369 Thr577<br />
RSK<br />
Ser386 Ser227<br />
P<br />
xanthine<br />
P<br />
Ser<br />
SOS<br />
Thr<br />
Ser63 Ser73<br />
c-Jun<br />
c-jun<br />
P<br />
Ser133<br />
CREB<br />
fatty acid<br />
synthetase<br />
Tyr706<br />
P<br />
P<br />
fatty acid synthetase<br />
NADPH<br />
oxidase<br />
PP2A<br />
c-fos<br />
P<br />
RasGAP<br />
GTP<br />
GDP<br />
Ras<br />
Grb2<br />
Ser21<br />
Ser32 Ser374<br />
c-Fos<br />
Ser42 Ser113<br />
Ser70<br />
Grb2<br />
PI3K<br />
PP2B<br />
Ser<br />
SOS<br />
Thr<br />
acyl CoA synthetase<br />
Fc RIa<br />
chain<br />
?<br />
Ser<br />
SOS<br />
Thr<br />
proteasome<br />
MKP<br />
P<br />
Tyr771<br />
Tyr783<br />
P<br />
P<br />
Ser383<br />
Elk-1<br />
Ser389<br />
P<br />
Site-2 protease<br />
PLC<br />
Pi<br />
Ser383<br />
Ser389 Elk-1<br />
Thr183<br />
Thr183 ERK1<br />
Tyr185ERK2<br />
Tyr185<br />
. O2 -<br />
RXR<br />
RasGAP<br />
SOD<br />
chain<br />
Tyr<br />
Tyr<br />
Tyr1254<br />
P<br />
chain<br />
Tyr<br />
P<br />
Tyr518<br />
GDP<br />
SREBP1c<br />
/bHLH<br />
SREBP1c<br />
/bHLH<br />
GTP<br />
Ras<br />
P<br />
P<br />
Thr183<br />
Thr183 ERK1<br />
Tyr185ERK2<br />
Tyr185<br />
P<br />
P<br />
MPO<br />
LOOH<br />
Golgi<br />
LXR<br />
LXR<br />
Tyr341 Ser4<br />
Raf<br />
Ser338 Ser62<br />
Tyr341 Ser4<br />
Raf<br />
Ser338 Ser62<br />
LXR<br />
27-hydroxyChol<br />
SREBP1c<br />
/bHLH<br />
Tyr518<br />
Tyr518<br />
Syk<br />
Tyr519<br />
Src<br />
P<br />
Ser<br />
Grb2<br />
SOS<br />
Thr<br />
P<br />
P<br />
Ser369<br />
P<br />
P<br />
Ser369<br />
Ser386<br />
P<br />
LXR<br />
SCAP<br />
HOCl<br />
P<br />
P<br />
Thr577<br />
P<br />
R<br />
9<br />
r<br />
Site<br />
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 589<br />
14<br />
<strong>Principles</strong> <strong>of</strong> <strong>cell</strong><br />
<strong>signaling</strong><br />
Melanie H. Cobb and Elliott M. Ross<br />
The University <strong>of</strong> Texas <strong>Southwestern</strong> Medical Center at Dallas<br />
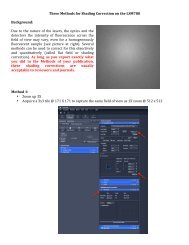
This image represents about 10% <strong>of</strong> the map <strong>of</strong> the known <strong>signaling</strong> interactions and<br />
reactions in the mouse macrophage. Preparing such a map in a computable format is<br />
the first step in analyzing a large <strong>signaling</strong> network. This map was prepared by the group<br />
led by Hiroaki Kitano at the Systems Biology Institute, Tokyo, using their CellDesigner<br />
program. Map courtesy <strong>of</strong> Kanae Oda, Yukiko Matsuoka, and Hiroaki Kitano (The Systems<br />
Biology Institute).<br />
Ub Lys63 TRAF6<br />
Lys63TRAF6<br />
Ub<br />
Lys63TRAF6<br />
Ub<br />
P<br />
Ser385<br />
IRF-3<br />
Ub Lys63 TRAF6<br />
Lys22 I B P<br />
P<br />
Tyr542 SHP-2<br />
Lys22 I B IL-10<br />
Ser385<br />
IRF-3<br />
Lys22 I B PKA<br />
Ser385<br />
IRF-3<br />
P Ser727<br />
TyrJAK1<br />
P<br />
Tyr697<br />
Tyr706<br />
P Tyr Gab2<br />
Tyr542 SHP-2<br />
Thr JNK<br />
Tyr542 SHP-2<br />
Syk<br />
Tyr519<br />
P<br />
Tyr519 Syk<br />
Ser386 R<br />
RSK Ser227<br />
14.1<br />
14.2<br />
14.3<br />
14.4<br />
14.5<br />
14.6<br />
14.7<br />
14.8<br />
14.9<br />
14.10<br />
14.11<br />
14.12<br />
14.13<br />
14.14<br />
14.15<br />
14.16<br />
14.17<br />
14.18<br />
14.19<br />
14.20<br />
CHAPTER O<strong>UT</strong>LINE<br />
Introduction<br />
Cellular <strong>signaling</strong> is primarily chemical<br />
Receptors sense diverse stimuli but initiate a limited<br />
repertoire <strong>of</strong> <strong>cell</strong>ular signals<br />
Receptors are catalysts and amplifiers<br />
Ligand binding changes receptor conformation<br />
Signals are sorted and integrated in <strong>signaling</strong> pathways<br />
and networks<br />
Cellular <strong>signaling</strong> pathways can be thought <strong>of</strong> as<br />
biochemical logic circuits<br />
Scaffolds increase <strong>signaling</strong> efficiency and enhance<br />
spatial organization <strong>of</strong> <strong>signaling</strong><br />
Independent, modular domains specify protein-protein<br />
interactions<br />
Cellular <strong>signaling</strong> is remarkably adaptive<br />
Signaling proteins are frequently expressed as multiple<br />
species<br />
Activating and deactivating reactions are separate and<br />
independently controlled<br />
Cellular <strong>signaling</strong> uses both allostery and covalent<br />
modification<br />
Second messengers provide readily diffusible pathways<br />
for information transfer<br />
Ca2+ <strong>signaling</strong> serves diverse purposes in all eukaryotic<br />
<strong>cell</strong>s<br />
Lipids and lipid-derived compounds are <strong>signaling</strong><br />
molecules<br />
PI 3-kinase regulates both <strong>cell</strong> shape and the activation<br />
<strong>of</strong> essential growth and metabolic functions<br />
Signaling through ion channel receptors is very fast<br />
Nuclear receptors regulate transcription<br />
G protein <strong>signaling</strong> modules are widely used and highly<br />
adaptable<br />
14.21<br />
14.22<br />
14.23<br />
14.24<br />
14.25<br />
14.26<br />
14.27<br />
14.28<br />
14.29<br />
14.30<br />
14.31<br />
14.32<br />
14.33<br />
14.34<br />
14.35<br />
14.36<br />
Tyr JAK1 GM-CSFR<br />
Tyr<br />
JAK2<br />
Tyr1007<br />
Heterotrimeric G proteins regulate a wide variety <strong>of</strong><br />
effectors<br />
NADP + Cl -<br />
P Ser727 STAT1 e<br />
O 2 H 2 O<br />
Heterotrimeric G proteins are controlled P<br />
pyruvate<br />
by a regulatory<br />
-<br />
2<br />
F<br />
hypoxanthine<br />
P Tyr STAT2<br />
NAD +<br />
Fe 3+<br />
H + H +<br />
e<br />
GTPase - Tyr542 SHP-2<br />
cycle<br />
O 2<br />
e -<br />
Small, monomeric GTP-binding proteins are multiuse<br />
switches<br />
Protein phosphorylation/dephosphorylation is a major<br />
regulatory mechanism in the <strong>cell</strong><br />
Two-component protein phosphorylation systems are<br />
<strong>signaling</strong> relays<br />
Pharmacological inhibitors <strong>of</strong> protein kinases may be<br />
used to understand and treat disease<br />
Phosphoprotein phosphatases reverse the actions <strong>of</strong><br />
kinases and are independently regulated<br />
Covalent modification by ubiquitin and ubiquitin-like<br />
proteins is another way <strong>of</strong> regulating protein function<br />
The Wnt pathway regulates <strong>cell</strong> fate during development<br />
and other processes in the adult<br />
Diverse <strong>signaling</strong> mechanisms are regulated by protein<br />
tyrosine kinases<br />
Src family protein kinases cooperate with receptor<br />
protein tyrosine kinases<br />
MAPKs are central to many <strong>signaling</strong> pathways<br />
Cyclin-dependent protein kinases control the <strong>cell</strong> cycle<br />
Diverse receptors recruit protein tyrosine kinases to the<br />
plasma membrane<br />
What’s next?<br />
Summary<br />
References<br />
589
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 590<br />
(GPCR)<br />
G protein<br />
coupled<br />
receptor<br />
Heterotrimeric<br />
G protein<br />
14.1<br />
Introduction<br />
All <strong>cell</strong>s, from prokaryotes through plants and<br />
animals, sense and react to stimuli in their environments<br />
with stereotyped responses that allow<br />
them to survive, adapt, and function in<br />
ways appropriate to the needs <strong>of</strong> the organism.<br />
These responses are not simply direct physical<br />
or metabolic consequences <strong>of</strong> changes in the<br />
local environment. Rather, <strong>cell</strong>s express arrays<br />
<strong>of</strong> sensing proteins, or receptors, that recognize<br />
specific extra<strong>cell</strong>ular stimuli. In response to<br />
these stimuli, receptors regulate the activities<br />
<strong>of</strong> diverse intra<strong>cell</strong>ular regulatory proteins that<br />
in turn initiate appropriate responses by the<br />
<strong>cell</strong>. The process <strong>of</strong> sensing external stimuli and<br />
conveying the inherent information to intra<strong>cell</strong>ular<br />
targets is referred to as <strong>cell</strong>ular signal<br />
transduction.<br />
Cells respond to all sorts <strong>of</strong> stimuli. Microbes<br />
respond to nutrients, toxins, heat, light, and<br />
chemical signals secreted by other microbes.<br />
Cells in multi<strong>cell</strong>ular organisms express receptors<br />
specific for hormones, neurotransmitters,<br />
autocrine and paracrine agents (hormonelike<br />
compounds from the secreting <strong>cell</strong> or <strong>cell</strong>s<br />
Overview <strong>of</strong> major receptor types in a <strong>cell</strong><br />
Receptor<br />
protein<br />
kinase<br />
Ion<br />
channel<br />
Transcription<br />
factor<br />
Twocomponent<br />
complex<br />
(<br />
Sensor<br />
Histidine<br />
kinase<br />
Response<br />
regulator<br />
NUCLEUS<br />
Transmembrane<br />
scaffold<br />
(<br />
E1<br />
E2<br />
E1<br />
E2<br />
Guanylyl<br />
cyclase<br />
FIGURE 14.1 Receptors form a rather small number <strong>of</strong> families that share common<br />
mechanisms <strong>of</strong> action and overall similar structures.<br />
nearby), odors, molecules that regulate growth<br />
or differentiation, and proteins on the outside<br />
<strong>of</strong> adjacent <strong>cell</strong>s. A mammalian <strong>cell</strong> typically<br />
expresses about fifty distinct receptors that sense<br />
different inputs, and, overall, mammals express<br />
several thousand receptors.<br />
Despite the diversity <strong>of</strong> <strong>cell</strong>ular lifestyles<br />
and the enormous number <strong>of</strong> substances sensed<br />
by different <strong>cell</strong>s, the general classes <strong>of</strong> proteins<br />
and mechanisms involved in signal transduction<br />
are conserved throughout living <strong>cell</strong>s, as<br />
shown in FIGURE 14.1.<br />
• G protein-coupled receptors,<br />
composed <strong>of</strong> seven membrane-spanning<br />
helices, promote activation <strong>of</strong> heterotrimeric<br />
GTP-binding proteins called<br />
G proteins, which associate with the inner<br />
face <strong>of</strong> the plasma membrane and<br />
convey signals to multiple intra<strong>cell</strong>ular<br />
proteins.<br />
• Receptor protein kinases are <strong>of</strong>ten<br />
dimers <strong>of</strong> single membrane-spanning<br />
proteins that phosphorylate their intra<strong>cell</strong>ular<br />
substrates and, thus, change<br />
the shape and function <strong>of</strong> the target proteins.<br />
These protein kinases frequently<br />
contain protein interaction domains that<br />
organize complexes <strong>of</strong> <strong>signaling</strong> proteins<br />
on the inner surface <strong>of</strong> the plasma<br />
membrane.<br />
• Phosphoprotein phosphatases reverse<br />
the effect <strong>of</strong> protein kinases by removing<br />
the phosphoryl groups added<br />
by protein kinases.<br />
• Other single membrane-spanning enzymes,<br />
such as guanylyl cyclase, have<br />
an overall architecture similar to the receptor<br />
protein kinases but different enzymatic<br />
activities. Guanylyl cyclase<br />
catalyzes the conversion <strong>of</strong> GTP to 3′:5′-<br />
cyclic GMP, which is used to propagate<br />
the signal.<br />
• Ion channel receptors, although diverse<br />
in detailed structure, are usually<br />
oligomers <strong>of</strong> subunits that each contain<br />
several membrane-spanning segments.<br />
The subunits change their conformations<br />
and relative orientations to permit<br />
ion flux through a central pore.<br />
• Two-component systems may either<br />
be membrane spanning or cytosolic. The<br />
number <strong>of</strong> their subunits is also variable,<br />
but each two-component system<br />
contains a histidine kinase domain or<br />
subunit that is regulated by a <strong>signaling</strong><br />
molecule and a response regulator that<br />
590 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 591<br />
contains a phosphorylatable aspartate<br />
(Asp) residue.<br />
• Some receptors are transmembrane<br />
scaffolds that change either the conformation<br />
or oligomerization <strong>of</strong> their<br />
intra<strong>cell</strong>ular scaffold domains in response<br />
to extra<strong>cell</strong>ular <strong>signaling</strong> molecules,<br />
or ligands, and, thus, recruit<br />
interacting regulatory proteins to a common<br />
site on the membrane.<br />
• Nuclear receptors are transcription<br />
factors, <strong>of</strong>ten heterodimers, that may<br />
reside in the cytoplasm until activated<br />
by agonists or may be permanently located<br />
in the nucleus.<br />
The biochemical processes <strong>of</strong> signal transduction<br />
are strikingly similar among <strong>cell</strong>s.<br />
Bacteria, fungi, plants, and animals use similar<br />
proteins and multiprotein modules to detect<br />
and process signals. For example, evolutionarily<br />
conserved heterotrimeric G proteins and G<br />
protein-coupled receptors are found in plants,<br />
fungi, and animals. Similarly, 3′:5′ cyclic AMP<br />
(cAMP) is an intra<strong>cell</strong>ular <strong>signaling</strong> molecule<br />
in bacteria, fungi, and animals; and Ca2+ serves<br />
a similar role in all eukaryotes. Protein kinases<br />
and phosphoprotein phosphatases are used to<br />
regulate enzymes in all <strong>cell</strong>s.<br />
Although the basic biochemical components<br />
and processes <strong>of</strong> signal transduction are conserved<br />
and reused, they are <strong>of</strong>ten used in wildly<br />
divergent patterns and for many different physiological<br />
purposes. For example, cAMP is synthesized<br />
by distantly related enzymes in bacteria,<br />
fungi, and animals, and acts on different proteins<br />
in each organism; it is a pheromone in<br />
some slime molds.<br />
Cells <strong>of</strong>ten use the same series <strong>of</strong> <strong>signaling</strong><br />
proteins to regulate a given process, such as<br />
transcription, ion transport, locomotion, and<br />
metabolism. Such <strong>signaling</strong> pathways are assembled<br />
into <strong>signaling</strong> networks to allow the<br />
<strong>cell</strong> to coordinate its responses to multiple inputs<br />
with its ongoing functions. It is now possible<br />
to discern conserved reaction sequences<br />
in and between pathways in <strong>signaling</strong> networks<br />
that are analogous to devices within the circuits<br />
<strong>of</strong> analog computers: amplifiers, logic gates,<br />
feedback and feed-forward controls, and memory.<br />
This chapter discusses the principles and<br />
strategies <strong>of</strong> <strong>cell</strong>ular <strong>signaling</strong> first and then discusses<br />
the conserved biochemical components<br />
and reactions <strong>of</strong> <strong>signaling</strong> pathways and how<br />
these principles are applied.<br />
14.2<br />
Cellular <strong>signaling</strong> is<br />
primarily chemical<br />
Key concepts<br />
• Cells can detect both chemical and physical<br />
signals.<br />
• Physical signals are generally converted to<br />
chemical signals at the level <strong>of</strong> the receptor.<br />
Most signals sensed by <strong>cell</strong>s are chemical, and,<br />
when physical signals are sensed, they are generally<br />
detected as chemical changes at the level<br />
<strong>of</strong> the receptor. For example, the visual photoreceptor<br />
rhodopsin is composed <strong>of</strong> the protein<br />
opsin, which binds to a second component,<br />
the colored vitamin A derivative cis-retinal (the<br />
chromophore). When cis-retinal absorbs a<br />
photon, it photoisomerizes to trans-retinal,<br />
which is an activating ligand <strong>of</strong> the opsin protein.<br />
(For more on rhodopsin <strong>signaling</strong> see 14.20<br />
G protein <strong>signaling</strong> modules are widely used and<br />
highly adaptable). Similarly, plants sense red and<br />
blue light using the photosensory proteins phytochrome<br />
and cryptochrome, which detect photons<br />
that are absorbed by their tetrapyrrole or<br />
flavin chromophores. Cryptochrome homologs<br />
are also expressed in animals, where they probably<br />
mediate adjustment <strong>of</strong> the diurnal cycle.<br />
A few receptors do respond directly to physical<br />
inputs. Pressure-sensing channels, which exist<br />
in one form or another in all organisms,<br />
mediate responses to pressure or shear by changing<br />
their ionic conductance. In mammals, hearing<br />
is mediated indirectly by a mechanically<br />
operated channel in the hair <strong>cell</strong> <strong>of</strong> the inner ear.<br />
The extra<strong>cell</strong>ular domain <strong>of</strong> a protein called cadherin<br />
is pulled in response to acoustic vibration,<br />
generating the force that opens the channel.<br />
Cells sense mechanical strain through a<br />
number <strong>of</strong> <strong>cell</strong> surface proteins, including integrins.<br />
Integrins provide signals to <strong>cell</strong>s based on<br />
their attachment to other <strong>cell</strong>s and to molecular<br />
complexes in the external milieu.<br />
One major group <strong>of</strong> physically responsive<br />
receptors is made up <strong>of</strong> channels that sense electric<br />
fields. Another interesting group are<br />
heat/pain-sensing ion channels; several <strong>of</strong> these<br />
heat-sensitive ion channels also respond to<br />
chemical compounds, such as capsaicin, the<br />
“hot” lipid irritant in hot peppers.<br />
Whether a signal is physical or chemical, the<br />
receptor initiates the reactions that change the<br />
behavior <strong>of</strong> the <strong>cell</strong>. We will discuss how these<br />
effects are generated in the rest <strong>of</strong> the chapter.<br />
14.2 Cellular <strong>signaling</strong> is primarily chemical 591
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 592<br />
Ligand A<br />
14.3<br />
Receptors sense diverse<br />
stimuli but initiate a<br />
limited repertoire <strong>of</strong><br />
<strong>cell</strong>ular signals<br />
Key concepts<br />
• Receptors contain a ligand-binding domain and an<br />
effector domain.<br />
• Receptor modularity allows a wide variety <strong>of</strong><br />
signals to use a limited number <strong>of</strong> regulatory<br />
mechanisms.<br />
• Cells may express different receptors for the same<br />
ligand.<br />
• The same ligand may have different effects on the<br />
<strong>cell</strong> depending on the effector domain <strong>of</strong> its<br />
receptor.<br />
Receptors mediate responses to amazingly diverse<br />
extra<strong>cell</strong>ular messenger molecules; hence,<br />
the <strong>cell</strong> must express a large number <strong>of</strong> receptor<br />
varieties, each able to bind its extra<strong>cell</strong>ular<br />
ligand. In addition, each receptor must be able<br />
to initiate a <strong>cell</strong>ular response. Receptors, thus,<br />
contain two functional domains: a ligandbinding<br />
domain and an effector domain,<br />
which may or may not correspond to definable<br />
structural domains within the protein.<br />
The separation <strong>of</strong> ligand-binding and effector<br />
functions allows receptors for diverse ligands<br />
to produce a limited number <strong>of</strong> evolutionarily<br />
conserved intra<strong>cell</strong>ular signals through the action<br />
<strong>of</strong> a few effector domains. In fact, there are<br />
Receptors have a ligand-binding domain and an effector domain<br />
Output<br />
1<br />
Output<br />
2<br />
Ligand A<br />
Output<br />
1<br />
Ligand B<br />
Output<br />
1<br />
CHIMERIC<br />
RECEPTOR<br />
Ligand C<br />
LBD1 LBD1 LBD1 LBD2 LBD3<br />
ED1 ED2<br />
ED1 ED1<br />
Output<br />
2<br />
ED2<br />
FIGURE 14.2 Receptors can be thought <strong>of</strong> as composed <strong>of</strong> two functional domains,<br />
a ligand-binding domain (LBD) and an effector domain (ED). The twodomain<br />
property implies that two receptors that respond to different ligands<br />
(middle) could initiate the same function by activating similar effector domains,<br />
or that a <strong>cell</strong> could express two receptor is<strong>of</strong>orms (left) that respond to<br />
the same ligand with distinct <strong>cell</strong>ular effects mediated by different effector domains.<br />
It also implies that one can create an artificial chimeric receptor with<br />
novel properties.<br />
only a limited number <strong>of</strong> receptor families, which<br />
are related by their conserved structures and <strong>signaling</strong><br />
functions (see Figure 14.1).<br />
There are several useful correlates to the<br />
two-domain nature <strong>of</strong> receptors. For example,<br />
a <strong>cell</strong> can control its responsiveness to an extra<strong>cell</strong>ular<br />
signal by regulating the synthesis or<br />
degradation <strong>of</strong> a receptor or by regulating the<br />
receptor’s activity (see 14.10 Cellular <strong>signaling</strong> is<br />
remarkably adaptive).<br />
In addition, the nature <strong>of</strong> a response is generally<br />
determined by the receptor and its effector<br />
domain rather than any physicochemical<br />
property <strong>of</strong> the ligand. FIGURE 14.2 illustrates the<br />
concept that a ligand may bind to more than<br />
one kind <strong>of</strong> receptor and elicit more than one<br />
type <strong>of</strong> response, or several different ligands<br />
may all act identically by binding to functionally<br />
similar receptors. For example, the neurotransmitter<br />
acetylcholine binds to two classes<br />
<strong>of</strong> receptors. Members <strong>of</strong> one class are ion channels;<br />
members <strong>of</strong> the other regulate G proteins.<br />
Similarly, steroid hormones bind both to nuclear<br />
receptors, which bind chromatin and regulate<br />
transcription, and to other receptors in<br />
the plasma membrane.<br />
Conversely, when multiple ligands bind to<br />
receptors <strong>of</strong> the same biochemical class, they<br />
generate similar intra<strong>cell</strong>ular responses. For example,<br />
it is not uncommon for a <strong>cell</strong> to express<br />
several distinct receptors that stimulate production<br />
<strong>of</strong> the intra<strong>cell</strong>ular <strong>signaling</strong> molecule cAMP.<br />
The effect <strong>of</strong> the receptor on the <strong>cell</strong> will also be<br />
determined significantly by the biology <strong>of</strong> the<br />
<strong>cell</strong> and its state at any given time.<br />
Ligand binding and effector domains may<br />
evolve independently in response to varied selective<br />
pressures. For example, mammalian and<br />
invertebrate rhodopsins transduce their signal<br />
through different effector G proteins (G t<br />
and<br />
G q<br />
, respectively). Another example is calmodulin,<br />
a small calcium-binding regulatory protein<br />
in animals, which in plants appears as a<br />
distinct domain in larger proteins.<br />
The receptor’s two-domain nature allows<br />
the <strong>cell</strong> to regulate the binding <strong>of</strong> ligand and<br />
the effect <strong>of</strong> ligand independently. Covalent<br />
modification or allosteric regulation can alter<br />
ligand-binding affinity, the ability <strong>of</strong> the ligand-bound<br />
receptor to generate its signal or<br />
both. We will discuss these concepts further in<br />
14.13 Cellular <strong>signaling</strong> uses both allostery and covalent<br />
modification.<br />
Receptors can be classified either according<br />
to the ligands they bind or the way in which<br />
they signal. Signal output, which is character-<br />
592 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 593<br />
istic <strong>of</strong> the effector domain, usually correlates<br />
best with overall structure and sequence conservation.<br />
(Receptor families grouped by their<br />
functions are the organizational basis <strong>of</strong> the second<br />
half <strong>of</strong> this chapter.) However, classifying<br />
receptors pharmacologically, according to their<br />
specificity for ligands, is particularly useful for<br />
understanding the organization <strong>of</strong> endocrine<br />
and neuronal systems and for categorizing the<br />
multiple physiological responses to drugs.<br />
Expression <strong>of</strong> a receptor that is not normally<br />
expressed in a <strong>cell</strong> is <strong>of</strong>ten sufficient to<br />
confer responsiveness to that receptor’s ligand.<br />
This responsiveness <strong>of</strong>ten occurs because the<br />
<strong>cell</strong> expresses the other components necessary<br />
for propagating the intra<strong>cell</strong>ular signal from the<br />
receptor. The precise nature <strong>of</strong> the response will<br />
reflect the biology <strong>of</strong> the <strong>cell</strong>. Experimentally,<br />
responsiveness to a compound can be induced<br />
by introducing the cDNA that encodes the receptor.<br />
For example, mammalian receptors may<br />
be expressed in yeast, such that the yeast respond<br />
visibly to receptor ligands, thus providing<br />
a way to screen for new chemicals (drugs)<br />
that activate the receptor.<br />
Finally, it is possible to create chimeric receptors<br />
by fusing the ligand-binding domain<br />
from one receptor with the effector domain<br />
from a different receptor (Figure 14.2). Such<br />
chimeras can mediate novel responses to the<br />
ligand. With genetic modification <strong>of</strong> the ligandbinding<br />
domain, receptors can be reengineered<br />
to respond to novel ligands. Thus, scientists can<br />
manipulate <strong>cell</strong> functions with nonbiological<br />
compounds.<br />
14.4<br />
Receptors are catalysts<br />
and amplifiers<br />
Key concepts<br />
• Receptors act by increasing the rates <strong>of</strong> key<br />
regulatory reactions.<br />
• Receptors act as molecular amplifiers.<br />
Receptors act to accelerate intra<strong>cell</strong>ular functions<br />
and are, thus, functionally analogous to enzymes<br />
or other catalysts. Some receptors,<br />
including the protein kinases, protein phosphatases,<br />
and guanylate cyclases, are themselves<br />
enzymes and thus classical biochemical catalysts.<br />
More generally, however, receptors use<br />
the relatively small energy <strong>of</strong> ligand binding to<br />
accelerate reactions that are driven by alternative<br />
energy sources. For example, receptors that<br />
are ion channels catalyze the movement <strong>of</strong> ions<br />
across membranes, a process driven by the electrochemical<br />
potential developed by distinct ion<br />
pumps. G protein-coupled receptors and other<br />
guanine nucleotide exchange factors catalyze<br />
the exchange <strong>of</strong> GDP for GTP on the G protein,<br />
an energetically favored process dictated by the<br />
<strong>cell</strong>’s nucleotide energy balance. Transcription<br />
factors accelerate the formation <strong>of</strong> the transcriptional<br />
initiation complex, but transcription itself<br />
is energetically driven by multiple steps <strong>of</strong><br />
ATP and dNTP hydrolysis.<br />
As catalysts, receptors enhance the rates <strong>of</strong><br />
reactions. Most <strong>signaling</strong> involves kinetic rather<br />
than thermodynamic regulation; that is, <strong>signaling</strong><br />
events change reaction rates rather than<br />
their equilibria (see the next section). Thus, <strong>signaling</strong><br />
is similar to metabolic regulation, in<br />
which specific reactions are chosen according to<br />
their rates, with thermodynamic driving forces<br />
playing only a supportive role.<br />
In all <strong>signaling</strong> reactions, receptors use their<br />
catalytic activities to function as molecular amplifiers.<br />
Directly or indirectly, a receptor generates<br />
a chemical signal that is huge, both<br />
energetically and with respect to the number<br />
<strong>of</strong> molecules recruited by a single receptor.<br />
Molecular amplification is a hallmark <strong>of</strong> receptors<br />
and many other steps in <strong>cell</strong>ular <strong>signaling</strong><br />
pathways.<br />
14.5<br />
Ligand binding changes<br />
receptor conformation<br />
Key concepts<br />
• Receptors can exist in active or inactive<br />
conformations.<br />
• Ligand binding drives the receptor toward the<br />
active conformation.<br />
A central mechanistic question in receptor function<br />
is how the binding <strong>of</strong> a <strong>signaling</strong> molecule<br />
to the ligand-binding domain increases the activity<br />
<strong>of</strong> the effector domain. The key to this<br />
question is that receptors can exist in multiple<br />
molecular conformations, some active for <strong>signaling</strong><br />
and others inactive. Ligands shift the<br />
conformational equilibrium among these conformations.<br />
The structural changes that occur<br />
during the receptor’s inactive-active isomerization<br />
and how ligand binding drives these<br />
changes are exciting areas <strong>of</strong> biophysical research.<br />
However, the basic concept can be described<br />
simply in terms <strong>of</strong> coupling the<br />
conformational isomerizations <strong>of</strong> the ligandbinding<br />
and effector domains.<br />
14.5 Ligand binding changes receptor conformation 593
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 594<br />
How do ligands activate (or not activate) a<br />
receptor? Most <strong>of</strong> the basic regulatory activities<br />
<strong>of</strong> receptors can be described by a simple scheme<br />
that considers the receptors as having two interconvertible<br />
conformations, inactive (R) and<br />
active (R*). R and R* are in equilibrium, which<br />
is described by the equilibrium constant J.<br />
Because unliganded receptors are usually<br />
minimally active, J1), then ligand binding<br />
will shift the conformation to the R* state to an<br />
equivalent extent (i.e., J*/J>>1). The relative<br />
activation by a saturating concentration <strong>of</strong> ligand,<br />
J*/J, will exactly equal the ligand’s relative<br />
selectivity for the active receptor conformation,<br />
K*/K. This argument is generally valid for the regulation<br />
<strong>of</strong> a protein’s activity by any regulatory<br />
ligand.<br />
This model explains many properties <strong>of</strong> receptors<br />
and their ligands both simply and quantitatively.<br />
• First, J must be greater than zero for the<br />
equilibrium to exist. Thus, even unliganded<br />
receptor has some activity.<br />
Overexpressed receptors frequently display<br />
their intrinsic low activity.<br />
• Because physiological receptors are<br />
nearly inactive in the absence <strong>of</strong> ligand,<br />
J must be much less than 1 and is probably<br />
less than 0.01; most receptors are<br />
less than 1% active without agonist.<br />
• Ligands can vary in their selectivities<br />
between R and R*. Their abilities to activate<br />
will also vary. Some ligands, referred<br />
to as agonists, can drive formation<br />
<strong>of</strong> appreciable R*. Others, known as partial<br />
agonists, will promote submaximal<br />
activation. Chemical manipulation<br />
<strong>of</strong> a ligand’s structure will <strong>of</strong>ten alter its<br />
activity as an agonist. These relationships<br />
are depicted graphically in FIGURE<br />
14.3.<br />
• A ligand that binds equally well to both<br />
the R and R* states will not cause activation.<br />
However, such a ligand may still<br />
occupy the binding site and thereby<br />
competitively inhibit binding <strong>of</strong> an activating<br />
ligand. Such competitive inhibitors,<br />
referred to as antagonists, are<br />
frequently used as drugs to block unwanted<br />
activation <strong>of</strong> a receptor in various<br />
disease states.<br />
• A ligand that binds preferentially to R<br />
594 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 595<br />
relative to R* will further shift the conformational<br />
equilibrium to the inactive<br />
state and cause net inhibition. Such ligands<br />
are called inverse agonists.<br />
Because J is already low, effects <strong>of</strong> inverse<br />
agonists may only be noticeable<br />
if a receptor is overexpressed or if the<br />
receptor is mutated to increase its intrinsic<br />
activity (i.e., the mutation increases<br />
J).<br />
• The extent to which an agonist stimulates<br />
a receptor is unrelated to its affinity.<br />
Both agonists and antagonists may<br />
bind with either high or low affinity.<br />
Affinity does determine the receptor’s<br />
sensitivity—that is, how low a concentration<br />
<strong>of</strong> ligand can the receptor detect.<br />
Affinities <strong>of</strong> receptors for natural regulatory<br />
ligands vary enormously, with<br />
physiologic K d<br />
values ranging from<br />
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 596<br />
RECEPTORS<br />
TRANSDUCERS<br />
EFFECTORS<br />
Convergent and divergent <strong>signaling</strong> pathways<br />
Linear,<br />
parallel<br />
Convergent Divergent Multiply<br />
branched<br />
FIGURE 14.4 Signaling pathways use convergent and divergent branching to coordinate<br />
information flow. The diagrams at top show how even a simple, threelevel<br />
<strong>signaling</strong> network can sort information. Convergence or divergence can<br />
take place at multiple points along a <strong>signaling</strong> pathway. As an example <strong>of</strong> complexity,<br />
the lower portion <strong>of</strong> the figure shows a small segment (~10%) <strong>of</strong> the G<br />
protein-mediated <strong>signaling</strong> network in a mouse macrophage <strong>cell</strong> line. It omits<br />
several interpathway regulatory mechanisms and completely ignores inputs from<br />
non-G protein-coupled receptors. Pathway map courtesy <strong>of</strong> Lily Jiang, University<br />
<strong>of</strong> Texas <strong>Southwestern</strong> Medical Center.<br />
maps. Signaling networks are also spatially complex.<br />
They may include components in various<br />
sub<strong>cell</strong>ular locations, with initial receptors and<br />
associated proteins in the plasma membrane, but<br />
with downstream proteins in the cytoplasm or intra<strong>cell</strong>ular<br />
organelles. Such complexity is necessary<br />
to allow the <strong>cell</strong>s to integrate and sort<br />
incoming signals and to regulate multiple intra<strong>cell</strong>ular<br />
functions simultaneously.<br />
The complexity and adaptability <strong>of</strong> <strong>signaling</strong><br />
networks, like the one shown in the lower<br />
half <strong>of</strong> Figure 14.4, make their dynamics at the<br />
whole-<strong>cell</strong> level difficult or impossible to grasp<br />
intuitively. Signaling networks resemble large<br />
analog computers, and investigators are increasingly<br />
depending on computational tools to understand<br />
<strong>cell</strong>ular information flow and its<br />
regulation. First, many <strong>signaling</strong> interactions<br />
that include only two or three proteins exert<br />
functions analogous to traditional computational<br />
logic circuits (see the next section). The<br />
theory and experience with such circuits in electronics<br />
facilitate understanding biological <strong>signaling</strong><br />
functions as well.<br />
The enormous complexity <strong>of</strong> <strong>cell</strong>ular <strong>signaling</strong><br />
networks can be simplified by considering<br />
them to be composed <strong>of</strong> interacting <strong>signaling</strong><br />
modules, i.e., groups <strong>of</strong> proteins that process signals<br />
in well-understood ways. A <strong>cell</strong>ular <strong>signaling</strong><br />
module is analogous to an integrated circuit<br />
in an electronic instrument that performs a<br />
known function, but whose exact components<br />
could be changed for similar use in another device.<br />
The concept <strong>of</strong> modular construction facilitates<br />
both qualitative and quantitative<br />
understanding <strong>of</strong> <strong>signaling</strong> networks. We will refer<br />
to many standard <strong>signaling</strong> modules later in<br />
the chapter. Examples include monomeric and<br />
heterotrimeric G protein modules, MAPK cascades,<br />
tyrosine (Tyr) kinase receptors and their<br />
binding proteins, and Ca2+ release/uptake modules.<br />
In each case, despite the numerous phylogenetic,<br />
developmental, and physiologic<br />
variations, understanding the basic function <strong>of</strong><br />
that class <strong>of</strong> module conveys understanding <strong>of</strong> all<br />
its incarnations. Last, the evolutionary importance<br />
<strong>of</strong> modules is significant; once the architecture<br />
<strong>of</strong> a module is established it can be reused.<br />
For larger-scale networks, multiplexed,<br />
high-throughput measurements on living <strong>cell</strong>s<br />
have been combined with powerful kinetic modeling<br />
strategies to allow an increasingly accurate<br />
quantitative depiction <strong>of</strong> information flow<br />
within <strong>signaling</strong> modules or entire networks.<br />
Such models, with sound and experimentally<br />
based parameter sets, can describe <strong>signaling</strong><br />
processes in systems too complex for intuitive<br />
or ad hoc analysis. They are also vital as tests <strong>of</strong><br />
understanding because they can predict experimental<br />
results in ways that can be used to test<br />
the validity <strong>of</strong> the model. Well-grounded models<br />
can then be used (cautiously) to suggest the<br />
mechanisms <strong>of</strong> systems for which data sets remain<br />
unattainable. At even greater levels <strong>of</strong><br />
complexity, the theories and tools <strong>of</strong> computer<br />
science are increasingly giving useful systemslevel<br />
analyses <strong>of</strong> signal flow in <strong>cell</strong>s. Using computational<br />
tools to analyze large arrays <strong>of</strong><br />
quantitative data allows us to understand <strong>cell</strong>ular<br />
information flow and its regulation.<br />
596 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 597<br />
Developing quantitative models <strong>of</strong> <strong>signaling</strong><br />
networks is a frontier in <strong>signaling</strong> biology. These<br />
models both help describe network function<br />
and pinpoint experiments to clarify mechanism.<br />
14.7<br />
Cellular <strong>signaling</strong><br />
pathways can be thought<br />
<strong>of</strong> as biochemical logic<br />
circuits<br />
Key concepts<br />
• Signaling networks are composed <strong>of</strong> groups <strong>of</strong><br />
biochemical reactions that function as<br />
mathematical logic functions to integrate<br />
information.<br />
• Combinations <strong>of</strong> such logic functions combine as<br />
<strong>signaling</strong> networks to process information at more<br />
complex levels.<br />
As introduced in the preceding section, processes<br />
that <strong>signaling</strong> pathways use to integrate and direct<br />
information to <strong>cell</strong>ular targets are strikingly analogous<br />
to the mathematical logic functions that are<br />
used to design the individual circuits <strong>of</strong> electronic<br />
computers. Indeed, there are biological equivalents<br />
<strong>of</strong> essentially all <strong>of</strong> the functional components<br />
that computer scientists and engineers<br />
consider in the design <strong>of</strong> computers and electronic<br />
control devices. To understand <strong>signaling</strong> pathways,<br />
it is, therefore, useful to consider groups <strong>of</strong><br />
reactions within a pathway as constituting logic circuits<br />
<strong>of</strong> the sort used in electronic computing, as<br />
illustrated in FIGURE 14.5. The simplest example is<br />
when two stimulatory pathways converge. If sufficient<br />
input from either is adequate to elicit the<br />
response, the convergence would constitute an<br />
“OR” function. If neither input is sufficient by itself<br />
but the combination <strong>of</strong> the two elicits the response,<br />
then the converging pathways would<br />
create “AND” functions. AND circuits are also referred<br />
to as coincidence detectors—a response<br />
is elicited only when two stimulating pathways<br />
are activated simultaneously.<br />
AND functions can result from the combination<br />
<strong>of</strong> two similar but quantitatively inadequate<br />
inputs. Alternatively, two mechanistically<br />
different inputs might both be required to elicit<br />
a response. An example <strong>of</strong> the latter would be<br />
a target protein that is allosterically activated<br />
only when phosphorylated, or that is activated<br />
by phosphorylation but is only functional when<br />
recruited to a specific sub<strong>cell</strong>ular location.<br />
The opposite <strong>of</strong> an AND circuit is a NOT<br />
function, where one pathway blocks the stim-<br />
Logical (Boolean)<br />
A<br />
B<br />
A + B<br />
A<br />
B<br />
A + B<br />
A<br />
B<br />
A + B<br />
A OR B<br />
A AND B<br />
A NOT B<br />
Response<br />
Response<br />
Response<br />
Response<br />
Response<br />
Simple logic circuits<br />
Response<br />
Quantitative (Analog)<br />
A + fixed [B]<br />
Response<br />
Response<br />
Additive<br />
ulatory effect <strong>of</strong> another. Simple logic gates are<br />
observed at many locations in <strong>cell</strong>ular <strong>signaling</strong><br />
pathways.<br />
We can also think about convergent <strong>signaling</strong><br />
in quantitative rather than Boolean terms<br />
by considering the additivity <strong>of</strong> inputs to a distinct<br />
process (see Figure 14.5, right). The OR<br />
function referred to above can be considered to<br />
be the additive positive inputs <strong>of</strong> two pathways.<br />
Such additivity could represent the ability <strong>of</strong><br />
several receptors to stimulate a pool <strong>of</strong> a particular<br />
G protein or the ability <strong>of</strong> two protein kinases<br />
to phosphorylate a single substrate.<br />
Additivity may be positive, as in the examples<br />
above, or negative, such as when two inhibitory<br />
inputs combine. Inhibition and stimulation may<br />
also combine additively to yield an algebraically<br />
balanced output. Alternatively, multiple inputs<br />
can combine with either more or less than an<br />
additive effect. The NOT function, discussed<br />
above, is analogous to describing a blockade <strong>of</strong><br />
stimulation. The AND function describes synergism,<br />
where one input potentiates another<br />
but alone has little effect.<br />
Even simple <strong>signaling</strong> networks can display<br />
complex patterns <strong>of</strong> information processing. One<br />
A<br />
log (agonist concentration)<br />
More than additive<br />
log (agonist concentration)<br />
Less than additive<br />
log (agonist concentration)<br />
B<br />
A + B<br />
A<br />
B<br />
A<br />
A + B<br />
B<br />
FIGURE 14.5 Signaling networks use simple logic functions to process<br />
information. Boolean OR, AND, and NOT functions (left) correspond to<br />
the quantitative interactions between converging signals that are shown<br />
on the right.<br />
14.7 Cellular <strong>signaling</strong> pathways can be thought <strong>of</strong> as biochemical logic circuits 597
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 598<br />
Input<br />
Positive feedback loop : irreversible ON switch<br />
Positive feed-forward loop : responds to prolonged input<br />
Input<br />
OH<br />
E<br />
G<br />
T<br />
T<br />
+<br />
Output<br />
Output<br />
Output<br />
Input strength<br />
Conformational lock - Dual control switch<br />
OH<br />
E<br />
OH<br />
+<br />
Kinase<br />
E<br />
E<br />
G<br />
G<br />
Phosphatase<br />
Signal processing circuits<br />
E<br />
E<br />
G<br />
P<br />
E<br />
P<br />
P<br />
E<br />
Output<br />
Output<br />
Time<br />
Time<br />
input<br />
G K G P K<br />
FIGURE 14.6 Relatively complex signal processing can be executed by simple<br />
multi-protein modules. The figure depicts three types <strong>of</strong> <strong>signaling</strong> modules<br />
(left) and their behavior in response to agonist (right). (top) In a positive<br />
feed-back module, a transducer protein (T) stimulates an effector (E) to produce<br />
a <strong>cell</strong>ular output, but the effector also stimulates the activity <strong>of</strong> the transducer.<br />
The result can be an all-or-none switch, where input up to a threshold<br />
has little effect, but then becomes committed when feedback from the effector<br />
is sufficient to maintain transducer activity even in the absence <strong>of</strong> continued<br />
input from the receptor. (center) In a positive feed-forward module, the<br />
effector requires input both from the transducer and from upstream in the pathway.<br />
When stimulation is brief (short horizontal bar under trace at right), significant<br />
amounts <strong>of</strong> active transducer do not accumulate and output is minimal.<br />
When stimulation is prolonged (longer bar), signal output is substantial. (bottom)<br />
In some dual-control switching modules, the binding <strong>of</strong> one regulator (G)<br />
can both activate the effector and expose another regulatory site, shown here<br />
as a Ser substrate site (-OH) for a protein kinase. The effector can only be phosphorylated<br />
or dephosphorylated when G is bound. Therefore, as shown at the<br />
right, addition <strong>of</strong> G alone will activate but activation <strong>of</strong> the kinase (K) alone<br />
will not. If kinase is active while G is bound, phosphorylation is resistant to<br />
phosphatase activity unless G is again present to reexpose the phosphoserine<br />
residue (shown on the graph at the right as a bold P).<br />
G P<br />
good example is the creation <strong>of</strong> “memory”: making<br />
the effect <strong>of</strong> a transient signal more or less<br />
permanent. Signaling pathways have multiple<br />
ways <strong>of</strong> setting memories, and <strong>of</strong> forgetting. One<br />
mechanism, common in protein kinase pathways,<br />
is the positive feedback loop, illustrated<br />
in the top panel <strong>of</strong> FIGURE 14.6. In a positive feedback<br />
loop, the input stimulates a transducer (T),<br />
which in turn stimulates the effector protein (E)<br />
to create the output. If the effector can also activate<br />
the transducer, sufficient initial signal can<br />
be fed back to the transducer that it can maintain<br />
the effector's full signal output even when<br />
input is removed. Such systems typically display<br />
a threshold behavior, as shown on the right.<br />
A positive feed-forward loop can generate<br />
memory <strong>of</strong> another type (Figure 14.6, middle<br />
panel), indicating the duration <strong>of</strong> input. In such<br />
circuits, the effector requires simultaneous input<br />
from both the receptor and from the intermediary<br />
transducer. If the pathway from<br />
receptor through transducer is relatively slow,<br />
or if it requires the accumulation <strong>of</strong> a substantial<br />
amount <strong>of</strong> transducer, only a prolonged input<br />
will trigger a response, as shown in the<br />
time-base output diagram at the right.<br />
A third way to establish memory is to allow<br />
one input to control the reversibility <strong>of</strong> a second<br />
regulatory event (Figure 14.6, bottom panel).<br />
WASP, a protein that initiates the polymerization<br />
<strong>of</strong> actin to drive <strong>cell</strong>ular motion and shape<br />
change, is activated both by phosphorylation<br />
and by the binding <strong>of</strong> Cdc42, a small GTP-binding<br />
protein (G). However, the phosphorylation<br />
site on WASP is only exposed when WASP is<br />
bound to Cdc42. Phosphorylation thus requires<br />
both activated Cdc42 and activated protein kinase.<br />
If Cdc42 dissociates, the phosphorylated<br />
state <strong>of</strong> WASP persists until another <strong>signaling</strong><br />
molecule, whose identity remains uncertain,<br />
binds again to expose the site to a protein phosphatase.<br />
As shown in the time-base graph, exposure<br />
to Cdc42 will activate, but exposure to<br />
kinase alone will not. If Cdc42 is present, then<br />
the kinase can activate WASP. Phospho-WASP<br />
is relatively insensitive to protein phosphatase<br />
(P) alone, but can be dephosphorylated if Cdc42<br />
or another G protein binds to expose the site to<br />
phosphatase.<br />
14.8<br />
Scaffolds increase<br />
<strong>signaling</strong> efficiency and<br />
enhance spatial<br />
organization <strong>of</strong> <strong>signaling</strong><br />
Key concepts<br />
• Scaffolds organize groups <strong>of</strong> <strong>signaling</strong> proteins and<br />
may create pathway specificity by sequestering<br />
components that have multiple partners.<br />
• Scaffolds increase the local concentration <strong>of</strong><br />
<strong>signaling</strong> proteins.<br />
• Scaffolds localize <strong>signaling</strong> pathways to sites <strong>of</strong><br />
action.<br />
598 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 599<br />
The INAD <strong>signaling</strong> complex<br />
TRP<br />
Rhodopsin<br />
Scaffolds concentrate and insulate <strong>signaling</strong> proteins<br />
Pheromone<br />
GPCR<br />
PKC<br />
PDZ PDZ<br />
PDZ<br />
INAD<br />
-<br />
PDZ PDZ<br />
CaM<br />
CaM<br />
PKC<br />
PDZ<br />
PDZ<br />
PDZ<br />
INAD<br />
PDZ PDZ<br />
CYTOSOL<br />
FIGURE 14.7 The scaffold InaD organizes proteins that transmit visual<br />
signals in the fly photoreceptor <strong>cell</strong>. InaD is localized to the photoreceptor<br />
membrane and coordinates light sensing and visual transduction. In<br />
invertebrate eyes, the visual <strong>signaling</strong> pathway goes from rhodopsin<br />
through G q<br />
to a phospholipase C-, and Ca2+ release triggered by PLC action<br />
initiates depolarization. This system is specialized for speed, and requires<br />
that the relevant proteins are nearby. InaD contains five PDZ<br />
domains, each <strong>of</strong> which binds to the C terminus <strong>of</strong> a signal transducing<br />
protein. The TRP channel, which mediates Ca2+ entry, PLC-, and a protein<br />
kinase C is<strong>of</strong>orm that is involved in rapid desensitization all bind constitutively<br />
to InaD. Rhodopsin and a myosin (NinaC) also bind, and G q<br />
binds indirectly.<br />
The proteins in a <strong>signaling</strong> pathway are frequently<br />
colocalized within <strong>cell</strong>s such that their<br />
mutual interactions are favored and their interactions<br />
with other proteins are minimized.<br />
Many <strong>signaling</strong> pathways are organized on scaffolds.<br />
Scaffolds bind several components <strong>of</strong> a<br />
<strong>signaling</strong> pathway in multiprotein complexes<br />
to enhance <strong>signaling</strong> efficiency. Scaffolds promote<br />
interactions <strong>of</strong> proteins that have a low<br />
affinity for each other, accelerate activation (and<br />
<strong>of</strong>ten inactivation) <strong>of</strong> the associated components,<br />
and localize the <strong>signaling</strong> proteins to appropriate<br />
sites <strong>of</strong> action. Colocalization may be<br />
tonic or regulated, and stimulus-dependent scaffolding<br />
<strong>of</strong>ten determines <strong>signaling</strong> outputs.<br />
The binding sites on a scaffolding protein<br />
are <strong>of</strong>ten localized in distinct modular proteinbinding<br />
domains, giving the impression that the<br />
protein is designed simply to hold the components<br />
<strong>of</strong> the pathway together. Many scaffolding<br />
proteins do lack intrinsic enzymatic activity,<br />
but some <strong>signaling</strong> enzymes also act as scaffolds.<br />
Binding to a scaffold facilitates <strong>signaling</strong> by<br />
increasing the local concentrations <strong>of</strong> the components,<br />
so that diffusion or transport <strong>of</strong> molecules<br />
to their sites <strong>of</strong> action is not necessary. In<br />
the photoreceptor <strong>cell</strong>s <strong>of</strong> Drosophila, scaffolding<br />
<strong>of</strong> <strong>signaling</strong> components is critical for rapid<br />
signal transmission. These <strong>cell</strong>s contain the InaD<br />
-<br />
Ste5p<br />
Ste11p<br />
Ste7p<br />
Fus3p<br />
G protein<br />
Pheromone<br />
Ste5p<br />
Mating<br />
response<br />
Ste11p<br />
Ste7p<br />
Fus3p<br />
Cdc42p<br />
Ste20p<br />
Scaffold organizes<br />
MAPK cascade<br />
Cdc42p<br />
Ste20p<br />
scaffolding protein, which has five modular<br />
binding domains, known as PDZ domains. Each<br />
<strong>of</strong> its PDZ domains binds to a C-terminal motif<br />
<strong>of</strong> a target protein, thereby facilitating interactions<br />
among the associated proteins. FIGURE 14.7<br />
shows a model for how InaD organizes the <strong>signaling</strong><br />
proteins. The mutational loss <strong>of</strong> InaD<br />
produces a nearly blind fly, and deletion <strong>of</strong> a<br />
single PDZ domain can yield a fly with a distinct<br />
visual defect characteristic <strong>of</strong> the protein<br />
that binds to the missing domain.<br />
A second example is Ste5p, a scaffold for the<br />
pheromone-induced mating response pathway<br />
in S. cerevisiae. FIGURE 14.8 illustrates how Ste5p<br />
binds and organizes components <strong>of</strong> a mitogen-<br />
Scaffold determines<br />
specificity <strong>of</strong> Ste11p<br />
<strong>signaling</strong><br />
Ste11p<br />
Ste7p<br />
Fus3p<br />
Mating<br />
response<br />
High osmolarity<br />
Ste11p<br />
Pbs2p<br />
Hog1p<br />
Osmoadaptation<br />
Cdc42p<br />
Ste20p<br />
Cdc42p<br />
Ste20p<br />
FIGURE 14.8 The scaffold Ste5p organizes the components <strong>of</strong> the MAPK<br />
cascade that mediates the pheromone-induced mating response in<br />
Saccharomyces cerevisiae. In the top left panel, Ste5p brings the components<br />
<strong>of</strong> the MAPK cascade to the membrane in response to pheromone. In<br />
the top right panel, binding to the heterotrimeric G protein brings loaded<br />
Ste5p in proximity to the protein kinase Ste20p bound to the activated small<br />
GTP binding protein Cdc42p. Their colocalization facilitates the sequential<br />
activation <strong>of</strong> the cascade components, resulting in activation <strong>of</strong> the MAPK<br />
Fus3p and the mating response. The MAP3K Ste11p can regulate not only<br />
the MAPK Fus3p in the mating pathway, but also the MAPK Hog1p in the<br />
high osmolarity pathway, as shown in the bottom two panels. The scaffold<br />
to which Ste11p binds, either Ste5p or Pbs2 (both a scaffold and a MAP2K),<br />
determines which MAPK and downstream events are activated as the output.<br />
14.8 Scaffolds increase <strong>signaling</strong> efficiency and enhance spatial organization <strong>of</strong> <strong>signaling</strong> 599
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 600<br />
activated protein kinase (MAPK) cascade, including<br />
a MAP3K (Ste11p), a MAP2K (Ste7p)<br />
and a MAPK (Fus3p). (The MAPK cascade will<br />
be discussed in 14.32 MAPKs are central to many<br />
<strong>signaling</strong> pathways). The function <strong>of</strong> Ste5p is partially<br />
retained even if the positions <strong>of</strong> its binding<br />
sites for the kinases are shuffled in the linear<br />
sequence <strong>of</strong> the protein, indicating that a major<br />
role is to bring the enzymes into proximity, rather<br />
than to precisely orient them. Ste5p also binds<br />
to the subunits <strong>of</strong> the heterotrimeric G protein<br />
that mediates the actions <strong>of</strong> mating<br />
pheromones, linking the membrane signal to<br />
the intra<strong>cell</strong>ular transducers. Yeast that lack<br />
Ste5p cannot mate, demonstrating that Ste5p is<br />
required for this biological function (but not all<br />
functions) carried out by the pathway.<br />
In addition to facilitating <strong>signaling</strong> in their<br />
own pathways, scaffolds can enhance <strong>signaling</strong><br />
specificity by limiting interactions with other<br />
<strong>signaling</strong> proteins. Scaffolds thus insulate components<br />
<strong>of</strong> a <strong>signaling</strong> pathway both from activation<br />
by inappropriate signals and from<br />
producing incorrect outputs. For example, the<br />
mating and osmosensing pathways in yeast<br />
share several components, including the MAP3K<br />
Ste11p, but each pathway maintains specificity<br />
because it employs different scaffolds that restrict<br />
signal transmission.<br />
In contrast, the presence <strong>of</strong> excess scaffold<br />
can inhibit <strong>signaling</strong> because the individual <strong>signaling</strong><br />
components will more frequently bind<br />
to distinct scaffold proteins rather than forming<br />
a functional complex. Such dilution among scaffolds<br />
causes separation rather than concentration<br />
<strong>of</strong> the components, preventing their<br />
productive interaction.<br />
14.9<br />
Independent, modular<br />
domains specify proteinprotein<br />
interactions<br />
Key concepts<br />
• Protein interactions may be mediated by small,<br />
conserved domains.<br />
• Modular interaction domains are essential for<br />
signal transmission.<br />
• Adaptors consist exclusively <strong>of</strong> binding domains or<br />
motifs.<br />
Modular protein interaction domains or motifs<br />
occur in many <strong>signaling</strong> proteins and confer the<br />
ability to bind structural motifs in other molecules,<br />
including proteins, lipids, and nucleic<br />
acids. Some <strong>of</strong> these domains are listed in FIGURE<br />
14.9. In contrast to scaffolds, which bind specific<br />
proteins with considerable selectivity, modular<br />
interaction domains generally recognize<br />
not a single molecule but a group <strong>of</strong> targets that<br />
share related structural features.<br />
Modular interaction domains important for<br />
signal transduction were first discovered in the<br />
protein tyrosine kinase proto-oncogene Src,<br />
which contains a protein tyrosine kinase domain<br />
and two domains named Src homology<br />
(SH) 2 and 3 domains. The modular SH2 and<br />
SH3 domains were originally identified by comparison<br />
<strong>of</strong> Src to two other tyrosine kinases, Fps<br />
and Abl. One or both <strong>of</strong> these domains appear<br />
in numerous proteins and both are critically involved<br />
in protein-protein interactions.<br />
SH3 domains, which consist <strong>of</strong> approximately<br />
50 residues, bind to specific short proline-rich<br />
sequences. Many cytoskeletal proteins<br />
and proteins found in focal adhesion complexes<br />
contain SH3 domains and proline rich sequences,<br />
suggesting that this targeting motif<br />
may send proteins with these domains to these<br />
sites <strong>of</strong> action within <strong>cell</strong>s. In contrast to phosphotyrosine-SH2<br />
binding, the proline-rich binding<br />
sites for SH3 domains are present in resting<br />
and activated <strong>cell</strong>s. However, SH3-proline interactions<br />
may be negatively regulated by phosphorylation<br />
within the proline-rich motif.<br />
SH2 domains, which consist <strong>of</strong> approximately<br />
100 residues, bind to Tyr phosphorylated<br />
proteins, such as cytoplasmic tyrosine<br />
kinases and receptor tyrosine kinases. Thus, Tyr<br />
phosphorylation regulates the appearance <strong>of</strong><br />
SH2 binding sites and, thereby, regulates a set<br />
<strong>of</strong> protein-protein interactions in a stimulusdependent<br />
manner.<br />
A clever strategy was used to identify the<br />
binding specificity <strong>of</strong> SH2 domains. An isolated<br />
recombinant SH2 domain was incubated with<br />
<strong>cell</strong> lysates and then recovered from the lysates<br />
using a purification tag. The proteins associated<br />
with the SH2 domain were some <strong>of</strong> the same<br />
proteins that were recognized by antiphosphotyrosine<br />
antibodies. By this and other methods,<br />
it was discovered that SH2 domains recognize<br />
sequences surrounding Tyr phosphorylation<br />
sites and require phosphorylation <strong>of</strong> the included<br />
Tyr for high affinity binding.<br />
Information on specific amino acid sequences<br />
that recognize and bind to modular<br />
binding domains is being accumulated as these<br />
individual interactions are identified. In addition,<br />
screening programs using cDNA and/or<br />
peptide libraries to assess binding capabilities<br />
600 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 601<br />
Characteristics <strong>of</strong> some common modular protein domains<br />
Domain Characteristics Cellular involvement<br />
14-3-3 Binds protein phosphoserine or<br />
phosphothreonine<br />
Bromo<br />
CARD<br />
C1<br />
C2<br />
EF hand<br />
F-Box<br />
FHA<br />
HECT<br />
LIM<br />
PDZ<br />
PH<br />
RING<br />
SAM<br />
SH2<br />
Binds acetylated lysine residues<br />
Dimerization<br />
Binds phorbol esters or diacylglycerol<br />
Binds phospholipids<br />
Binds calcium<br />
Binds Skp1 in a ubiquitin-ligase<br />
complex<br />
Binds protein phosphothreonine<br />
or phosphoserine<br />
Binds E2 ubiquitin-conjugating<br />
enzymes to transfer ubiquitin to<br />
the substrate or to ubiquitin<br />
chains<br />
Zinc-binding cysteine-rich motif<br />
that forms two tandemly<br />
repeated zinc fingers<br />
Binds to the C-terminal 4-5<br />
residues <strong>of</strong> proteins that have a<br />
hydrophobic residue at the<br />
terminus; may bind to PIP2<br />
Binds to specific phosphoinositides,<br />
esp. PI-4,5-P 2 , PI-3,4-P 2 or<br />
PI-3,4,5-P 3 .<br />
Binds zinc and may be found in<br />
E3 ubiquitin ligases<br />
Homo- and heterooligomerization<br />
Binds to protein phosphotyrosine<br />
(pY)<br />
Protein sequestration<br />
Chromatin-associated<br />
proteins<br />
Caspase activation<br />
Recruitment to membranes<br />
Signal transduction,<br />
vesicular trafficking<br />
Calcium-dependent<br />
processes<br />
Ubiquitination<br />
Various; DNA damage<br />
FYVE Binds to PI(3)P Membrane trafficking,<br />
TGF- <strong>signaling</strong><br />
Ubiquitination<br />
Wide variety <strong>of</strong><br />
processes<br />
Scaffolding diverse<br />
protein complexes<br />
<strong>of</strong>ten at the membrane<br />
Recruitment to membranes<br />
and motility<br />
Ubiquitination,<br />
transcription<br />
Wide variety <strong>of</strong><br />
processes<br />
Tyrosine protein kinase<br />
<strong>signaling</strong><br />
SH3 Binds to PXXP motifs Various processes<br />
TPR<br />
WW<br />
Degenerate sequence <strong>of</strong> ~34<br />
amino acids with residues<br />
WL/GYAFAP; forms a scaffold<br />
Binds proline-rich sequences<br />
Wide variety <strong>of</strong><br />
processes<br />
Alternative to SH3;<br />
vesicular trafficking<br />
FIGURE 14.9 The table describes a subset <strong>of</strong> known modular protein interaction<br />
domains found in many proteins. Interactions mediated by these<br />
domains are essential to controlling <strong>cell</strong> function. Few if any <strong>of</strong> these domains<br />
exist in prokaryotes. Adapted from the Pawson Lab, Protein Interaction<br />
Domains, Mount Sinai Hospital (http://pawsonlab.mshri.on.ca/).<br />
yield such motifs. Consensus target sequences<br />
for individual domains have been identified<br />
based on the sequence specificity <strong>of</strong> their binding<br />
to arrayed sequences. These consensus sequences<br />
can then be used to predict whether<br />
the domain will bind a site in a candidate protein.<br />
Adaptor proteins, which lack enzymatic<br />
activity, link <strong>signaling</strong> molecules and target<br />
them in a manner that is responsive to extra<strong>cell</strong>ular<br />
signals. Adaptor proteins are generally<br />
made up <strong>of</strong> two or more modular interaction<br />
domains or the complementary recognition<br />
motifs. Unlike scaffolds, adaptors are usually<br />
14.9 Independent, modular domains specify protein-protein interactions 601
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 602<br />
multifunctional because their modular interaction<br />
domains and motifs are not as highly specific.<br />
Adaptors bind to two or more other<br />
<strong>signaling</strong> proteins via their protein-protein interaction<br />
domains to colocalize them or to facilitate<br />
additional interactions.<br />
Grb2 is a prototypical adaptor protein that<br />
was identified as a protein that bound to the C-<br />
terminal region <strong>of</strong> the EGF receptor. Grb2 has<br />
one SH2 and two SH3 domains. It binds constitutively<br />
to specific proline-rich segments <strong>of</strong> proteins<br />
through its SH3 domain, although this<br />
binding can be negatively regulated. One target<br />
<strong>of</strong> Grb2 is SOS, a guanine nucleotide exchange<br />
factor that activates the small GTP-binding protein<br />
Ras in response to EGF <strong>signaling</strong>. Through<br />
its SH2 domain, Grb2 binds Tyr-phosphorylated<br />
proteins, including the receptors themselves in<br />
a stimulus-dependent manner. Thus, Tyr phosphorylation<br />
<strong>of</strong> these receptors in response to<br />
ligand will enable the binding <strong>of</strong> Grb2 to the receptors,<br />
which, in turn, will recruit SOS to the<br />
membrane-localized receptor. Once at the membrane,<br />
SOS can activate its target, Ras.<br />
14.10<br />
Cellular <strong>signaling</strong> is<br />
remarkably adaptive<br />
Key concepts<br />
• Sensitivity <strong>of</strong> <strong>signaling</strong> pathways is regulated to<br />
allow responses to change over a wide range <strong>of</strong><br />
signal strengths.<br />
• Feedback mechanisms execute this function in all<br />
<strong>signaling</strong> pathways.<br />
• Most pathways contain multiple adaptive feedback<br />
loops to cope with signals <strong>of</strong> various strengths and<br />
durations.<br />
A universal property <strong>of</strong> <strong>cell</strong>ular <strong>signaling</strong> pathways<br />
is adaptation to the incoming signal. Cells continuously<br />
adjust their sensitivity to signals to maintain<br />
their ability to detect changes in input. Typically,<br />
when a <strong>cell</strong> is exposed to a new input, it initiates<br />
a process <strong>of</strong> desensitization that dampens the <strong>cell</strong>ular<br />
response to a new plateau lower than the initial<br />
peak response, as illustrated in FIGURE 14.10.<br />
When the stimulus is removed, the desensitized<br />
state can persist, with sensitivity slowly returning<br />
to normal. Similarly, the removal <strong>of</strong> a tonic stimulus<br />
can hypersensitize <strong>signaling</strong> systems.<br />
FIGURE 14.10 Top: Upon exposure to<br />
a stimulus, <strong>signaling</strong> pathways adjust<br />
their sensitivities to adapt to the new<br />
level <strong>of</strong> input. Thus, the response decays<br />
after initial stimulation. A second<br />
similar stimulus will elicit a smaller<br />
response unless adequate time is allowed<br />
for recovery. Bottom: Some adaptation<br />
mechanisms feed back only on<br />
the receptor that is stimulated and do<br />
not alter parallel pathways. Such mechanisms<br />
are referred to as homologous.<br />
At left, agonist a for receptor R1 can<br />
initiate either <strong>of</strong> two feedback events<br />
that desensitize R1 alone. In other<br />
cases, a stimulus will also cause parallel<br />
or related systems to desensitize.<br />
At the right, agonist a initiates desensitization<br />
<strong>of</strong> both R1 and R2. The response<br />
to agonist b, which binds to<br />
R2, is also desensitized. Such heterologous<br />
desensitization is common.<br />
K<br />
a<br />
Homologous<br />
desensitization<br />
Initial<br />
response<br />
R1 R 2<br />
R1 R 2<br />
X 1 X 2<br />
R esponse<br />
Response<br />
Patterns <strong>of</strong> adaptation in <strong>signaling</strong> networks<br />
Agonist a<br />
for R1<br />
Agonist Agonist Agonist<br />
Reapply<br />
a or b<br />
Desensitization<br />
b<br />
a<br />
a<br />
R1 R 2<br />
X 1 X 2<br />
Time<br />
Heterologous<br />
desensitization<br />
Response<br />
R1<br />
Agonist a<br />
for R1<br />
or R1 R 2<br />
Reapply<br />
a or b<br />
Y<br />
Time<br />
Y<br />
Time<br />
Z<br />
Z<br />
602 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 603<br />
Adaptation in <strong>signaling</strong> is one <strong>of</strong> the best examples<br />
<strong>of</strong> biological homeostasis. The adaptability<br />
<strong>of</strong> <strong>cell</strong>ular <strong>signaling</strong> can be quite impressive.<br />
Cells commonly regulate their sensitivity to physiological<br />
stimuli over more than a 100-fold range,<br />
and the mammalian visual response can adapt to<br />
incoming light over a 107-fold range. This remarkable<br />
ability allows a photoreceptor <strong>cell</strong> to<br />
detect a single photon, and allows a person to<br />
read in both very dim light and intense sunlight.<br />
Adaptability is observed in bacteria, plants, fungi,<br />
and animals. Many <strong>of</strong> its properties are conserved<br />
throughout biology, although the most complex<br />
adaptive mechanisms are found in animals. The<br />
general mechanism for adaptation is the negative<br />
feedback loop, which biochemically samples<br />
the signal and controls the adaptive process.<br />
Adaptation varies with both the intensity and<br />
the duration <strong>of</strong> the incoming signal. Stronger or<br />
more persistent inputs tend to drive greater adaptive<br />
change and, <strong>of</strong>ten, adaptation that persists<br />
for a longer time. Cells can modulate adaptation<br />
in this way because adaptation is exerted by a<br />
succession <strong>of</strong> independent mechanisms, each with<br />
its own sensitivity and kinetic parameters.<br />
G protein pathways <strong>of</strong>fer ex<strong>cell</strong>ent examples<br />
<strong>of</strong> adaptation. FIGURE 14.11 shows that the earliest<br />
step in adaptation is receptor phosphorylation,<br />
which is catalyzed by G protein-coupled<br />
receptor kinases (GRKs) that selectively recognize<br />
the receptor’s ligand-activated conformation.<br />
Phosphorylation inhibits the receptor’s ability<br />
to stimulate G protein activation and also promotes<br />
binding <strong>of</strong> arrestin, a protein that further<br />
inhibits G protein activation. Moreover, arrestin<br />
binding primes receptors for endocytosis, which<br />
removes them from the <strong>cell</strong> surface. Endocytosis<br />
can also be the first step in receptor proteolysis.<br />
Along with these direct effects, many receptor<br />
genes display feedback inhibition <strong>of</strong> transcription,<br />
such that <strong>signaling</strong> by a receptor decreases<br />
its own expression.<br />
Stimulation thus causes multiple adaptive<br />
processes that range from immediate (phosphorylation,<br />
arrestin binding) through delayed (transcriptional<br />
regulation), and include both reversible<br />
and irreversible events. This array <strong>of</strong> adaptive<br />
events has been demonstrated for many G protein-coupled<br />
receptors, and many <strong>cell</strong>s may use<br />
all <strong>of</strong> them to control output from one receptor.<br />
The speed, extent, and reversibility <strong>of</strong> adaptation<br />
are selected by a <strong>cell</strong>’s developmental program.<br />
Cells can change their patterns <strong>of</strong> adaptation<br />
both qualitatively and quantitatively by altering<br />
the points in a pathway where feedback is initiated<br />
and exerted. In a linear pathway, changing<br />
Multiple adaptation processes occur after a stimulus<br />
Relative<br />
response<br />
Receptor phosphorylation<br />
Arrestin binding<br />
Receptor<br />
endocytosis<br />
Endosomal receptor<br />
degradation<br />
5 Receptor transcription<br />
inhibited<br />
0 1 10 100 1000<br />
Time (seconds)<br />
Agonist<br />
added<br />
Agonist<br />
binds<br />
1<br />
2<br />
Agonist<br />
GPCR<br />
G protein<br />
Receptor<br />
recycling<br />
Lysosome<br />
3<br />
G protein<br />
active<br />
4<br />
EFFECTORS<br />
CYTOPLASM<br />
4 Receptor<br />
degradation<br />
DNA<br />
GRK<br />
1 Receptor<br />
Arrestin<br />
phosphorylation<br />
2 Arrestin<br />
binding<br />
NUCLEUS<br />
Early<br />
endosome<br />
GPCR gene<br />
3 Receptor<br />
endocytosis<br />
5 Receptor<br />
transcription<br />
inhibited<br />
FIGURE 14.11 Multiple adaptation processes are invoked during a stimulus,<br />
and multiple nested mechanisms for adaptation are the rule. They are usually<br />
invoked sequentially according to the duration and intensity <strong>of</strong> the stimulus.<br />
For GPCRs, at least five desensitizing mechanisms are known, with others acting<br />
on the G protein and effectors.<br />
these points will alter the kinetics or extent <strong>of</strong><br />
adaptation (Figure 14.10). In branched pathways,<br />
changing these points can determine whether<br />
adaptation is unique to one input or is exerted<br />
for many similar inputs. If receptor activation triggers<br />
its desensitization directly, or if an event<br />
downstream on an unbranched pathway triggers<br />
desensitization, then only signals that initiate with<br />
that receptor will be altered. Receptor-selective<br />
adaptation is referred to as homologous adaptation<br />
(Figure 14.10).<br />
14.10 Cellular <strong>signaling</strong> is remarkably adaptive 603
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 604<br />
Alternatively, feedback control can initiate<br />
downstream from multiple receptors in a convergent<br />
pathway and thus regulate both the<br />
initiating receptor and the others. Such heterologous<br />
adaptation regulates all the possible<br />
inputs to a given control point. A common<br />
example is the phosphorylation <strong>of</strong> G proteincoupled<br />
receptors by either protein kinase A or<br />
protein kinase C, which are activated by downstream<br />
signals cAMP or Ca2+ plus the lipid diacylglycerol,<br />
respectively. Like GRK, these kinases<br />
both attenuate receptor activity and promote<br />
arrestin binding.<br />
Cells also alter their responses to incoming<br />
signals for homeostatic reasons. These considerations<br />
include phase <strong>of</strong> the <strong>cell</strong> cycle, metabolic<br />
status, or other aspects <strong>of</strong> <strong>cell</strong>ular activity.<br />
Again, all these adaptive processes may be displayed<br />
to a greater or lesser extent in different<br />
<strong>cell</strong>s, different pathways within a <strong>cell</strong> or different<br />
situations during the <strong>cell</strong>’s lifetime.<br />
14.11<br />
Signaling proteins are<br />
frequently expressed as<br />
multiple species<br />
Key concepts<br />
• Distinct species (is<strong>of</strong>orms) <strong>of</strong> similar <strong>signaling</strong><br />
proteins expand the regulatory mechanisms<br />
possible in <strong>signaling</strong> pathways.<br />
• Is<strong>of</strong>orms may differ in function, susceptibility to<br />
regulation or expression.<br />
• Cells may express one or several is<strong>of</strong>orms to fulfill<br />
their <strong>signaling</strong> needs.<br />
Cells increase the richness, adaptability, and<br />
regulation <strong>of</strong> their <strong>signaling</strong> pathways by expressing<br />
multiple species <strong>of</strong> individual <strong>signaling</strong><br />
proteins that display distinct biochemical<br />
properties. These species may be encoded by<br />
multiple genes or by multiple mRNAs derived<br />
from a single gene by alternative splicing or<br />
mRNA editing. The numerical complexity implicit<br />
in these choices is impressive. Consider<br />
the neurotransmitter serotonin: In mammals,<br />
there are thirteen serotonin receptors, each <strong>of</strong><br />
which stimulates a distinct spectrum <strong>of</strong> G proteins<br />
<strong>of</strong> the G i<br />
, G s<br />
, and G q<br />
families. (A fourteenth<br />
serotonin receptor is an ion channel.)<br />
FIGURE 14.12 shows the relationship <strong>of</strong> serotonin<br />
receptors to these G protein families.<br />
There is also tremendous diversity among<br />
the G proteins and adenylyl cyclases. There are<br />
three genes for Gα i<br />
and one each for the closely<br />
related Gα z<br />
and Gα o<br />
. Furthermore, the Gα o<br />
mRNA is multiply spliced. There are four G q<br />
members. In addition, there are five genes for<br />
Gβ and twelve for Gγ, and most <strong>of</strong> the possible<br />
Gβγ dimers are expressed naturally. There are<br />
ten genes for adenylyl cyclases, which are direct<br />
targets <strong>of</strong> G s<br />
and either direct or indirect targets<br />
<strong>of</strong> the other G proteins. While all nine membrane-bound<br />
adenylyl cyclase is<strong>of</strong>orms are stimulated<br />
by Gα s<br />
, they display diverse stimulatory<br />
and inhibitory responses to Gβγ, Gα i<br />
, Ca2+,<br />
calmodulin, and several protein kinases, as illustrated<br />
in FIGURE 14.13. Thus, stimulation by<br />
serotonin can lead to diverse responses depending<br />
upon the various forms <strong>of</strong> the proteins that<br />
are engaged at a particular time and location.<br />
FIGURE 14.12 Receptors for serotonin have<br />
evolved in mammals as a family <strong>of</strong> 13 genes that<br />
regulate three <strong>of</strong> the four major classes <strong>of</strong> G proteins.<br />
While all respond to the natural ligand<br />
serotonin, the binding sites have evolved sufficient<br />
differences that drugs have been developed<br />
that specifically target one or more<br />
is<strong>of</strong>orms. The type 3 serotonin receptors, not<br />
shown here, are ligand-gated ion channels and<br />
are not obviously related to the others.<br />
Evolutionary relationship <strong>of</strong> serotonin receptor is<strong>of</strong>orms<br />
Is<strong>of</strong>orms G protein<br />
1B<br />
1D<br />
1E G i<br />
1F<br />
1A<br />
5A<br />
7<br />
5B<br />
G s<br />
4<br />
2A<br />
2C<br />
G q<br />
6<br />
2B<br />
G s<br />
120 100 80 60 40 20 0<br />
Nucleotide substitution distance<br />
604 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 605<br />
Different is<strong>of</strong>orms <strong>of</strong> adenylyl cyclase are regulated differently<br />
Gα s<br />
Gβγ<br />
PKC<br />
Ca 2+ NO<br />
PKA<br />
Regulators<br />
CaMK<br />
inhibit<br />
Gα i<br />
activate<br />
CaM<br />
FIGURE 14.13 All <strong>of</strong> the mammalian membrane-bound adenylyl cyclases are<br />
structurally homologous and catalyze the same reaction, and all are stimulated<br />
by G s<br />
. Their responses to other inputs (protein kinases CaMK, PKA and PKC;<br />
Ca2+; calmodulin (CaM); NO • ) are specific to each is<strong>of</strong>orm, allowing a rich combinatoric<br />
input to <strong>cell</strong>ular cAMP <strong>signaling</strong>.<br />
Sometimes is<strong>of</strong>orms <strong>of</strong> a <strong>signaling</strong> protein<br />
are subject to quite different kinds <strong>of</strong> inputs.<br />
For example, all <strong>of</strong> the members <strong>of</strong> the phospholipase<br />
C family (PLC) hydrolyze phosphatidylinositol-4,5-bisphosphate<br />
to form two second<br />
messengers, diacylglycerol and inositol-1,4,5<br />
trisphosphate (see 14.16 Lipids and lipid-derived<br />
compounds are <strong>signaling</strong> molecules). The distinct<br />
is<strong>of</strong>orms may be regulated by diverse combinations<br />
<strong>of</strong> Gα q<br />
, Gβγ, phosphorylation, monomeric<br />
G proteins, or Ca2+.<br />
Because a <strong>cell</strong> has multiple options when<br />
expressing a form <strong>of</strong> a <strong>signaling</strong> protein, it can<br />
use expression <strong>of</strong> particular is<strong>of</strong>orms to alter<br />
how it performs otherwise identical <strong>signaling</strong><br />
functions. Different <strong>cell</strong>s express one or more<br />
is<strong>of</strong>orms to allow appropriate responses, and expression<br />
can vary according to other inputs or<br />
the <strong>cell</strong>’s metabolic status. In addition, <strong>signaling</strong><br />
pathways are remarkably resistant to mutational<br />
or other injuries because loss <strong>of</strong> a single species<br />
or is<strong>of</strong>orm <strong>of</strong> a <strong>signaling</strong> protein can <strong>of</strong>ten be<br />
compensated for by increased expression or activity<br />
<strong>of</strong> another species. Similarly, engineered<br />
overexpression can result in the reduced expression<br />
<strong>of</strong> endogenous proteins. The existence <strong>of</strong><br />
multiple receptor species can, thus, substantially<br />
add to adaptability and the consequent resistance<br />
<strong>of</strong> <strong>signaling</strong> networks to damage.<br />
14.12<br />
Activating and<br />
deactivating reactions<br />
are separate and<br />
independently controlled<br />
Key concepts<br />
• Activating and deactivating reactions are usually<br />
executed by different regulatory proteins.<br />
• Separating activation and inactivation allows for<br />
fine-tuned regulation <strong>of</strong> amplitude and timing.<br />
In <strong>signaling</strong> networks, individual proteins are<br />
frequently activated and deactivated by distinct<br />
reactions, a feature that facilitates separate regulation.<br />
Common examples include using protein<br />
kinases and phosphoprotein phosphatases<br />
14.12 Activating and deactivating reactions are separate and independently controlled 605
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 606<br />
to catalyze protein phosphorylation and dephosphorylation;<br />
using adenylyl cyclase to create<br />
cAMP while using phosphodiesterases to<br />
hydrolyze it or anion transporters to pump it<br />
out <strong>of</strong> the <strong>cell</strong>; or using GTP/GDP exchange factors<br />
(GEFs) to activate G proteins and GTPaseactivating<br />
proteins (GAPs) to deactivate them.<br />
Depending on stoichiometry and detailed mechanism,<br />
these strategies can convey either additive<br />
or nonadditive inputs while maintaining<br />
fine control over the kinetics <strong>of</strong> activation and<br />
deactivation <strong>of</strong> a <strong>signaling</strong> pathway. The use <strong>of</strong><br />
distinct reactions for activation and deactivation<br />
is analogous to the use <strong>of</strong> distinct anabolic<br />
and catabolic enzymes in reversible metabolic<br />
pathways.<br />
14.13<br />
Cellular <strong>signaling</strong> uses<br />
both allostery and<br />
covalent modification<br />
Key concepts<br />
• Allostery refers to the ability <strong>of</strong> a molecule to alter<br />
the conformation <strong>of</strong> a target protein when it binds<br />
noncovalently to that protein.<br />
• Modification <strong>of</strong> a protein’s chemical structure is<br />
also frequently used to regulate its activity.<br />
Cellular <strong>signaling</strong> uses almost every imaginable<br />
mechanism for regulating the activities <strong>of</strong><br />
intra<strong>cell</strong>ular proteins, but most can be described<br />
as either allosteric or covalent. Individual <strong>signaling</strong><br />
proteins typically respond to multiple allosteric<br />
and covalent inputs.<br />
Allostery refers to the ability <strong>of</strong> a molecule<br />
to alter the conformation <strong>of</strong> a target protein<br />
when it binds noncovalently to that protein.<br />
Because a protein’s activity reflects its conformation,<br />
the binding <strong>of</strong> any molecule that alters<br />
conformation can change the target protein’s<br />
activity. Any molecule can have allosteric effects:<br />
protons or Ca2+, small organic molecules,<br />
or other proteins. Allosteric regulation can be<br />
both inhibitory or stimulatory.<br />
Covalent modification <strong>of</strong> a protein’s chemical<br />
structure is also frequently used to regulate<br />
its activity. The change in the protein’s chemical<br />
structure alters its conformation and, thus,<br />
its activity. Most regulatory covalent modification<br />
is reversible. The classic and most common<br />
regulatory covalent event is phosphorylation,<br />
in which a phosphoryl group is transferred from<br />
ATP to the protein, most <strong>of</strong>ten to the hydroxyl<br />
group <strong>of</strong> serine (Ser), threonine(Thr), or tyrosine<br />
(Tyr). Enzymes that phosphorylate proteins<br />
are known as protein kinases. Their actions are<br />
opposed by phosphoprotein phosphatases, which<br />
catalyze the hydrolysis <strong>of</strong> the phosphoryl group<br />
to yield free phosphate and restore the unmodified<br />
hydroxyl residue. Other forms <strong>of</strong> covalent<br />
modification are also common and will be addressed<br />
throughout the chapter.<br />
14.14<br />
Second messengers<br />
provide readily diffusible<br />
pathways for information<br />
transfer<br />
Key concepts<br />
• Second messengers can propagate signals between<br />
proteins that are at a distance.<br />
• cAMP and Ca2+ are widely used second messengers.<br />
Signaling pathways make use <strong>of</strong> both proteins<br />
and small molecules according to their distinctive<br />
attributes. A small molecule used as an intra<strong>cell</strong>ular<br />
signal, or second messenger, has a<br />
number <strong>of</strong> advantages over a protein as a <strong>signaling</strong><br />
intermediary. Small molecules can be<br />
synthesized and destroyed quickly. Because they<br />
can be made readily, they can act at high concentrations<br />
so that their affinities for target proteins<br />
can be low. Low affinity permits rapid<br />
dissociation, such that their signals can be terminated<br />
promptly when free second messenger<br />
molecules are destroyed or sequestered. Because<br />
second messengers are small, they also can diffuse<br />
quickly within the <strong>cell</strong>, although many <strong>cell</strong>s<br />
have developed mechanisms to spatially restrict<br />
such diffusion. Second messengers are, thus,<br />
superior to proteins in mediating fast responses,<br />
particularly at a distance. Second messengers<br />
are also useful when signals have to be addressed<br />
to large numbers <strong>of</strong> target proteins simultaneously.<br />
These advantages <strong>of</strong>ten overcome their<br />
lack <strong>of</strong> catalytic activity and their inability to<br />
bind multiple molecules simultaneously.<br />
FIGURE 14.14 lists intra<strong>cell</strong>ular second messengers<br />
developed through evolution. This number<br />
is surprisingly low. Several are nucleotides<br />
synthesized from major metabolic nucleotide<br />
precursors. They include cAMP, cyclic GMP,<br />
ppGppp, and cyclic ADP-ribose. Other soluble<br />
second messengers include a sugar phosphate,<br />
inositol-1,4,5-trisphosphate (IP 3<br />
), a divalent metal<br />
ion Ca2+, and a free radical gas nitric oxide (NO • ).<br />
Lipid second messengers include diacylglycerol<br />
and phosphatidylinositol-3,4,5-trisphosphate,<br />
606 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 607<br />
Second<br />
messenger<br />
3':5'-cyclic AMP<br />
(cAMP)<br />
RNA polymerase<br />
Magic spot<br />
(ppGpp, ppGppp) ObgE transcription<br />
arrest<br />
detector<br />
Cyclic di-GMP<br />
phosphodiesterase<br />
Inositol-1,3,5-<br />
trisphosphate<br />
(IP 3 )<br />
Diacylglycerol<br />
(DAG)<br />
Phosphatidyl-<br />
inositol-4,5-<br />
bisphosphate<br />
(PIP 2 )<br />
3':5'-Cyclic GMP<br />
(cGMP)<br />
Cyclic ADP-ribose<br />
Nitric oxide (NO. )<br />
Ca2+<br />
Cyclic<br />
diguanosinemonophosphate<br />
Phosphatidyl-<br />
inositol-3,4,5-<br />
trisphosphate<br />
Adenylyl<br />
Protein kinase A<br />
cyclase<br />
Bacterial transcription<br />
factors<br />
Cation channel<br />
Cyclic nucleotide<br />
phosphodiesterase<br />
Rap GDP/GTP<br />
exchange factor<br />
(Epac)<br />
IP 3 -gated Ca 2+<br />
channel<br />
Protein<br />
kinase C<br />
Trp cation<br />
channel<br />
Ion channel<br />
Transporters<br />
Protein kinase G<br />
Ca2+ channel<br />
Various two<br />
component<br />
system proteins<br />
Guanylyl cyclase<br />
Numerous<br />
calmodulin<br />
Akt (protein<br />
kinase B)<br />
Second messengers<br />
Targets<br />
Other PH<br />
domains/proteins<br />
Synthesis/<br />
Release<br />
Rel1A<br />
SpoT<br />
Cation channel<br />
Cyclic nucleotide<br />
phosphodiesterase<br />
Phospholipase<br />
C<br />
Phospholipase<br />
C<br />
PIP 5-kinase<br />
Guanylyl<br />
cyclase<br />
ADP-ribose<br />
cyclase<br />
Diguanylate<br />
cyclase<br />
PI 3-kinase<br />
GTP<br />
PIP 2<br />
PIP 2<br />
PI-4-P<br />
GTP<br />
NAD<br />
GTP<br />
Stored<br />
Ca2+<br />
PIP 2<br />
phosphatidylinositol-4,5-diphosphate, sphingosine-1-phosphate<br />
and phosphatidic acid.<br />
The first <strong>signaling</strong> compound to be described<br />
as a second messenger was cAMP. The name<br />
arose because cAMP is synthesized in animal<br />
<strong>cell</strong>s as a second, intra<strong>cell</strong>ular signal in response<br />
to numerous extra<strong>cell</strong>ular hormones, the first<br />
messengers in the pathway. cAMP is used by<br />
prokaryotes, fungi, and animals to convey information<br />
to a variety <strong>of</strong> regulatory proteins.<br />
(Its occurrence in higher plants has still not been<br />
proved.)<br />
Adenylyl cyclases, the enzymes that synthesize<br />
cAMP from ATP, are regulated in various<br />
ways depending on the organism in which<br />
they occur. In animals, adenylyl cyclase is an<br />
integral protein <strong>of</strong> the plasma membrane whose<br />
multiple is<strong>of</strong>orms are stimulated by diverse<br />
agents (see Figure 14.13). In animal <strong>cell</strong>s, adenylyl<br />
cyclase is generally stimulated by G s<br />
, which<br />
was originally discovered as an adenylyl cyclase<br />
regulator. Some fungal adenylyl cyclases are<br />
also stimulated by G proteins. Bacterial cyclases<br />
are far more diverse in their regulation.<br />
cAMP is removed from <strong>cell</strong>s in two ways.<br />
It may be extruded from <strong>cell</strong>s by an ATP-driven<br />
anion pump but is more <strong>of</strong>ten hydrolyzed to 5′-<br />
AMP by members <strong>of</strong> the cyclic nucleotide phosphodiesterase<br />
family, a large group <strong>of</strong> proteins<br />
that are themselves under multiple regulatory<br />
controls.<br />
The prototypical downstream regulator for<br />
cAMP in animals is the cAMP-dependent protein<br />
kinase, but a bacterial cAMP-regulated transcription<br />
factor was discovered shortly thereafter,<br />
and other effectors are now known (Figure<br />
14.14). The cAMP system remains the prototypical<br />
eukaryotic <strong>signaling</strong> pathway in that its<br />
components exemplify almost all <strong>of</strong> the recognized<br />
varieties <strong>of</strong> <strong>signaling</strong> molecules and their<br />
interactions: hormone, receptor, G protein,<br />
adenylyl cyclase, protein kinase, phosphodiesterase,<br />
and extrusion pump.<br />
The second messenger-stimulated protein<br />
kinase PKA is a tetramer composed <strong>of</strong> two catalytic<br />
(C) subunits and two regulatory (R) subunits,<br />
as illustrated in FIGURE 14.15. The R subunit<br />
binds to the catalytic subunit in the substratebinding<br />
region, maintaining C in an inhibited<br />
state. Each R subunit binds two molecules <strong>of</strong><br />
cAMP, four cAMP molecules per PKA holoenzyme.<br />
When these sites are filled, the R subunit<br />
dimer dissociates rapidly, leaving two free catalytic<br />
subunits with high activity. The difference<br />
in affinity <strong>of</strong> R for C in the presence and absence<br />
<strong>of</strong> cAMP is ~10,000-fold. The strongly cooperative<br />
binding <strong>of</strong> cAMP generates a very steep<br />
activation curve with an apparent threshold below<br />
which no significant activation <strong>of</strong> PKA occurs,<br />
as illustrated in Figure 14.15. PKA activity,<br />
thus, increases dramatically over a narrow range<br />
<strong>of</strong> cAMP concentrations. PKA is also regulated<br />
Precursor<br />
Removal<br />
Phosphodiesterase<br />
ATP<br />
Organic<br />
anion<br />
transporter<br />
SpoTcatalyzed<br />
hydrolysis<br />
Phosphatase<br />
Diacylglycerol<br />
kinase<br />
Diacylglycerol<br />
lipase<br />
Phospholipase<br />
C<br />
Phosphatase<br />
Phosphodiesterase<br />
Hydrolysis<br />
NO . synthase arginine Reduction<br />
Release from<br />
storage<br />
organelles<br />
or plasma<br />
membrane<br />
channels<br />
Reuptake<br />
and<br />
extrusion<br />
pumps<br />
Phosphatase<br />
FIGURE 14.14 Major second messengers, some <strong>of</strong> the proteins that they regulate,<br />
their sources and their disposition.<br />
14.14 Second messengers provide readily diffusible pathways for information transfer 607
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 608<br />
(C)<br />
Catalytic<br />
subunits<br />
PKA<br />
C<br />
R<br />
R<br />
C<br />
Activation <strong>of</strong> PKA by cAMP<br />
(R)<br />
R egulatory<br />
subunits<br />
4 cAMP<br />
R<br />
R<br />
- cAMP<br />
- cAMP<br />
- cAMP<br />
- cAMP<br />
Activated<br />
PKA<br />
C<br />
C<br />
teins throughout the <strong>cell</strong> ranging from ion channels<br />
to transcription factors, and its conserved<br />
substrate preference frequently permits prediction<br />
<strong>of</strong> substrates by sequence analysis. The<br />
cAMP response element binding protein CREB<br />
is phosphorylated by PKA on Ser 133 and is<br />
largely responsible for the impact <strong>of</strong> cAMP on<br />
transcription <strong>of</strong> numerous genes.<br />
R 2 C 2 + 4 cAMP<br />
Kinase activity as a<br />
function <strong>of</strong> [cAMP] (%)<br />
100<br />
80<br />
60<br />
40<br />
20<br />
10%<br />
R 2<br />
. cAMP 4 + 2C<br />
90%<br />
14.15<br />
Ca2+ <strong>signaling</strong> serves<br />
diverse purposes in all<br />
eukaryotic <strong>cell</strong>s<br />
Key concepts<br />
• Ca2+ serves as a second messenger and regulatory<br />
molecule in essentially all <strong>cell</strong>s.<br />
• Ca2+ acts directly on many target proteins and also<br />
regulates the activity <strong>of</strong> a regulatory protein<br />
calmodulin.<br />
• The cytosolic concentration <strong>of</strong> Ca2+ is controlled by<br />
organellar sequestration and release.<br />
2 x 10 -9 2 x 10 -8 2 x 10 -7<br />
[cAMP]<br />
FIGURE 14.15 PKA is a heterotetramer composed <strong>of</strong> two catalytic (C) and<br />
two regulatory (R) subunits. Binding <strong>of</strong> four molecules <strong>of</strong> cAMP to the regulatory<br />
subunits induces dissociation <strong>of</strong> two molecules <strong>of</strong> C, the active form<br />
<strong>of</strong> PKA, from the cAMP-bound regulatory subunit dimer. In the bottom panel,<br />
the cooperative binding <strong>of</strong> four molecules <strong>of</strong> cAMP generates a steep activation<br />
pr<strong>of</strong>ile. Activity increases from approximately 10% to 90% as the<br />
cAMP concentration increases only 10-fold. An apparent threshold is introduced<br />
because there is little change in activity at low concentrations <strong>of</strong><br />
cAMP.<br />
by phosphorylation <strong>of</strong> its activation loop.<br />
Phosphorylation occurs cotranslationally, and<br />
the activation loop phosphorylation is required<br />
for assembly <strong>of</strong> the R 2<br />
C 2<br />
tetramer.<br />
The PKAs are mostly cytosolic and are also<br />
targeted to specific locations by binding organelle-associated<br />
scaffolds (A-kinase anchoring<br />
proteins, or AKAPs). These AKAPs facilitate<br />
phosphorylation <strong>of</strong> membrane proteins including<br />
GPCRs, transporters, and ion channels.<br />
AKAPs can also target PKA to other <strong>cell</strong>ular locations<br />
including mitochondria, the cytoskeleton,<br />
and the centrosome. AKAPs <strong>of</strong>ten harbor<br />
binding sites for other regulatory molecules<br />
such as phosphoprotein phosphatases and additional<br />
protein kinases, which allows for coordination<br />
<strong>of</strong> multiple <strong>signaling</strong> pathways and<br />
integration <strong>of</strong> their outputs.<br />
PKA generally phosphorylates substrates<br />
with a primary consensus motif <strong>of</strong> Arg-Arg-<br />
Xaa-Ser-Hydrophobic, placing it in a large group<br />
<strong>of</strong> kinases that recognize basic residues preceding<br />
the phosphorylation site. PKA regulates pro-<br />
Ca2+ is used as a second messenger in all <strong>cell</strong>s,<br />
and is, thus, an even more widespread second<br />
messenger than cAMP. Many proteins bind Ca2+<br />
with consequent allosteric changes in their enzymatic<br />
activities, sub<strong>cell</strong>ular localization, or<br />
interaction with other proteins or with lipids.<br />
Direct targets <strong>of</strong> Ca2+ regulation include almost<br />
all classes <strong>of</strong> <strong>signaling</strong> proteins described in this<br />
chapter, numerous metabolic enzymes, ion<br />
channels and pumps, and contractile proteins.<br />
Most noteworthy may be muscle actomyosin<br />
fibers, which are triggered to contract in response<br />
to cytosolic Ca2+ (see 8.21 Myosin-II functions<br />
in muscle contraction).<br />
Although free Ca2+ is found at concentrations<br />
near 1 mM in most extra<strong>cell</strong>ular fluids, intra<strong>cell</strong>ular<br />
Ca2+ concentrations are maintained<br />
near 100 nanomolar levels by the combined action<br />
<strong>of</strong> pumps and transporters that either extrude<br />
free Ca2+ or sequester it in the endoplasmic<br />
reticulum or mitochondria. Ca2+ <strong>signaling</strong> is initiated<br />
when Ca2+-selective channels in the endoplasmic<br />
reticulum or plasma membrane are<br />
opened to allow Ca2+ to enter the cytoplasm.<br />
The most important entrance channels include<br />
electrically gated channels in animal plasma<br />
membranes; a Ca2+ channel in the endoplasmic<br />
reticulum that is opened by another second messenger,<br />
inositol 1,4,5-trisphosphate (see below);<br />
and an electrically gated channel in the endoplasmic<br />
(sarcoplasmic) reticulum <strong>of</strong> muscle<br />
that opens in response to depolarization <strong>of</strong> nearby<br />
plasma membrane, a process known as excitation-contraction<br />
coupling (see 2.9 Plasma mem-<br />
608 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 609<br />
Calcium binding causes a conformational change in calmodulin<br />
Calcium-free<br />
calmodulin<br />
calmodulin free + 4 Ca 2+<br />
Calcium-bound<br />
calmodulin bound to<br />
target peptide <strong>of</strong> CaMK<br />
Ca 2+<br />
target<br />
(Ca 2+ ) 4<br />
. calmodulin . active target<br />
FIGURE 14.16 Ribbon diagrams representing<br />
the crystal structures <strong>of</strong> calmodulin free<br />
<strong>of</strong> Ca2+ and bound to four Ca2+ ions reveal<br />
the huge conformational change that<br />
calmodulin undergoes upon Ca2+ binding.<br />
Ca2+-calmodulin causes activity changes in<br />
target proteins. The bottom panel shows<br />
the activation <strong>of</strong> a target by calmodulin as<br />
a function <strong>of</strong> the intra<strong>cell</strong>ular free Ca2+ concentration.<br />
The requirement for binding<br />
four Ca2+ ions to induce the conformational<br />
transition results in cooperative activation<br />
<strong>of</strong> targets. Activity increases from 10% to<br />
90% as the Ca2+ concentration increases<br />
only 10-fold. Structures generated from<br />
Protein Data Bank files 1CFD and 1MXE.<br />
Activation <strong>of</strong> target<br />
by calmodulin (%)<br />
100<br />
90%<br />
80<br />
60<br />
40<br />
20<br />
10%<br />
3 x 10 -8 3 x 10 -7 3 x 10 -6<br />
[Ca 2+ ]<br />
brane Ca2+ channels activate intra<strong>cell</strong>ular functions).<br />
In addition to the proteins that are regulated<br />
by binding Ca2+ directly, many other proteins respond<br />
to Ca2+ by binding a widespread Ca2+ sensor,<br />
the small, ~17 kDa protein calmodulin.<br />
Calmodulin requires the binding <strong>of</strong> four molecules<br />
<strong>of</strong> Ca2+ to become fully active, and binding<br />
is highly cooperative, generating a sigmoid<br />
activation pr<strong>of</strong>ile illustrated in FIGURE 14.16.<br />
Calmodulin generally binds its targets in a Ca2+dependent<br />
manner, but Ca2+-free calmodulin<br />
may remain bound but inactive in some cases.<br />
For example, calmodulin is a constitutive subunit<br />
<strong>of</strong> phosphorylase kinase that is activated<br />
upon Ca2+ binding. Higher plants again make<br />
major modifications to this paradigm. Calmodulin<br />
is not expressed as a distinct protein but, instead,<br />
is found as a domain in Ca2+-regulated proteins.<br />
In yet another variation, the adenylyl cyclase secreted<br />
by the pathogenic bacterium Bordetella pertussis<br />
is inactive outside <strong>cell</strong>s but is activated by<br />
Ca2+-free calmodulin in animal <strong>cell</strong>s, where its<br />
rapid production <strong>of</strong> cAMP is highly toxic.<br />
14.16<br />
Lipids and lipid-derived<br />
compounds are <strong>signaling</strong><br />
molecules<br />
Key concepts<br />
• Multiple lipid-derived second messengers are<br />
produced in membranes.<br />
• Phospholipase Cs release soluble and lipid second<br />
messengers in response to diverse inputs.<br />
• Channels and transporters are modulated by<br />
different lipids in addition to inputs from other<br />
sources.<br />
• PI 3-kinase synthesizes PIP 3<br />
to modulate <strong>cell</strong><br />
shape and motility.<br />
• PLD and PLA 2<br />
create other lipid second<br />
messengers.<br />
Signals that originate at the plasma membrane<br />
may have soluble regulatory targets in the cytoplasm<br />
or intra<strong>cell</strong>ular organelles, but integral<br />
plasma membrane proteins are also subject to<br />
acute controls. For these targets, lipid second<br />
messengers may be primary inputs. Lipids derived<br />
from membrane phospholipids or other<br />
14.16 Lipids and lipid-derived compounds are <strong>signaling</strong> molecules 609
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 610<br />
lipid species play numerous roles in <strong>cell</strong> <strong>signaling</strong>.<br />
Because their analysis has been more difficult<br />
than for soluble messengers, many<br />
probably remain to be discovered and understood.<br />
FIGURE 14.17 shows the structure <strong>of</strong> some<br />
<strong>of</strong> these lipids.<br />
Phospholipase Cs (PLCs) are the prototypical<br />
lipid <strong>signaling</strong> enzymes. PLC is<strong>of</strong>orms catalyze<br />
the hydrolysis <strong>of</strong> phospholipids between<br />
the 3-sn-hydroxyl and the phosphate group to<br />
yield a diacylglycerol and phosphate ester. In<br />
animals and fungi, PLCs specific for the substrate<br />
Structures <strong>of</strong> some lipid second messengers<br />
O<br />
Phosphatidylinositol (PI)<br />
O<br />
O<br />
O<br />
HO<br />
OH<br />
O<br />
2<br />
O<br />
3<br />
P<br />
1<br />
O -<br />
O<br />
6 OH<br />
4 5<br />
OH<br />
OH<br />
O<br />
Phosphatidylinositol-3,4,5-trisphosphate (PIP 3 )<br />
O<br />
O<br />
O<br />
H - O 3 PO<br />
HO<br />
OH<br />
OH<br />
O -<br />
O<br />
P<br />
O<br />
O<br />
6 OH<br />
2 1 4 5<br />
3<br />
OPO 3 H -<br />
OPO 3 H -<br />
O<br />
Diacylglycerol (DAG)<br />
O<br />
O<br />
O<br />
OH<br />
OPO 3 H -<br />
6 OH<br />
2 1 4 5<br />
3<br />
Inositol trisphosphate (IP 3 )<br />
OPO 3 H -<br />
OPO 3 H -<br />
O<br />
Phosphatidic acid (PA)<br />
O<br />
O<br />
O<br />
O<br />
O-<br />
P<br />
O -<br />
O<br />
FIGURE 14.17 Structures <strong>of</strong> some lipid second messengers and the common precursor phosphatidylinositol.<br />
The acyl side chain structures shown here are the most common for mammalian PI lipids. Much <strong>of</strong> the PA in<br />
<strong>cell</strong>s is derived from PC, and its acyl chains may differ from those shown.<br />
610 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 611<br />
phosphatidylinositol-4,5-bisphosphate (PIP 2<br />
)<br />
hydrolyze PIP 2<br />
to form two second messengers:<br />
1,2-sn-diacylglycerol (DAG) and inositol-1,4,5-<br />
trisphosphate (IP 3<br />
). The PLC substrate PIP 2<br />
is itself<br />
an important regulatory ligand that<br />
modulates the activity <strong>of</strong> several ion channels,<br />
transporters, and enzymes. Thus, PLC alters concentration<br />
<strong>of</strong> three second messengers; its net<br />
effect depends on the net turnover <strong>of</strong> the substrate<br />
and products.<br />
DAG is probably the best known lipid second<br />
messenger; its hydrophobicity limits it to action<br />
in membranes. DAG activates some is<strong>of</strong>orms<br />
<strong>of</strong> protein kinase C (PKC), modulates the activity<br />
<strong>of</strong> several cation channels and activates at<br />
least one other protein kinase. DAG can be further<br />
hydrolyzed to release arachidonic acid,<br />
which can regulate some ion channels.<br />
Arachidonic acid is also the precursor <strong>of</strong> oxidation<br />
products, such as prostaglandins and thromboxanes,<br />
which are potent extra<strong>cell</strong>ular <strong>signaling</strong><br />
agents. In addition to DAG, PKCs require interaction<br />
with Ca2+ and an acidic phospholipid,<br />
such as phosphatidylserine, to become activated.<br />
Thus, activation <strong>of</strong> PKC requires the coincidence<br />
<strong>of</strong> multiple inputs both to generate DAG and to<br />
increase intra<strong>cell</strong>ular Ca2+. There are more than<br />
a dozen PKCs, classified together according to<br />
highly conserved sequences in the catalytic domain.<br />
Three subgroups <strong>of</strong> PKCs, also identifiable<br />
by sequence, share different patterns <strong>of</strong> regulation.<br />
Their regulation provides examples <strong>of</strong><br />
many ways in which other mammalian protein<br />
kinases are regulated.<br />
The first <strong>of</strong> these groups, canonical PKCs,<br />
are generally soluble or very loosely associated<br />
with membranes prior to the appearance <strong>of</strong><br />
DAG. DAG causes their association with membranes<br />
and permits activation upon binding <strong>of</strong><br />
other regulators. The second group <strong>of</strong> PKCs requires<br />
similar lipids but not Ca2+, and the third<br />
group requires other lipids but neither DAG nor<br />
Ca2+ for activation.<br />
The N-terminal region <strong>of</strong> PKCs contains a<br />
pseudosubstrate domain, a sequence that resembles<br />
that <strong>of</strong> a typical substrate except that<br />
the target Ser is replaced with Ala. The pseudosubstrate<br />
region binds to the active site to inhibit<br />
the kinase. Activators cause the<br />
pseudosubstrate domain to flip out <strong>of</strong> the active<br />
site. PKCs are also activated by proteolysis,<br />
as are many protein kinases with discrete<br />
autoinhibitory domains. Proteases clip a flexible<br />
hinge region, which results in loss <strong>of</strong> the<br />
regulatory domain and consequent activation<br />
<strong>of</strong> the kinase.<br />
PKC is the major receptor for phorbol esters,<br />
a class <strong>of</strong> powerful tumor promoters. Phorbol<br />
esters mimic DAG and cause a more massive<br />
and prolonged activation than physiological<br />
stimuli. This massive stimulation can induce<br />
proteolysis <strong>of</strong> PKC, resulting in downregulation,<br />
or loss <strong>of</strong> the kinase. (For a personal description<br />
on the discovery <strong>of</strong> protein kinase C<br />
see EXP : 14-0001 )<br />
IP 3<br />
, the second product <strong>of</strong> the PLC reaction,<br />
is a soluble second messenger. The most significant<br />
IP 3<br />
target is a Ca2+ channel in the endoplasmic<br />
reticulum. IP 3<br />
causes this channel to<br />
open and release stored Ca2+ into the cytoplasm,<br />
thereby rapidly elevating the cytosolic Ca2+ over<br />
100-fold and, in turn, causing the activation <strong>of</strong><br />
numerous targets <strong>of</strong> Ca2+ <strong>signaling</strong>.<br />
There are at least six families <strong>of</strong> PIP 2<br />
-selective<br />
PLC enzymes, defined by their distinct forms<br />
<strong>of</strong> regulation, domain compositions, and overall<br />
sequence conservation. Their catalytic domains<br />
are all quite similar. The PLC-βs are<br />
stimulated primarily by Gα q<br />
and Gβγ (to individually<br />
varying extents). Several are also modulated<br />
by phosphorylation. PLC-γ is<strong>of</strong>orms are<br />
stimulated by phosphorylation on Tyr residues,<br />
frequently by receptor tyrosine kinases. The<br />
PLC-ε is<strong>of</strong>orms are regulated by small,<br />
monomeric G proteins <strong>of</strong> the Rho family. The<br />
regulation <strong>of</strong> the PLC-δs is still incompletely understood.<br />
Two other classes similar to the PLCδs,<br />
PLC-η and -ζ, have also been defined recently.<br />
(There is no PLC-α.) In addition to their distinct<br />
modes <strong>of</strong> regulation, all <strong>of</strong> the PLCs are stimulated<br />
by Ca2+, and Ca2+ <strong>of</strong>ten acts synergistically<br />
with other stimulatory inputs. This synergy<br />
underlies the intensification and prolongation<br />
<strong>of</strong> Ca2+ <strong>signaling</strong> observed in many <strong>cell</strong>s.<br />
Phospholipases A 2<br />
and D (PLA 2<br />
and PLD)<br />
also hydrolyze glycerol phospholipids in <strong>cell</strong><br />
membranes to form important <strong>signaling</strong> compounds.<br />
PLA 2<br />
hydrolyzes the fatty acid at the sn-<br />
2 position <strong>of</strong> multiple phospholipids to produce<br />
the cognate lysophospholipid and the free fatty<br />
acid, which is generally unsaturated. The free<br />
fatty acid is <strong>of</strong>ten arachidonic acid, a precursor<br />
<strong>of</strong> extra<strong>cell</strong>ular signals. The biological roles <strong>of</strong><br />
free lysophospholipids are not understood in<br />
detail but have been linked to effects on the<br />
structure <strong>of</strong> the membrane bilayer.<br />
PLD catalyzes a reaction much like that <strong>of</strong><br />
PLC but instead hydrolyzes the phosphodiester<br />
on the substituent side <strong>of</strong> the phosphate group<br />
to form 3-sn-phosphatidic acid. Cellular PLDs act<br />
on multiple glycerol phospholipid substrates,<br />
but phosphatidylcholine is probably the sub-<br />
14.16 Lipids and lipid-derived compounds are <strong>signaling</strong> molecules 611
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 612<br />
FIGURE 14.18 Activated PI 3-<br />
kinase phosphorylates PIP 2<br />
to produce<br />
PIP 3<br />
. The PH domain-containing<br />
protein kinases PDK1 and Akt<br />
bind to PIP 3<br />
at the plasma membrane.<br />
Their colocalization facilitates<br />
the phosphorylation <strong>of</strong> Akt<br />
by PDK1. A second phosphorylation<br />
within a hydrophobic motif results<br />
in Akt activation by one <strong>of</strong><br />
several candidate protein kinases.<br />
The Akt-2 is<strong>of</strong>orm is required to<br />
elicit hallmark actions <strong>of</strong> insulin.<br />
p85 p110<br />
PI 3-kinase<br />
PIP 2 PIP 3<br />
strate most relevant to <strong>signaling</strong> functions. The<br />
functions <strong>of</strong> the phosphatidic acid product,<br />
which is also formed by phosphorylation <strong>of</strong><br />
DAG, remain poorly understood but appear to<br />
include a role in secretion and the fusion <strong>of</strong> intra<strong>cell</strong>ular<br />
membranes.<br />
14.17<br />
PI 3-kinase regulates<br />
both <strong>cell</strong> shape and the<br />
activation <strong>of</strong> essential<br />
growth and metabolic<br />
functions<br />
Key concepts<br />
• Phosphorylation <strong>of</strong> some lipid second messengers<br />
changes their activity.<br />
• PIP 3<br />
is recognized by proteins with a pleckstrin<br />
homology domain.<br />
Lipid second messengers may also be modified<br />
by phosphorylation. PI 3-kinase phosphorylates<br />
PIP 2<br />
on the 3-position <strong>of</strong> the inositol ring to<br />
form PI 3,4,5-P 3<br />
, another lipid second messenger.<br />
The total activity <strong>of</strong> PI 3-kinase is too low<br />
to significantly deplete total PIP 2<br />
, but formation<br />
<strong>of</strong> small amounts <strong>of</strong> PIP 3<br />
in localized membrane<br />
domains is vital for altering <strong>cell</strong> shape<br />
and <strong>cell</strong>ular motility.<br />
PIP 3<br />
acts by recruiting proteins that contain<br />
PIP 3<br />
binding domains, including pleckstrin<br />
homology (PH) and FYVE domains, to sites<br />
where they regulate cytoskeletal remodeling,<br />
contractile protein function, or other regulatory<br />
events. These proteins anchor and/or orient<br />
the structural or motor proteins involved<br />
in <strong>cell</strong>ular movement and localize <strong>signaling</strong> proteins<br />
to sites <strong>of</strong> action at the membrane. PIP 3<br />
PIP 3 binding brings Akt and PDK1 to the membrane<br />
PIP 2 phosphorylated<br />
Akt<br />
Akt and PDK1<br />
bind PIP3 through<br />
PH domains<br />
Akt<br />
PH<br />
domains<br />
PDK1<br />
PDK1<br />
Akt is activated by<br />
phosphorylation<br />
Other<br />
kinase<br />
Glucose uptake<br />
Glycogen synthesis<br />
Antilipolysis<br />
Antiapoptosis<br />
<strong>signaling</strong> can be fast and dramatic; it largely accounts<br />
for directing the mobility <strong>of</strong> motile mammalian<br />
<strong>cell</strong>s.<br />
Lipid mediators are essential in the insulin<br />
<strong>signaling</strong> pathway. The binding <strong>of</strong> insulin stimulates<br />
the Tyr autophosphorylation <strong>of</strong> its receptor<br />
and the activation <strong>of</strong> effectors through insulin receptor<br />
substrate (IRS) proteins (see 14.30 Diverse<br />
<strong>signaling</strong> mechanisms are regulated by protein tyrosine<br />
kinases). PI 3-kinase is activated when its p85<br />
subunit binds to IRS1. The PIP 3<br />
generated by PI3-<br />
kinase binds the protein kinases Akt and phosphoinositide-dependent<br />
kinase-1 (PDK-1) via their PH<br />
domains. This interaction results in the localization<br />
<strong>of</strong> Akt to the membrane where it is activated<br />
by PDK1, as illustrated in FIGURE 14.18. Akt phosphorylates<br />
downstream targets, including protein<br />
kinases, GAPs, and transcription factors.<br />
Activation <strong>of</strong> Akt, specifically Akt-2, is required<br />
for the hallmark actions <strong>of</strong> insulin including regulation<br />
<strong>of</strong> glucose transporter translocation, enhanced<br />
protein synthesis, and expression <strong>of</strong><br />
gluconeogenic and lipogenic enzymes.<br />
14.18<br />
Signaling through ion<br />
channel receptors is very<br />
fast<br />
Key concepts<br />
• Ion channels allow the passage <strong>of</strong> ions through a<br />
pore, resulting in rapid (microsecond) changes in<br />
membrane potential.<br />
• Channels are selective for particular ions or for<br />
cations or anions.<br />
• Channels regulate intra<strong>cell</strong>ular concentrations <strong>of</strong><br />
regulatory ions, such as Ca2+.<br />
Ligand-gated ion channels are multisubunit,<br />
membrane-spanning proteins that create and<br />
regulate a water-filled pore through the membrane,<br />
as illustrated in the X-ray crystal structure<br />
<strong>of</strong> the nicotinic acetylcholine receptor in<br />
FIGURE 14.19. When stimulated by extra<strong>cell</strong>ular<br />
agonists, the subunits rearrange their conformations<br />
and orientations to open the pore and, thus,<br />
connect the aqueous spaces on either side <strong>of</strong> the<br />
membrane. The pore has a diameter that allows<br />
ions to diffuse freely from one side <strong>of</strong> the membrane<br />
to the other, driven by the electrical and<br />
chemical gradients that have been established<br />
by ion pumps and transporters. (For more about<br />
channel, pump and transporter mechanics see 2<br />
Transport <strong>of</strong> ions and small molecules across membranes.)<br />
Channels maintain selectivity among<br />
ions by regulating the pore diameter precisely<br />
612 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 613<br />
and by lining the walls <strong>of</strong> the pore with appropriate<br />
hydrophilic residues. Receptor ion channels<br />
can, thus, provide a diffusion path for only<br />
cations or anions, or select among different ions.<br />
Ligand-gated ion channels provide the<br />
fastest signal transduction mechanism found in<br />
biology. Upon binding an agonist ligand, channels<br />
open within microseconds. At synapses,<br />
where neurotransmitters need to diffuse less<br />
than 0.1 micron, a signal in the postsynaptic<br />
<strong>cell</strong> can be generated in 100 microseconds. In<br />
contrast, receptor-stimulated G proteins require<br />
about 100 milliseconds to exchange GDP for<br />
GTP, and the action <strong>of</strong> receptor protein kinases<br />
is even slower. Ligand-gated ion channels are<br />
important receptors in many <strong>cell</strong>s in addition<br />
to neurons and muscle, and other ion channels<br />
play equally vital roles in <strong>signaling</strong> pathways<br />
triggered by other classes <strong>of</strong> ligands.<br />
Ion channel <strong>signaling</strong> differs from that <strong>of</strong><br />
the other receptors mentioned in this chapter<br />
in that there is no immediate protein target nor,<br />
in most cases, is there a specific second messenger<br />
involved. In most cases, channel-mediated<br />
ion flow acts to increase or decrease the <strong>cell</strong>’s<br />
membrane potential and, thus, modulates all<br />
transport processes for metabolites or ions that<br />
are electrically driven.<br />
Animal <strong>cell</strong>s maintain an inside-negative<br />
membrane potential by pumping out Na + ions<br />
and pumping in K + ions (for more on membrane<br />
potential see 2.4 Electrochemical gradients<br />
across the <strong>cell</strong> membrane generate the membrane potential).<br />
The opening <strong>of</strong> a channel selective for<br />
Na+ will thus depolarize <strong>cell</strong>s, and the opening<br />
<strong>of</strong> a channel for K + will hyperpolarize <strong>cell</strong>s.<br />
Similarly, because Cl - is primarily extra<strong>cell</strong>ular,<br />
opening Cl - channels will also cause hyperpolarization.<br />
These electrical effects convey information<br />
to effector proteins that are energetically<br />
coupled to the membrane potential, or to specific<br />
ion gradients, or that bind a specific ion<br />
(such as Ca2+) whose concentration changes<br />
upon channel opening.<br />
The nicotinic acetylcholine receptor is the prototypical<br />
receptor ion channel and was the first<br />
receptor that was shown to be a channel. It is a relatively<br />
unselective cation channel that causes depolarization<br />
<strong>of</strong> the target <strong>cell</strong> by allowing Na+<br />
influx. It is best known as the excitatory receptor<br />
at the neuromuscular synapse, where it triggers<br />
contraction, but alternative is<strong>of</strong>orms are also active<br />
in neurons and many other <strong>cell</strong>s. In muscles,<br />
nicotinic depolarization acts via a voltage-sensitive<br />
Ca2+ channel to allow Ca2+ release from the sarcoplasmic<br />
reticulum into the cytosol. Calcium acts<br />
Nicotinic acetylcholine receptor structure<br />
CLOSED<br />
CYTOSOL<br />
Pore<br />
OPEN<br />
Pore<br />
FIGURE 14.19 The nicotinic cholinergic receptor is a cation-selective channel<br />
that is composed <strong>of</strong> five homologous but usually nonidentical subunits that<br />
oligomerize to form a primarily -helical membrane-spanning core. The channel<br />
itself is created within this core, and its opening and closing are executed<br />
by cooperative changes in subunit arrangement. Structure generated from<br />
Protein Data Bank file 2BG9.<br />
as a second (or third) messenger to initiate contraction<br />
(see 2.13 Cardiac and skeletal muscles are activated<br />
by excitation-contraction coupling). Nicotinic<br />
receptors promote exocytosis in some secretory<br />
<strong>cell</strong>s by a similar mechanism, where Ca2+ triggers<br />
the exocytic event. In neurons, where nicotinic<br />
stimulation causes an action potential (depolarization<br />
that is rapidly propagated along the neuron),<br />
the initial depolarization is sensed by<br />
voltage-sensitive Na+ channels. Their opening<br />
(along with the action <strong>of</strong> other channels) propagates<br />
the action potential along the neuron.<br />
The nervous system is rich in receptor cation<br />
channels that respond to other neurotransmitters,<br />
the most common <strong>of</strong> which is the amino<br />
acid glutamate (Glu). The three different families<br />
<strong>of</strong> glutamate receptors share the property<br />
<strong>of</strong> cation conductance, but each family has its<br />
own spectrum <strong>of</strong> drug responses. All operate as<br />
neuronal activators, with one interesting twist:<br />
The NMDA family <strong>of</strong> receptors, named for their<br />
response to a selective drug, is permeant to Ca2+<br />
in addition to Na+. A significant component <strong>of</strong><br />
its activity is to permit the inward flow <strong>of</strong> Ca2+,<br />
which acts as a second messenger on a wide variety<br />
<strong>of</strong> targets. Persistant stimulation <strong>of</strong> NMDA<br />
channels by glutamate released during injury,<br />
or by drugs, can cause toxic amounts <strong>of</strong> Ca2+ to<br />
enter, resulting in neuronal death.<br />
A second functional group <strong>of</strong> receptor channels<br />
is selective for anions and, by allowing inward<br />
flux <strong>of</strong> Cl - , hyperpolarizes the target <strong>cell</strong>.<br />
Anion-selective receptors include those for γ-<br />
aminobutyric acid (GABA) and glycine (Gly). In<br />
neurons, hyperpolarization can inhibit the initiation<br />
<strong>of</strong> an action potential and/or neurotransmitter<br />
release.<br />
14.18 Signaling through ion channel receptors is very fast 613
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 614<br />
Perhaps the most diverse family <strong>of</strong> ligandgated<br />
channels is that <strong>of</strong> the TRP and TRP-like<br />
family, <strong>of</strong> which about 30 have been found in<br />
mammals. Distinct forms are found in invertebrates.<br />
The TRP channels are Ca2+-selective<br />
channels that are formed by tetramers <strong>of</strong> identical<br />
subunits that surround the central channel.<br />
Each subunit is composed <strong>of</strong> a homologous<br />
bundle <strong>of</strong> six membrane-spanning helices, but<br />
the N and C termini contain a diverse collection<br />
<strong>of</strong> regulatory and protein interaction domains,<br />
including protein kinase domains (whose<br />
substrates are currently unknown).<br />
All TRP channels allow transmembrane flux<br />
<strong>of</strong> Ca2+ to permit its action as a second messenger,<br />
but different TRP is<strong>of</strong>orms serve numerous<br />
physiological functions. The prototypical TRP,<br />
found in invertebrate photoreceptors, gates Ca2+<br />
flow from intra<strong>cell</strong>ular stores into the cytoplasm<br />
to initiate visual <strong>signaling</strong>. Others admit Ca2+<br />
from outside the <strong>cell</strong>, and still others allow Ca2+<br />
to enter the endoplasmic reticulum virtually directly<br />
from the extra<strong>cell</strong>ular space because they<br />
form a bridge between the plasma membrane<br />
and channels in the endoplasmic reticulum at<br />
points where the membranes abut each other.<br />
Regulation <strong>of</strong> TRP channels is perhaps even<br />
more diverse. Various TRP channels respond to<br />
heat, cold, painful stimuli, pressure, and high<br />
or low osmolarity. Many TRPs are regulated either<br />
positively or negatively by lipids, such as<br />
eicosanoids, diacylglycerol, and PIP 2<br />
. For example,<br />
capsaicin, the hot compound in chilis, is<br />
an agonist for some vanilloid receptors (TRPVs).<br />
Still other TRP channels are mechanosensors<br />
that allow cilia to sense fluid flow. The most famous<br />
<strong>of</strong> these is the sensory channel <strong>of</strong> the hair<br />
<strong>cell</strong> <strong>of</strong> the inner ear. This channel opens when<br />
the apical cilia on the hair <strong>cell</strong> are bent in response<br />
to sound-driven fluid flow.<br />
14.19<br />
Nuclear receptors<br />
regulate transcription<br />
Key concepts<br />
• Nuclear receptors modulate transcription by<br />
binding to distinct short sequences in<br />
chromosomal DNA known as response elements.<br />
• Receptor binding to other receptors, inhibitors, or<br />
coactivators leads to complex transcriptional<br />
control circuits.<br />
• Signaling through nuclear receptors is relatively<br />
slow, consistent with their roles in adaptive<br />
responses.<br />
Nuclear receptors are unique among <strong>cell</strong>ular<br />
receptors in that their ligands pass unaided<br />
through the plasma membrane. These receptors,<br />
when complexed with their ligands, enter<br />
the nucleus and regulate gene transcription.<br />
Ligands for nuclear receptors include sex steroids<br />
(estrogen and testosterone) and other steroid<br />
hormones, vitamins A and D, retinoids and<br />
other fatty acids, oxysterols, and bile acids.<br />
Nuclear receptors are structurally conserved.<br />
They consist <strong>of</strong> a C-terminal ligand binding domain,<br />
an N-terminal interaction region that recognizes<br />
components <strong>of</strong> the transcriptional<br />
machinery and acts as a transactivation domain,<br />
a centrally located zinc finger domain that binds<br />
DNA, and, <strong>of</strong>ten, another transactivation domain<br />
nearer the C-terminus. In the absence <strong>of</strong><br />
ligand, these receptors are bound to corepressor<br />
proteins that suppress their activity. Upon<br />
hormone binding, corepressors dissociate and<br />
the receptors are assembled in multiprotein<br />
complexes with coactivators that modulate receptor<br />
action and facilitate transcriptional regulation.<br />
As illustrated in FIGURE 14.20, agonists<br />
and antagonists bind to distinct receptor conformations<br />
(see 14.5 Ligand binding changes receptor<br />
conformation). Receptor agonists favor the binding<br />
<strong>of</strong> receptors to coactivators and DNA, and<br />
antagonists favor conformations that block coactivator-receptor<br />
binding.<br />
Nuclear receptors bind with high specificity<br />
to hormone response elements in the 5’ untranscribed<br />
region <strong>of</strong> regulated genes. Response<br />
elements are typically short direct or inverted<br />
repeat sequences, and a gene may contain response<br />
elements for several different receptors<br />
in addition to binding sites for other transcriptional<br />
regulatory proteins.<br />
The sex steroid estrogen can bind to two<br />
different nuclear receptors, the estrogen receptors<br />
ER and ER. Coactivator and corepressor<br />
proteins differentially regulate ER and ER in<br />
transcriptional complexes that are expressed in<br />
specific <strong>cell</strong> types. Other ligands that bind to<br />
these receptors include valuable therapeutic<br />
agents. For example, 4 hydroxy-tamoxifen is<br />
an estrogen receptor antagonist used in the therapy<br />
<strong>of</strong> estrogen-receptor-positive breast cancer<br />
to inhibit growth <strong>of</strong> residual cancer <strong>cell</strong>s.<br />
However, unlike its antagonistic effects on the<br />
estrogen receptor in breast, 4 hydroxy-tamoxifen<br />
displays weak partial agonist activity in<br />
uterus. In the estrogen receptor system, partial<br />
agonists are known as selective estrogen receptor<br />
modulators (SERMs). Properties that contribute<br />
to partial agonist activity include the<br />
relative expression <strong>of</strong> the two estrogen recep-<br />
614 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 615<br />
Estrogen receptor conformation depends on which ligand is bound<br />
Agonist-bound<br />
conformation<br />
Antagonist-bound<br />
conformation<br />
N<br />
N<br />
K362<br />
H11<br />
H5<br />
K362<br />
H5<br />
545<br />
C<br />
H6<br />
545<br />
542<br />
H3<br />
H11<br />
H6<br />
538<br />
542<br />
H3<br />
538<br />
FIGURE 14.20 The estrogen receptor adopts different conformations when<br />
bound to agonists and antagonists. The ligand-binding domain <strong>of</strong> the estrogen<br />
receptor is bound to the agonist estradiol on the left and to the antagonist<br />
raloxifene on the right. Note the marked difference in position <strong>of</strong> helix 12,<br />
shown in blue in the active structure and green in the inhibited structure.<br />
Reproduced from Brzozowski, A. M., et al. 1997. Molecular basis <strong>of</strong> agonism and<br />
antagonism in the oestrogen receptor. Nature. 389: 753–758. Photo courtesy<br />
<strong>of</strong> M. Brzozowski, University <strong>of</strong> New York.<br />
tors, ER and ER, as well as the expression <strong>of</strong><br />
repressors and coactivators that interact with<br />
each receptor type. Thus, the behavior <strong>of</strong> nuclear<br />
receptor ligands must be considered in the<br />
tissue, <strong>cell</strong>ular, and <strong>signaling</strong> context.<br />
14.20<br />
G protein <strong>signaling</strong><br />
modules are widely used<br />
and highly adaptable<br />
Key concepts<br />
• The basic module is a receptor, a G protein and an<br />
effector protein.<br />
• Cells express several varieties <strong>of</strong> each class <strong>of</strong><br />
proteins.<br />
• Effectors are heterogeneous and initiate diverse<br />
<strong>cell</strong>ular functions.<br />
Activation <strong>of</strong> G protein-coupled receptors<br />
(GPCRs) and their associated heterotrimeric G<br />
proteins is one <strong>of</strong> the most widespread mechanisms<br />
<strong>of</strong> communicating extra<strong>cell</strong>ular signals<br />
to the intra<strong>cell</strong>ular environment. G protein <strong>signaling</strong><br />
modules are found in all eukaryotes.<br />
Depending on the species, mammals express<br />
500-1000 GPCRs that respond to hormones,<br />
neurotransmitters, pheromones, metabolites,<br />
local <strong>signaling</strong> substances, and other regulatory<br />
molecules. Essentially all chemical classes are<br />
represented among the GPCR ligands. In addition,<br />
a roughly equal number <strong>of</strong> olfactory GPCRs<br />
are expressed in olfactory neurons and work in<br />
combination to screen compounds in the animal’s<br />
environment via the sense <strong>of</strong> smell.<br />
Because GPCRs are involved in many kinds <strong>of</strong><br />
physiologic responses, they are also one <strong>of</strong> the<br />
most widely used targets for drugs.<br />
A minimal G protein <strong>signaling</strong> module consists<br />
<strong>of</strong> three proteins: a G protein-coupled receptor,<br />
the heterotrimeric G protein, and an<br />
effector protein, as illustrated in FIGURE 14.21. The<br />
receptor activates the G protein on the inner face<br />
<strong>of</strong> the plasma membrane in response to an extra<strong>cell</strong>ular<br />
ligand. The G protein then activates (or<br />
occasionally inhibits) an effector protein that<br />
propagates a signal within the <strong>cell</strong>. Thus, signal<br />
conduction in the simplest G protein module is<br />
linear. However, as depicted in FIGURE 14.22, a<br />
typical animal <strong>cell</strong> may express a dozen GPCRs,<br />
more than six G proteins, and a dozen effectors.<br />
Each GPCR regulates one or more G proteins,<br />
and each G protein regulates several effectors.<br />
Moreover, distinct efficiencies and rates govern<br />
each interaction. Thus, a <strong>cell</strong>’s G protein network<br />
is actually a signal-integrating computer whose<br />
14.20 G protein <strong>signaling</strong> modules are widely used and highly adaptable 615
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 616<br />
Agonist<br />
GPCR<br />
Trimeric<br />
G protein<br />
Receptor<br />
activated<br />
CYTOSOL<br />
Heterotrimeric G protein <strong>signaling</strong><br />
Activated<br />
G protein<br />
dissociates<br />
IP 3 -gated<br />
Ca 2+ channel<br />
ENDOPLASMIC<br />
RETICULUM<br />
PIP 2<br />
IP 3<br />
Ca 2+<br />
DAG<br />
Hydrolysis <strong>of</strong> PIP 2<br />
to IP 3 and DAG<br />
Release <strong>of</strong> Ca 2+<br />
FIGURE 14.21 G protein-mediated signal transduction follows a path <strong>of</strong> agonist<br />
to receptor to heterotrimeric G protein to effector to the effector's output.<br />
Both G and G subunits regulate distinct effectors. In the example<br />
shown here, G q<br />
regulates a phospholipase C- to produce two second messengers,<br />
diacyglycerol (DAG) and inositol-trisphosphate (IP 3<br />
). IP 3<br />
triggers Ca2+ release<br />
from the endoplasmic reticulum.<br />
-<br />
output is a spectrum <strong>of</strong> <strong>cell</strong>ular signals that is<br />
complex in both amplitude and kinetics. Because<br />
<strong>of</strong> their conserved parts list, G protein modules<br />
are well suited to initiating a wide variety <strong>of</strong> intra<strong>cell</strong>ular<br />
signals in response to diverse molecular<br />
inputs and can do so over a wide range <strong>of</strong><br />
time scales (milliseconds to minutes).<br />
GPCRs are integral plasma membrane proteins<br />
composed <strong>of</strong> a bundle <strong>of</strong> seven hydrophobic<br />
membrane-spanning helices with an<br />
extra<strong>cell</strong>ular N terminus and cytosolic C terminus,<br />
as depicted in FIGURE 14.23. Based on the<br />
three-dimensional structure <strong>of</strong> rhodopsin and<br />
on copious biochemical and genetic data, it is<br />
likely that all GPCRs share the same basic mechanism<br />
<strong>of</strong> conformational activation and deactivation<br />
in response to activating ligands (see 14.5<br />
Ligand binding changes receptor conformation).<br />
Binding <strong>of</strong> agonist ligand on the extra<strong>cell</strong>ular<br />
face <strong>of</strong> the receptor drives realignment <strong>of</strong> the helices<br />
to alter the structure <strong>of</strong> a binding site for the<br />
heterotrimeric G protein on the cytoplasmic face,<br />
and this altered conformation <strong>of</strong> the G proteinbinding<br />
surface promotes G protein activation.<br />
FIGURE 14.22 A portion <strong>of</strong> the G protein-mediated<br />
<strong>signaling</strong> network in<br />
macrophages highlights some <strong>of</strong> the<br />
complexity <strong>of</strong> interactions possible in<br />
such systems. Several receptors and G<br />
protein subunits are omitted. Where a<br />
named G protein is shown, its <strong>signaling</strong><br />
output is probably mediated by its G<br />
subunit. Activation <strong>of</strong> any G protein also<br />
activates its G subunit, although Gmediated<br />
<strong>signaling</strong> is usually most prominent<br />
from G i<br />
trimers. In addition, several<br />
G proteins modulate the activities <strong>of</strong><br />
others through poorly understood pathways.<br />
Only a small sampling <strong>of</strong> effectors<br />
is shown, and the only adaptive mechanism<br />
shown is GRK-catalyzed phosphorylation<br />
<strong>of</strong> receptors. Data from Paul<br />
Sternweis, Alliance for Cellular Signaling.<br />
Agonist C5a ISO PGE S1P UDP <strong>UT</strong>P PAF LPA<br />
GPCR<br />
G Protein<br />
Effector<br />
Partial G protein <strong>signaling</strong> network in mouse macrophages<br />
C5aR β 2 AR E2R EDG P2YR P2YR PAFR EDG<br />
Gi Gs G12<br />
Gq<br />
Ad<br />
Cyc<br />
cAMP<br />
ATP<br />
PI 3-<br />
Kinase<br />
PIP 2<br />
PLC-β<br />
G12<br />
??<br />
PIP 3<br />
DAG +<br />
IP 3 IP 2 + P i<br />
Phosphatase<br />
Inactivation<br />
mechanisms<br />
cAMP<br />
AMP<br />
PDE<br />
Ca2+<br />
Ca2+<br />
pump<br />
IP 3 R<br />
GRK<br />
616 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 617<br />
Structure <strong>of</strong> rhodopsin<br />
Heterotrimeric G protein structure<br />
CYTOPLASM<br />
MEMBRANE<br />
Retinal<br />
FIGURE 14.23 The figure shows the crystal structure<br />
<strong>of</strong> the GPCR rhodopsin. Each membrane-spanning helix<br />
is a different color; most structures on the cytoplasmic<br />
face are not shown. The retinal chromophore is<br />
shown within the helix bundle. GPCR sequence similarity<br />
separates the mammalian GPCRs into at least four<br />
structural families that are so diverse that there may<br />
be little sequence similarity among the classes. Within<br />
a family, similarity is greatest in the membrane-spanning<br />
helices, less in the interhelical loops, and least in<br />
the N- and C-terminal domains and in the cytoplasmic<br />
loop that connects spans five and six. Regardless, the<br />
generalizations about functional domains in receptors<br />
seem to hold true within different families. GPCRs frequently<br />
form dimers, occasionally heterodimers, and<br />
dimerization can be crucial for function. Structure generated<br />
from Protein Data Bank file 1F88.<br />
The heterotrimeric G proteins to which<br />
GPCRs are coupled are composed <strong>of</strong> a nucleotide-binding<br />
Gα subunit and a Gβγ subunit<br />
dimer, as illustrated in FIGURE 14.24. The structure<br />
<strong>of</strong> the trimer and each subunit is known for<br />
several states <strong>of</strong> activation and in complex with<br />
several interacting proteins. A Gαβγ heterotrimer<br />
is named according to its α subunit, which largely<br />
defines the G protein’s selectivity among receptors.<br />
Each subunit also regulates a distinct group<br />
<strong>of</strong> effector proteins.<br />
Gα subunits are globular, two-domain proteins<br />
<strong>of</strong> 38-44 kDa. The GTP-binding domain<br />
belongs to the GTP-binding protein superfamily<br />
that includes the small, monomeric G proteins<br />
(such as Ras, Rho, Arf, Rab; see 14.23 Small,<br />
monomeric GTP-binding proteins are multiuse<br />
switches) as well as the GTP-binding translational<br />
initiation and elongation factors. A second domain<br />
modulates GTP binding and hydrolysis.<br />
Gα subunits are only slightly hydrophobic, but<br />
they are predominantly membrane-associated<br />
FIGURE 14.24 The structure <strong>of</strong> the nonactivated G i<br />
heterotrimer,<br />
the G protein that is responsible for inhibition <strong>of</strong><br />
adenylyl cyclase and for most G-mediated <strong>signaling</strong>, is<br />
shown with each subunit colored as shown. GDP is shown<br />
bound to the G i<br />
subunit. Structure generated from Protein<br />
Data Bank file 1GP2.<br />
G<br />
protein<br />
Gs<br />
Golf<br />
Gi (3)<br />
Go<br />
Gz<br />
Ggus<br />
Gt (2)<br />
Gq (4)<br />
G 12<br />
G 13<br />
Adenylyl cyclase<br />
EFFECTOR PROTEIN<br />
K+channel, PI 3-kinase<br />
Other cation channel<br />
Rho GEF<br />
G protein targets<br />
Stimulated<br />
Cyclic GMP phosphodiesterase<br />
Phospholipase-Cβ<br />
Inhibited<br />
Adenylyl cyclase<br />
FIGURE 14.25 G protein-regulated effectors do not share structural<br />
similarities. They may be ion channels or membrane spanning<br />
enzymes in the plasma membrane, peripheral proteins on the<br />
inner face <strong>of</strong> the membrane, or fundamentally soluble proteins<br />
that can bind to G subunits. The chart shows the major groups<br />
<strong>of</strong> G proteins, sorted according to sequence similarity, and some<br />
<strong>of</strong> the effectors that they are known to regulate.<br />
because <strong>of</strong> constitutive N-terminal fatty acylation<br />
and because they bind to the membraneattached<br />
Gβγ subunits. Mammals have 16 Gα<br />
genes that are grouped in subfamilies according<br />
to similar sequence and function (e.g., s, i, q,<br />
and 12). These subfamilies are listed in FIGURE<br />
14.25.<br />
Gβ and Gγ subunits associate irreversibly<br />
soon after translation to form stable Gβγ dimers,<br />
which then associate reversibly with a Gα. Gβ<br />
subunits are 35 kDa proteins composed <strong>of</strong> seven<br />
14.20 G protein <strong>signaling</strong> modules are widely used and highly adaptable 617
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 618<br />
-strand repeats that form a cylindrical structure<br />
known as a propeller. There are five Gβ genes<br />
in mammals. Four encode strikingly similar proteins<br />
that naturally dimerize with the twelve Gγ<br />
subunits (Figure 14.24). The fifth, Gβ5, is less<br />
closely related to the others and interacts primarily<br />
with a Gγ-like domain in other proteins rather<br />
than with Gγ subunits themselves.<br />
Gγ subunits are smaller (~7 kDa) and far<br />
more diverse in sequence than are the Gβ’s. The<br />
last three amino acid residues <strong>of</strong> Gγ subunits<br />
are proteolyzed to leave a conserved C-terminal<br />
cysteine that is irreversibly S-prenylated<br />
and carboxymethylated, helping to anchor Gβγ<br />
to the membrane. Gβ and Gγ subunits can associate<br />
in most possible combinations. Because<br />
almost all <strong>cell</strong>s express multiple Gβ and Gγ subunits,<br />
it has been difficult to assign specific roles<br />
to individual Gβγ combinations. The best recognized<br />
interactions <strong>of</strong> Gβγ subunits occur at<br />
sites on Gβ, although distinct functions <strong>of</strong> Gγ<br />
have also been supported.<br />
14.21<br />
Heterotrimeric G proteins<br />
regulate a wide variety <strong>of</strong><br />
effectors<br />
Key concepts<br />
• G proteins convey signals by regulating the<br />
activities <strong>of</strong> multiple intra<strong>cell</strong>ular <strong>signaling</strong><br />
proteins known as effectors.<br />
• Effectors are structurally and functionally diverse.<br />
• A common G-protein binding domain has not been<br />
identified among effector proteins.<br />
• Effector proteins integrate signals from multiple G<br />
protein pathways.<br />
G protein-regulated effectors include enzymes<br />
that create or destroy intra<strong>cell</strong>ular second<br />
messengers (adenylyl cyclase, cyclic GMP phosphodiesterase,<br />
phospholipase C-β, phosphatidylinositol-3-kinase),<br />
protein kinases, ion channels<br />
(K+, Ca2+) and possibly membrane transport<br />
proteins (see Figure 14.25). Effectors may be<br />
integral membrane proteins or intrinsically soluble<br />
proteins that bind G proteins at the membrane<br />
surface. No conserved G protein-binding<br />
domain or sequence motif has been identified<br />
among effector proteins, and most effectors are<br />
related to proteins that have similar functions<br />
but that are not regulated by G proteins.<br />
Sensitivity to G protein regulation, thus, evolved<br />
independently in multiple families <strong>of</strong> regulatory<br />
proteins.<br />
Because they can respond to a variety <strong>of</strong><br />
Gα and Gβγ subunits, effector proteins can integrate<br />
signals from multiple G protein pathways.<br />
The different Gα or Gβγ subunits may<br />
have opposite or synergistic effects on a given<br />
effector. For example, some <strong>of</strong> the membranebound<br />
adenylyl cyclases in mammals are stimulated<br />
by Gα s<br />
and inhibited by Gα i<br />
(see Figure<br />
14.13). Many effectors are further regulated by<br />
other allosteric ligands (e.g., lipids, calmodulin)<br />
and by phosphorylation, contributing even more<br />
to integration <strong>of</strong> information.<br />
Effectors are usually represented as multiple<br />
is<strong>of</strong>orms, and each is<strong>of</strong>orm may be regulated<br />
differently, adding to the complexity <strong>of</strong> G<br />
protein networks. For example, some is<strong>of</strong>orms<br />
<strong>of</strong> adenylyl cyclase are stimulated by Gβγ,<br />
whereas others are inhibited. All phospholipase<br />
C-βs are stimulated both by Gα q<br />
family members<br />
and by Gβγ, but the potency and maximal<br />
effect <strong>of</strong> these two inputs vary dramatically<br />
among the four PLC-β is<strong>of</strong>orms.<br />
14.22<br />
Heterotrimeric G proteins<br />
are controlled by a<br />
regulatory GTPase cycle<br />
Key concepts<br />
• Heterotrimeric G proteins are activated when the<br />
Gα subunit binds GTP.<br />
• GTP hydrolysis to GDP inactivates the G protein.<br />
• GTP hydrolysis is slow, but is accelerated by<br />
proteins called GAPs.<br />
• Receptors promote activation by allowing GDP<br />
dissociation and GTP association; spontaneous<br />
exchange is very slow.<br />
• RGS proteins and phospholipase C-βs are GAPs for<br />
G proteins.<br />
The key event in heterotrimeric G protein <strong>signaling</strong><br />
is the binding <strong>of</strong> GTP to the Gα subunit.<br />
GTP binding activates the Gα subunit, which<br />
allows both it and the Gβγ subunit to bind and<br />
regulate effectors. The Gα subunit remains active<br />
as long as GTP is bound, but Gα also has<br />
GTPase activity and hydrolyzes bound GTP to<br />
GDP. Gα-GDP is inactive. G proteins thus traverse<br />
a GTPase cycle <strong>of</strong> GTP binding/activation and hydrolysis/deactivation,<br />
as depicted in FIGURE 14.26.<br />
Therefore, the control <strong>of</strong> G protein <strong>signaling</strong> is<br />
intrinsically kinetic. The relative signal strength,<br />
or amplitude, is proportional to the fraction <strong>of</strong><br />
G protein that is in the active, GTP-bound form.<br />
This fraction equals the balance <strong>of</strong> the rates <strong>of</strong><br />
GTP binding and GTP hydrolysis, the activating<br />
618 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 619<br />
and deactivating arms <strong>of</strong> the GTPase cycle. Both<br />
limbs are highly regulated over a range <strong>of</strong> rates<br />
greater than 1000-fold.<br />
Receptors promote G protein activation by<br />
opening the nucleotide-binding site on the G<br />
protein, thus accelerating both GDP dissociation<br />
and GTP association. This process is referred to<br />
as GDP/GTP exchange catalysis. Exchange proceeds<br />
in the direction <strong>of</strong> activation because the<br />
affinity <strong>of</strong> G proteins for GTP is much higher than<br />
that for GDP and because the cytosolic concentration<br />
<strong>of</strong> GTP is about 20-fold higher than that<br />
<strong>of</strong> GDP. Spontaneous GDP/GTP exchange is very<br />
slow for most G proteins (many minutes), which<br />
maintains basal signal output at a low level. In<br />
contrast, receptor-catalyzed exchange can take<br />
place in a few tens <strong>of</strong> milliseconds, which allows<br />
rapid responses in <strong>cell</strong>s such as visual photoreceptors,<br />
other neurons, or muscle.<br />
Because receptors are not directly required<br />
for a G protein’s <strong>signaling</strong> activity, a receptor can<br />
dissociate after GDP/GTP exchange and catalyze<br />
the activation <strong>of</strong> additional G protein molecules.<br />
In this way, a single receptor may maintain the<br />
activation <strong>of</strong> multiple G proteins, providing molecular<br />
amplification <strong>of</strong> the incoming signal.<br />
Other receptors may remain bound to their G<br />
protein targets, which means that they do not<br />
act as amplifiers. However, more tightly bound<br />
receptors can initiate <strong>signaling</strong> more quickly and<br />
promote G protein reactivation when hydrolysis<br />
<strong>of</strong> bound GTP is rapid.<br />
In the absence <strong>of</strong> stimulus, Gα subunits<br />
hydrolyze bound GTP slowly. The average activation<br />
lifetime <strong>of</strong> the Gα-GTP complex is<br />
about 10-150 seconds, depending on the G<br />
protein. This rate is far slower than rates <strong>of</strong> deactivation<br />
<strong>of</strong>ten observed in <strong>cell</strong>s when an agonist<br />
is removed. For example, visual <strong>signaling</strong><br />
terminates in about 10 ms after stimulation by<br />
a photon, and many other G protein systems<br />
are almost as fast. GTP hydrolysis is accelerated<br />
by GTPase-activating proteins (GAPs),<br />
which directly bind Gα subunits. In some cases<br />
acceleration exceeds 2000-fold. Such speed is<br />
necessary in systems like vision or neurotransmission,<br />
which must respond to quickly changing<br />
stimuli. Because G protein <strong>signaling</strong> is a<br />
balance <strong>of</strong> activation and deactivation, GAPs<br />
deplete the pool <strong>of</strong> GTP-activated G protein<br />
and can thereby also act to inhibit G protein<br />
<strong>signaling</strong>. GAPs can thus inhibit <strong>signaling</strong>,<br />
quench output upon signal termination, or<br />
both. What behavior predominates depends<br />
on the GAP’s intrinsic activity and its regulation.<br />
Receptor<br />
+ agonist<br />
GDP<br />
G protein-GDP<br />
Pi<br />
The regulatory GTPase cycle<br />
Receptor<br />
- agonist<br />
G protein<br />
GAP<br />
GTP<br />
G protein-GTP<br />
*ACTIVE*<br />
Effector protein<br />
There are two families <strong>of</strong> GAPs for heterotrimeric<br />
G proteins. The RGS proteins (regulators<br />
<strong>of</strong> G protein <strong>signaling</strong>) are a family <strong>of</strong><br />
about 30 proteins, most or all <strong>of</strong> which have<br />
GAP activity and regulate G protein <strong>signaling</strong><br />
rates and amplitudes. The role <strong>of</strong> RGS proteins<br />
in terminating the G protein signal can be seen<br />
in FIGURE 14.27. Some proteins with RGS domains<br />
also act as G protein-regulated effectors.<br />
These include activators <strong>of</strong> the Rho family <strong>of</strong><br />
monomeric GTP-binding proteins (see below)<br />
and GPCR kinases, which are feedback regulators<br />
<strong>of</strong> GPCR function. The second group <strong>of</strong> G<br />
protein GAPs are phospholipase C-βs. These enzymes<br />
are effectors that are stimulated by both<br />
Gα q<br />
and by Gβγ, but they also act as G q<br />
GAPs,<br />
probably to control output kinetics.<br />
G protein-GTP-<br />
Effector protein<br />
*ACTIVE*<br />
FIGURE 14.26 G proteins are activated when GTP binds to the G subunit, such<br />
that both G-GTP and G can bind and regulate the activities <strong>of</strong> appropriate<br />
effector proteins. G subunits also have intrinsic GTPase activities, and the primary<br />
deactivating reaction is hydrolysis <strong>of</strong> bound GTP to GDP (rather than GTP<br />
dissociation). Thus, the steady-state signal output from a receptor-G protein<br />
module is the fraction <strong>of</strong> the G protein in the GTP-bound state, which reflects<br />
the balance <strong>of</strong> the activation and deactivation rates. Both GTP binding and GTP<br />
hydrolysis are intrinsically slow and highly regulated. GDP binds tightly to G,<br />
such that GDP dissociation is rate-limiting for binding <strong>of</strong> a new molecule <strong>of</strong><br />
GTP and consequent reactivation. Both GDP release and GTP binding are catalyzed<br />
by GPCRs. Hydrolysis <strong>of</strong> bound GTP is accelerated by GTPase-activating<br />
proteins (GAPs). Receptors and GAPs coordinately control both the steady-state<br />
level <strong>of</strong> signal output and the rates <strong>of</strong> activation and deactivation <strong>of</strong> the module.<br />
14.22 Heterotrimeric G proteins are controlled by a regulatory GTPase cycle 619
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 620<br />
Single photon responses <strong>of</strong> GAP-deficient mice<br />
Current (pA)<br />
0.50<br />
0.25<br />
0.00<br />
knockout<br />
heterozygous<br />
wild-type<br />
0 2 Time (s) 4<br />
Light flash<br />
FIGURE 14.27 G protein GAPs can accelerate signal termination<br />
upon removal <strong>of</strong> agonist, and <strong>of</strong>ten do not act as inhibitors<br />
during the response to receptor. The figure shows<br />
the electrical response <strong>of</strong> a mouse photoreceptor (rod) <strong>cell</strong><br />
to a single photon <strong>of</strong> light. In mice that lack RGS9, the GAP<br />
for the photoreceptor G protein G t<br />
, the signal is prolonged<br />
for many seconds because hydrolysis <strong>of</strong> GTP bound to G t<br />
is<br />
slow. In wild-type or heterozygous mice, hydrolysis takes<br />
place in about 15 milliseconds, and the decay <strong>of</strong> the signal<br />
is much faster. Note that the maximal output is similar in<br />
wild-type and mutant mice, indicating that the GAP does<br />
not act as an inhibitor in rod <strong>cell</strong>s. In humans, genetic loss<br />
<strong>of</strong> RGS9 leads to severe loss <strong>of</strong> vision that is particularly<br />
marked in bright light. Reproduced from Chen et al. Nature.<br />
2000. 403:557–560. Permission also granted by Ching-Kang<br />
Jason Chen, Virginia Commonwealth University.<br />
While the GTPase cycle described in Figure<br />
14.26 is general, it is highly simplified. Interactions<br />
among receptor, Gα, Gβγ, GAP, and effector are<br />
frequently simultaneous and <strong>of</strong>ten demonstrate<br />
complex cooperative interactions. For example,<br />
Gβγ inhibits the release <strong>of</strong> GDP (to minimize<br />
spontaneous activation), promotes the exchange<br />
catalyst activity <strong>of</strong> the receptor, inhibits GAP activity,<br />
and helps initiate receptor phosphorylation<br />
that leads to desensitization. The other<br />
components can be nearly this multifunctional.<br />
In addition, inputs from other proteins can alter<br />
the dynamics <strong>of</strong> the GTPase cycle at several points.<br />
The core G protein module is, thus, functionally<br />
versatile as a signal processor in addition to being<br />
versatile in the scope <strong>of</strong> its targets.<br />
14.23<br />
Small, monomeric GTPbinding<br />
proteins are<br />
multiuse switches<br />
Key concepts<br />
• Small GTP-binding proteins are active when bound<br />
to GTP and inactive when bound to GDP.<br />
• GDP/GTP exchange catalysts known as GEFs<br />
(guanine nucleotide exchange factors) promote<br />
activation.<br />
• GAPs accelerate hydrolysis and deactivation.<br />
• GDP dissociation inhibitors (GDIs) slow<br />
spontaneous nucleotide exchange.<br />
Monomeric GTP-binding proteins, which are<br />
encoded by about 150 genes in animals, modulate<br />
a wide variety <strong>of</strong> <strong>cell</strong>ular processes including<br />
signal transduction, organellar trafficking,<br />
intra-organellar transport, cytoskeletal assembly,<br />
and morphogenesis. The small GTP-binding<br />
proteins that most clearly function in signal<br />
transduction are the Ras and Ras-related proteins<br />
(Ral, Rap) and the Rho/Rac/Cdc42 proteins,<br />
about 10-15 in all. They are usually about<br />
20-25 kDa in size and are homologous to the<br />
GTP-binding domains <strong>of</strong> Gα subunits.<br />
The regulatory activities <strong>of</strong> the small GTPbinding<br />
proteins are controlled by a GTP binding<br />
and hydrolysis cycle like that <strong>of</strong> the<br />
heterotrimeric G proteins, with similar regulatory<br />
inputs. They are activated by GTP, and hydrolysis<br />
<strong>of</strong> bound GTP to GDP terminates<br />
activation. GDP/GTP exchange catalysts, known<br />
as GEFs (guanine nucleotide exchange factors,<br />
functionally analogous to GPCRs) promote activation,<br />
and GAPs accelerate hydrolysis and<br />
consequent deactivation. In addition, GDP dissociation<br />
inhibitors (GDIs) slow spontaneous<br />
nucleotide exchange and activation to dampen<br />
basal activity, an activity shared by Gβγ subunits<br />
for the heterotrimeric G proteins.<br />
While the underlying biochemical regulatory<br />
events are essentially identical for<br />
monomeric and heterotrimeric G proteins,<br />
monomeric G proteins use the basic GTPase cycle<br />
in additional ways. Signal output by heterotrimeric<br />
G proteins and many monomeric<br />
G proteins is usually thought to reflect a balance<br />
<strong>of</strong> their active (GTP-bound) and inactive<br />
(GDP-bound) states in a rapidly turning-over<br />
GTPase cycle. GEFs favor formation <strong>of</strong> more active<br />
G protein, and GAPs favor the inactive state.<br />
In contrast, probably an equal number <strong>of</strong> the<br />
monomeric G proteins behave as acute on-<strong>of</strong>f<br />
switches. Upon binding GTP, they initiate a<br />
process (regulation, recruitment <strong>of</strong> other pro-<br />
620 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 621<br />
teins). They then maintain this activity, sometimes<br />
for many seconds or minutes, until they<br />
are acted upon by a GAP. For example, the<br />
monomeric G protein Ran regulates nucleocytoplasmic<br />
trafficking <strong>of</strong> protein and RNA in both<br />
directions, cooperating with carrier proteins<br />
known as karyopherins (see 5.15 The Ran GTPase<br />
controls the direction <strong>of</strong> nuclear transport). In the<br />
nucleus, high Ran GEF activity promotes GTP<br />
binding. Nuclear Ran-GTP then binds import<br />
karyopherins to drive dissociation <strong>of</strong> newly arrived<br />
cargo and promote return <strong>of</strong> the karyopherin<br />
to the cytoplasm. It also binds export<br />
karyopherins to permit binding <strong>of</strong> outgoing<br />
cargo. Outside the nucleus, high Ran GAP activity<br />
promotes GTP hydrolysis. Cytoplasmic<br />
Ran-GDP dissociates from both the export karyopherins<br />
to allow dissociation <strong>of</strong> outgoing cargo<br />
and from the import karyopherins to allow them<br />
to bind cargo for import. Thus, for monomeric<br />
G proteins such as Ran, each phase <strong>of</strong> the GTPase<br />
cycle determines a specific, coupled step in a<br />
parallel regulatory cycle.<br />
A second major difference between the<br />
monomeric and the heterotrimeric G proteins<br />
is the structures <strong>of</strong> the GEFs, GAPs, and GDIs.<br />
Both GEFs and GAPs for monomeric GTP-binding<br />
proteins are structurally heterogeneous (although<br />
some clearly related families are<br />
evident). In addition, mechanisms for regulating<br />
these GEFs and GAPs are equally diverse.<br />
They include phosphorylation by protein kinases;<br />
allosteric regulation by heterotrimeric<br />
and/or monomeric G proteins, by second messengers<br />
and by other regulatory proteins; sub<strong>cell</strong>ular<br />
sequestration or recruitment to scaffolds;<br />
and assorted other mechanisms.<br />
The Ras proteins were the first small GTPbinding<br />
proteins to be discovered. They were<br />
identified as oncogene products because they<br />
cause malignant growth if they are either overexpressed<br />
or persistently activated by mutation;<br />
they are among the most commonly mutated<br />
genes in human tumors. Several viral ras genes<br />
figure prominently as oncogenes.<br />
Mammalian <strong>cell</strong>s contain three ras genes (H,<br />
N, and K). They may share inputs and outputs<br />
to varying extents, and they can compensate for<br />
each other in some genetic screens. It has been<br />
difficult to assign unique functions to the individual<br />
Ras proteins. Inputs to the Ras proteins<br />
are diverse and speak to the importance <strong>of</strong> Ras<br />
proteins as a crucial node in <strong>signaling</strong>.<br />
Ras GEFs and GAPS are regulated by both<br />
receptor and nonreceptor Tyr kinases through<br />
direct phosphorylation and by recruitment <strong>of</strong><br />
the regulators to the plasma membrane. Other<br />
cytoplasmic serine/threonine kinases also converge<br />
on Ras activation. Rap1, another member<br />
<strong>of</strong> the Ras family, may also fit directly into<br />
this network because it is suspected <strong>of</strong> competing<br />
with Ras proteins for protein kinase targets;<br />
in vivo it can suppress the oncogenic activity <strong>of</strong><br />
Ras. Rap1 is regulated independently, however,<br />
and acts on independent <strong>signaling</strong> pathways as<br />
well. One <strong>of</strong> its GAPs is stimulated by the G i<br />
class <strong>of</strong> G proteins, for example, and its several<br />
GEFs are stimulated by Ca2+, diacylglycerol, and<br />
cAMP.<br />
Ras proteins generally regulate <strong>cell</strong> growth,<br />
proliferation, and differentiation by modulating<br />
the activities <strong>of</strong> multiple effector proteins.<br />
The best known and best studied Ras effector is<br />
the protein kinase Raf, which initiates a MAPK<br />
cascade. FIGURE 14.28 shows well established Ras<br />
effectors.<br />
Rho, Rac, and Cdc42 are related monomeric<br />
GTP-binding proteins that are involved in generating<br />
signals that affect <strong>cell</strong> morphology. Each<br />
class <strong>of</strong> proteins regulates its own array <strong>of</strong> effectors<br />
and is controlled by separate groups <strong>of</strong> GEFs,<br />
GAPs, and GDIs. Effectors regulated by this family<br />
include phospholipases C and D, multiple<br />
protein and lipid kinases, proteins that nucleate<br />
or reorganize actin filaments, and components<br />
<strong>of</strong> the neutrophil oxygen activating<br />
system, among others (see 8.14 Small G proteins<br />
regulate actin polymerization).<br />
14.24<br />
Ras has three main effectors<br />
Function<br />
Effector Target<br />
Protein kinase cascade Raf MAPK<br />
Lipid kinase PI 3-kinase Akt<br />
Exchange factor RalGDS Exocyst<br />
Protein phosphorylation/<br />
dephosphorylation is a<br />
major regulatory<br />
mechanism in the <strong>cell</strong><br />
Key concepts<br />
• Protein kinases are a large protein family.<br />
• Protein kinases phosphorylate Ser and Thr, or Tyr,<br />
or all three.<br />
• Protein kinases may recognize the primary<br />
sequence surrounding the phosphorylation site.<br />
• Protein kinases may preferentially recognize<br />
phosphorylation sites within folded domains.<br />
FIGURE 14.28 Ras-GTP binds<br />
to many proteins. Three well<br />
established effectors include<br />
Raf, PI 3-kinase, and RalGDS.<br />
Activation <strong>of</strong> these effectors<br />
activates a MAPK pathway, increases<br />
PI 3-kinase activity,<br />
and promotes assembly <strong>of</strong> a<br />
protein complex involved in<br />
exocytosis <strong>of</strong> secretory vesicles.<br />
14.24 Protein phosphorylation/dephosphorylation is a major regulatory mechanism in the <strong>cell</strong> 621
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 622<br />
SUBSTRATE 1<br />
(Protein SER)<br />
H<br />
OH<br />
PRODUCT 1<br />
(Phosphorylated protein)<br />
H<br />
N<br />
C<br />
C<br />
N<br />
C<br />
N<br />
C<br />
C<br />
N<br />
C<br />
H<br />
O<br />
CH 2<br />
H<br />
O<br />
H<br />
O<br />
CH 2<br />
H<br />
O<br />
Protein kinases are two substrate enzymes<br />
O - P O -<br />
O<br />
+<br />
O - P O<br />
O<br />
SUBSTRATE<br />
(Mg 2+. 2<br />
ATP)<br />
Mg 2+<br />
O - P O<br />
O<br />
O - O -<br />
O - P O<br />
O<br />
O - P O<br />
O<br />
CH 2<br />
(Mg 2+. Triphosphate)<br />
Ribose<br />
PROTEIN KINASE<br />
PRODUCT<br />
(Mg 2+. 2<br />
ADP)<br />
Mg 2+<br />
O - P O<br />
O<br />
CH 2<br />
Adenine<br />
Adenine<br />
O<br />
(Mg 2+. Diphosphate)<br />
Rib<br />
FIGURE 14.29 Protein kinases transfer the -phosphoryl group from ATP to<br />
serine, threonine, or tyrosine residues in protein substrates.<br />
Protein phosphorylation is the most common<br />
form <strong>of</strong> regulatory posttranslational modification.<br />
It occurs in all organisms, and it is estimated<br />
that about one-third <strong>of</strong> proteins in animals are<br />
at some time phosphorylated. Phosphorylation<br />
can stimulate or inhibit the catalytic activity <strong>of</strong><br />
an enzyme, the affinity with which a protein<br />
binds other molecules, its sub<strong>cell</strong>ular localization,<br />
its ability to be further covalently modified,<br />
or its stability. Single phosphorylations may cause<br />
500-fold or greater changes in activity, and proteins<br />
are <strong>of</strong>ten phosphorylated on multiple<br />
residues in complex and interacting patterns.<br />
Most protein phosphorylation in eukaryotes,<br />
and essentially all in animals, is catalyzed<br />
by protein kinases; dephosphorylation is catalyzed<br />
by phosphoprotein phosphatases. Both<br />
classes <strong>of</strong> enzymes are controlled by diverse<br />
mechanisms. In addition, proteins are <strong>of</strong>ten<br />
phosphorylated by multiple protein kinases, resulting<br />
in the generation <strong>of</strong> a range <strong>of</strong> activity<br />
states. This complexity allows inputs from different<br />
<strong>signaling</strong> pathways to be integrated into<br />
the resulting activity <strong>of</strong> the target.<br />
In bacteria, plants, and fungi, an additional<br />
protein phosphorylating system known as twocomponent<br />
<strong>signaling</strong> is vital. The protein kinases<br />
involved in two-component <strong>signaling</strong> are<br />
unrelated to the eukaryotic protein kinase superfamily<br />
and phosphorylate aspartate residues<br />
rather than serine, threonine, or tyrosine.<br />
Protein kinases transfer a phosphoryl group<br />
from ATP to Ser, Thr, and Tyr residues <strong>of</strong> protein<br />
substrates to form chemically stable phosphate<br />
esters, as shown in FIGURE 14.29. In<br />
animals, the distribution <strong>of</strong> phosphate among<br />
these three amino acid residues is uneven:<br />
~90%-95% is on Ser, 5%-8% on Thr, and less<br />
than 1% on Tyr residues. The human genome<br />
contains approximately 500 genes that encode<br />
protein kinases, and many protein kinase<br />
mRNAs undergo alternative splicing. This makes<br />
the protein kinase gene superfamily one <strong>of</strong> the<br />
largest functional gene groups. The number and<br />
diversity <strong>of</strong> these enzymes emphasize the great<br />
and varied uses <strong>of</strong> protein kinases to regulate <strong>cell</strong>ular<br />
functions. Although some protein kinases<br />
have a limited tissue and/or developmental distribution,<br />
many are ubiquitously expressed.<br />
Protein kinases are grouped according to<br />
their residue specificity. Protein kinases that<br />
phosphorylate Ser will usually also recognize<br />
Thr, hence the name protein Ser/Thr kinase.<br />
Multi<strong>cell</strong>ular organisms have protein Tyr kinases,<br />
which only recognize Tyr. Dual specificity<br />
protein kinases can phosphorylate Ser,<br />
Thr, and Tyr in the appropriately restricted substrate<br />
conformational context and are generally<br />
the most selective <strong>of</strong> the protein kinases.<br />
The analysis <strong>of</strong> the kinomes <strong>of</strong> several organisms<br />
has led to a more elaborate grouping<br />
derived from sequence relationships, shown in<br />
FIGURE 14.30, that also reflects to some extent<br />
on regulatory mechanisms and substrate specificity.<br />
For example, the AGC group is named<br />
for its founding members, cAMP-dependent<br />
protein kinase (PKA), cyclic GMP-dependent<br />
protein kinase (PKG), Ca2+, and phospholipiddependent<br />
protein kinase (PKC). These protein<br />
kinases are regulated by second messengers and<br />
prefer substrates that contain basic residues near<br />
the phosphorylation site.<br />
In addition to substrate specificity for amino<br />
acid residues, most protein kinases are also selective<br />
for local sequence surrounding the substrate<br />
site. Screening strategies have resulted in methods<br />
to predict if proteins contain consensus substrate<br />
sites for a wide variety <strong>of</strong> protein kinases.<br />
Antibodies can be used to identify and roughly<br />
quantitate protein phosphorylation at specific<br />
sites in proteins. Beyond local recognition, protein<br />
kinases may display marked substrate selectivity<br />
among similar proteins based on overall<br />
622 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 623<br />
Human kinome tree<br />
FIGURE 14.30 The protein kinases in<br />
the human genome can be grouped according<br />
to sequence relationships that<br />
reveal seven major branches. The tyrosine<br />
kinases are contained within one<br />
major branch. The others are Ser/Thrspecific<br />
or dual specificity, and are named<br />
for the best described members: AGC<br />
from PKA, PKG, and PKC; CAMK from the<br />
calcium, calmodulin-dependent kinases;<br />
CMGC from CDKs, MAPKs, GSK3, Clks; CK1<br />
from casein kinase 1; STE from Ste20,<br />
Ste11, and Ste7, the MAP4K, MAP3K,<br />
and MAP2K in the yeast mating pathway;<br />
and TKL, the Tyr kinase-like enzymes.<br />
Reproduced with permission from<br />
G. Manning, et al. 2002. Science. 298:<br />
1912-1934. © 2002 AAAS. Photo courtesy<br />
<strong>of</strong> Gerard Manning, Salk Institute,<br />
and reprinted with permission <strong>of</strong> Cell<br />
Signaling Technology, Inc. (www.<strong>cell</strong>signal.com).<br />
three-dimensional structure, for example, or<br />
among proteins that have been differentially covalently<br />
modified by phosphorylation or ubiquitination.<br />
In animal <strong>cell</strong>s, some protein kinases are<br />
hormone receptors that span the plasma membrane.<br />
Some protein kinase receptors are protein<br />
serine/threonine kinases, such as the<br />
transforming growth factor- (TGF-)receptor,<br />
but the majority are protein tyrosine kinases, including<br />
receptors for insulin, epidermal growth<br />
factor (EGF), platelet-derived growth factor<br />
(PDGF), and other regulators <strong>of</strong> <strong>cell</strong> growth and<br />
differentiation. Other protein kinases are intrinsically<br />
soluble intra<strong>cell</strong>ular enzymes, although<br />
they may bind to one or more organellar<br />
membranes.<br />
X-ray crystallographic structures <strong>of</strong> protein<br />
kinases have revealed a wealth <strong>of</strong> information<br />
about their mechanism <strong>of</strong> activation. The conserved<br />
minimum catalytic core <strong>of</strong> a protein kinase<br />
contains about 270 amino acids, yielding<br />
a minimum molecular mass <strong>of</strong> about 30,000<br />
Da. Within this core, there are two folded domains<br />
that form the active site at their interface,<br />
as shown in FIGURE 14.31. One or both <strong>of</strong><br />
the conserved lysine (Lys) or aspartate (Asp)<br />
residues that are required for phosphoryl transfer<br />
are frequently mutated to disrupt kinase activity.<br />
A sequence near the active site, referred<br />
to as the activation loop, <strong>of</strong>ten undergoes a conformational<br />
rearrangement to generate active<br />
forms <strong>of</strong> the protein kinases and is the most<br />
common site <strong>of</strong> regulatory phosphorylation in<br />
14.24 Protein phosphorylation/dephosphorylation is a major regulatory mechanism in the <strong>cell</strong> 623
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 624<br />
ERK2 inactive and active conformations<br />
INACTIVE (ERK2)<br />
N terminal<br />
domain<br />
Tyr185<br />
Thr183<br />
C terminal<br />
domain<br />
ACTIVE (ERK2-P2)<br />
Tyr185<br />
Thr183<br />
FIGURE 14.31 The structures <strong>of</strong> unphosphorylated, inactive MAPK ERK2 and<br />
phosphorylated, active ERK2 are compared. ERK2 has a typical protein kinase<br />
structure. The smaller N-terminal domain is composed primarily <strong>of</strong> strands<br />
and the larger C-terminal domain is primarily -helical. The active site is formed<br />
at the interface <strong>of</strong> the two domains. The activation loop emerges from the active<br />
site and is refolded following phosphorylation <strong>of</strong> the Tyr and Thr residues,<br />
inducing the repositioning <strong>of</strong> active site residues. ATP (not shown) binds in<br />
the interior <strong>of</strong> the active site; productive binding <strong>of</strong> protein substrates to the<br />
surface <strong>of</strong> the C-terminal domain is also facilitated by the reorganization <strong>of</strong><br />
the activation loop. Structures generated from Protein Data Bank files 1ERK<br />
and 2ERK.<br />
Sensor/<br />
Histidine<br />
kinase<br />
Response<br />
regulator<br />
His<br />
Asp<br />
Ligand<br />
P<br />
ATP<br />
His phosphorylation<br />
Two-component <strong>signaling</strong> systems<br />
ADP<br />
His<br />
Asp<br />
P<br />
P<br />
Transfer <strong>of</strong> phosphate<br />
to Asp: response<br />
regulator active<br />
His<br />
Asp<br />
P<br />
P<br />
H 2 O<br />
Response regulator<br />
deactivated<br />
FIGURE 14.32 The basic two-component system is composed <strong>of</strong> a signal-activated<br />
histidine kinase, referred to as a sensor, and an effector protein, the response<br />
regulator, that is activated when it is phosphorylated on an aspartate<br />
residue by the sensor. The activity <strong>of</strong> the response regulator is terminated when<br />
the aspartyl-phosphate is hydrolyzed.<br />
the protein kinase family. There are unique inserts<br />
on the surface <strong>of</strong> protein kinases that generate<br />
specificity in localization, interaction with<br />
other regulatory molecules, and recognition <strong>of</strong><br />
substrates. These landmarks allow both classification<br />
and genetic manipulation <strong>of</strong> protein kinases.<br />
Protein kinases have evolved numerous<br />
and diverse regulatory mechanisms to complement<br />
their number and multiple functions.<br />
These mechanisms include allosteric activation<br />
and inhibition by lipids, soluble small molecules<br />
and other proteins; activating and inhibitory<br />
phosphorylation and other covalent modifications,<br />
including proteolysis; and binding to scaffolds<br />
and adaptors to enhance activity or limit<br />
nonspecific activities. Many such inputs may<br />
regulate a single protein kinase in a complex<br />
combinatoric code. Further, multiple protein<br />
kinases that act sequentially, such as in a protein<br />
kinase cascade (see Figure 14.38), can create<br />
uniquely complex <strong>signaling</strong> patterns.<br />
14.25<br />
Two-component protein<br />
phosphorylation systems<br />
are <strong>signaling</strong> relays<br />
Key concepts<br />
• Two-component <strong>signaling</strong> systems are composed <strong>of</strong><br />
sensor and response regulator components.<br />
• Upon receiving a stimulus, sensor components<br />
undergo autophosphorylation on a histidine (His)<br />
residue.<br />
• Transfer <strong>of</strong> the phosphate to an aspartyl residue on<br />
the response regulator serves to activate the<br />
regulator.<br />
Prokaryotes, plants, and fungi share an alternative<br />
mechanism for regulatory phosphorylation<br />
and dephosphorylation known as two-component<br />
<strong>signaling</strong>. FIGURE 14.32 shows a typical twocomponent<br />
system. In this system, the receptor,<br />
referred to as a sensor, responds to a stimulus by<br />
catalyzing its own phosphorylation on a His<br />
residue. Sensors include chemoattractant receptors<br />
in bacteria, a regulator <strong>of</strong> osmolarity in fungi,<br />
light-sensitive proteins, the receptor for the plantripening<br />
hormone ethylene, and other receptors<br />
for diverse environmental, hormonal, and metabolic<br />
signals. The mammalian mitochondrial dehydrogenase<br />
kinases are related in sequence to<br />
the bacterial histidine kinases, although the mammalian<br />
enzymes phosphorylate serine or threonine<br />
residues, not histidine. The phosphorylated<br />
sensor next transfers its covalently bound phos-<br />
624 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 625<br />
phate to an aspartyl residue on a second protein<br />
known as a response regulator. Response regulators<br />
initiate <strong>cell</strong>ular responses, usually by binding<br />
to other cytoplasmic proteins and allosterically<br />
regulating their activities.<br />
Although all two-component systems follow<br />
this same general pattern, their structures<br />
and precise reaction pathways vary enormously.<br />
Some two-component systems are composed<br />
<strong>of</strong> only one protein (sensor and response regulator<br />
in a single polypeptide chain). Others are<br />
composed <strong>of</strong> a sensor protein and two aspartylphosphorylated<br />
proteins, in which the first or<br />
the second may display response regulatory activity.<br />
Finally, two-component systems usually<br />
lack conventional protein phosphatases.<br />
Hydrolysis <strong>of</strong> the aspartyl-phosphate bond may<br />
be spontaneous or regulated by the response<br />
regulator itself.<br />
14.26<br />
Pharmacological<br />
inhibitors <strong>of</strong> protein<br />
kinases may be used to<br />
understand and treat<br />
disease<br />
Key concepts<br />
• Protein kinase inhibitors are useful both for<br />
<strong>signaling</strong> research and as drugs.<br />
• Protein kinase inhibitors usually bind in the ATP<br />
binding site.<br />
Many inhibitors have been developed for basic<br />
research purposes to explore the functions <strong>of</strong><br />
protein kinases. The importance <strong>of</strong> these enzymes<br />
in disease processes has also made them<br />
targets <strong>of</strong> drug screening projects yielding inhibitors<br />
for many protein kinases. The majority<br />
<strong>of</strong> pharmacological inhibitors <strong>of</strong> protein<br />
kinases compete with ATP binding. Because <strong>of</strong><br />
the huge number <strong>of</strong> ATP-binding proteins in a<br />
<strong>cell</strong>, there are inevitable concerns about inhibitor<br />
specificity not only with respect to the<br />
other protein kinases but also to the other proteins<br />
that bind nucleotides. This problem has<br />
been mitigated with variable success through<br />
chemical library screening, structure-based modification<br />
<strong>of</strong> lead compounds, and inhibitor testing<br />
against panels <strong>of</strong> protein kinases.<br />
Many inhibitors with actions on PKA or<br />
PKCs, for example, have effects on several other<br />
members <strong>of</strong> the AGC family. Although pharmacological<br />
inhibitors with effects on PKA<br />
abound, the most selective are derived from the<br />
naturally occurring small inhibitory protein<br />
known as PKI or the Walsh inhibitor. In vitro<br />
and <strong>cell</strong>-based screens have identified much<br />
more selective inhibitors for MAP2Ks in the<br />
ERK1/2 pathway. These inhibitors have fewer<br />
known protein kinase cross reactivities, probably<br />
due to the fact that they do not bind in the<br />
ATP site. Among inhibitors that have progressed<br />
in the clinic, compounds developed against the<br />
EGF receptor and certain other protein tyrosine<br />
kinases have had considerable success.<br />
14.27<br />
Phosphoprotein<br />
phosphatases reverse the<br />
actions <strong>of</strong> kinases and are<br />
independently regulated<br />
Key concepts<br />
• Phosphoprotein phosphatases reverse the actions<br />
<strong>of</strong> protein kinases.<br />
• Phosphoprotein phosphatases may<br />
dephosphorylate phosphoserine/threonine,<br />
phosphotyrosine, or all three.<br />
• Phosphoprotein phosphatase specificity is <strong>of</strong>ten<br />
achieved through the formation <strong>of</strong> specific protein<br />
complexes.<br />
Protein phosphorylation is reversed by phosphoprotein<br />
phosphatases. These enzymes display distinct<br />
specificities and modes <strong>of</strong> regulation.<br />
Phosphoprotein phosphatases can be considered<br />
in two broad groups based on their specificity and<br />
sequence relationships: protein-serine/threonine<br />
phosphatases and protein-tyrosine phosphatases.<br />
Most protein-serine/threonine phosphatases<br />
are regulated by association with other proteins.<br />
Targeted localization is the major determinant <strong>of</strong><br />
substrate specificity. Phosphoprotein phosphatase<br />
1 (PP1) associates with a variety <strong>of</strong> regulatory<br />
subunits that specifically direct it to relevant organelles.<br />
One subunit (known as the G subunit),<br />
for example, specifies association with glycogen<br />
particles. The interaction with this subunit is itself<br />
regulated by phosphorylation. Small protein<br />
inhibitors can suppress PP1 activity.<br />
Phosphoprotein phosphatase 2A (PP2A) is<br />
composed <strong>of</strong> a catalytic subunit, a scaffolding<br />
subunit, and one <strong>of</strong> a large number <strong>of</strong> regulatory<br />
subunits. The regulatory subunit modulates activity<br />
and localization <strong>of</strong> the phosphatase. Some<br />
viruses alter the behavior <strong>of</strong> the <strong>cell</strong>s they infect<br />
by interfering with phosphatase activity. For example,<br />
<strong>cell</strong>s transformed with the SV40 virus<br />
express a viral protein known as small t anti-<br />
14.27 Phosphoprotein phosphatases reverse the actions <strong>of</strong> kinases and are independently regulated 625
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 626<br />
gen. Small t displaces the regulatory subunit<br />
from PP2A and alters the activity and the sub<strong>cell</strong>ular<br />
localization <strong>of</strong> the phosphatase. In addition,<br />
natural toxins such as okadaic acid,<br />
calyculin, and microcystin inhibit PP2A and PP1<br />
to varying extents both in vitro and in intact <strong>cell</strong>s.<br />
Another major protein-serine/threonine<br />
phosphatase, called calcineurin (also known as<br />
phosphoprotein phosphatase 2B), is regulated<br />
by Ca2+-calmodulin (see 14.15 Ca2+ <strong>signaling</strong><br />
serves diverse purposes in all eukaryotic <strong>cell</strong>s) and<br />
plays essential roles in cardiac development and<br />
T <strong>cell</strong> activation, among other events. The major<br />
mechanism <strong>of</strong> action <strong>of</strong> the immunosuppressants<br />
cyclosporin and FK506 is to inhibit<br />
calcineurin.<br />
The protein tyrosine phosphatases (PTPs)<br />
are cysteine-dependent enzymes that utilize a<br />
conserved Cys-Xaa-Arg motif to hydrolyze phosphoester<br />
bonds in their substrates. The PTPs are<br />
encoded by over 100 genes in humans and are<br />
classified in four subfamilies: the phosphotyrosine-specific<br />
phosphatases, the Cdc25 phosphatases,<br />
the dual specificity phosphatases<br />
(DSPs), and the low molecular weight phosphatases.<br />
Thirty-eight <strong>of</strong> the PTPs are highly selective<br />
for phosphotyrosine residues within substrates.<br />
Some <strong>of</strong> the phosphotyrosine-selective<br />
phosphatases are transmembrane proteins,<br />
whereas others are membrane associated. The<br />
most obvious function <strong>of</strong> the PTPs is to reverse<br />
the functions <strong>of</strong> tyrosine kinases; however, some<br />
have primary functions in transducing tyrosine<br />
kinase signals. For example, the protein tyrosine<br />
phosphatase SHP2 (also known as SHPTP2),<br />
binds to certain tyrosine kinase receptors<br />
through its SH2 domain and is itself tyrosine<br />
phosphorylated, thereby creating a binding site<br />
for the SH2 domain-containing adaptor protein,<br />
Grb2, which leads to activation <strong>of</strong> Ras (see<br />
14.32 MAPKs are central to many <strong>signaling</strong> pathways).<br />
The Cdc25 phosphatases recognize cyclindependent<br />
kinase (CDK) family members as<br />
substrates and play a critical role in increasing<br />
CDK activity at key junctures <strong>of</strong> the <strong>cell</strong> cycle<br />
(see Figure 14.39 and 11.4 The <strong>cell</strong> cycle is a cycle<br />
<strong>of</strong> CDK function). Similar to the dual specificity<br />
kinases, the dual specificity phosphatases are<br />
specific for a restricted number <strong>of</strong> substrates. A<br />
number <strong>of</strong> DSPs dephosphorylate MAPKs; these<br />
DSPs are called MAP kinase phosphatases, or<br />
MKPs. Several <strong>of</strong> these have been implicated<br />
in MAPK nuclear entry and exit. Some MKPs<br />
are encoded by early response genes, whose<br />
products are active near the initiation <strong>of</strong> the <strong>cell</strong><br />
cycle (see 11.7 Entry into <strong>cell</strong> cycle and S phase is<br />
tightly regulated).<br />
Substrates <strong>of</strong> other PTP family members,<br />
such as the tumor suppressor PTEN, include<br />
phosphoinositides, which are phosphorylated<br />
derivatives <strong>of</strong> the glycerolipid phosphatidylinositol<br />
that serve as second messengers<br />
(see 14.16 Lipids and lipid-derived compounds are<br />
<strong>signaling</strong> molecules). Removal <strong>of</strong> the phosphate<br />
group inactivates the second messenger. It remains<br />
unclear whether members <strong>of</strong> this group<br />
work exclusively on phophoinositides or also<br />
on protein tyrosine phosphate.<br />
14.28<br />
Covalent modification by<br />
ubiquitin and ubiquitinlike<br />
proteins is another<br />
way <strong>of</strong> regulating protein<br />
function<br />
Key concepts<br />
• Ubiquitin and related small proteins, may be<br />
covalently attached to other proteins as a<br />
targeting signal.<br />
• Ubiquitin is recognized by diverse ubiquitin<br />
binding proteins.<br />
• Ubiquitination can cooperate with other covalent<br />
modifications.<br />
• Ubiquitination regulates <strong>signaling</strong> in addition to<br />
its role in protein degradation.<br />
An important mechanism for control <strong>of</strong> protein<br />
function is through covalent modification with<br />
small proteins <strong>of</strong> the ubiquitin family. Ubiquitin<br />
is one <strong>of</strong> a family <strong>of</strong> proteins referred to as ubiquitin-like<br />
(Ubl) proteins. Ubiquitin itself is highly<br />
conserved among species, suggesting the functional<br />
importance <strong>of</strong> all <strong>of</strong> its 76 residues. In addition<br />
to the long-established role <strong>of</strong> ubiquitin<br />
in initiating protein degradation, ubiquitin modification<br />
also has a variety <strong>of</strong> functions in signal<br />
transduction.<br />
Ubl proteins are conjugated to the substrate<br />
protein by an isopeptide bond between an amino<br />
group on the substrate, usually from a Lys side<br />
chain, and the C-terminal Gly residue <strong>of</strong> the<br />
processed Ubl protein. E1, E2, and E3 proteins<br />
are required to catalyze conjugation to Ubl proteins<br />
(see Biochem 4.3 Ubiquitin attachment to substrates<br />
requires multiple enzymes). Several Ubl<br />
proteins may be attached to one substrate, <strong>of</strong>ten<br />
by serial formation <strong>of</strong> a polyubiquitin chain.<br />
Mono- and polyubiquitination both change the<br />
626 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 627<br />
protein’s behavior to induce downstream signals.<br />
Monoubiquitination is a significant regulatory<br />
modification in vesicular trafficking and DNA repair.<br />
For example, the monoubiquitinated form<br />
<strong>of</strong> the FANCD2 protein becomes associated with<br />
the repair protein BRCA1 at sites <strong>of</strong> DNA repair.<br />
Modification by the Ubl protein SUMO has<br />
roles in nuclear transport, transcription, and<br />
<strong>cell</strong> cycle progression.<br />
Polyubiquitin chains are formed when the<br />
Lys residues <strong>of</strong> ubiquitin itself, particularly K48<br />
and K63, are ubiquitinated. Addition <strong>of</strong> polyubiquitin<br />
with a K48 linkage generally directs proteins<br />
to the proteasome for degradation, whereas<br />
conjugation to polyubiquitin chains with a K63<br />
linkage promotes signal transmission, not proteolysis.<br />
Protein-bound ubiquitin is recognized by<br />
a variety <strong>of</strong> ubiquitin binding domains, including<br />
UIM (ubiquitin-interacting motif), UBA (ubiquitin<br />
association), and certain zinc finger domains.<br />
Such domains have the capacity to act as receptors<br />
for ubiquitin within modified proteins.<br />
Activation <strong>of</strong> the transcription factor NF-κB<br />
occurs by a mechanism dependent on modification<br />
both by the addition <strong>of</strong> Ubl proteins and phosphorylation.<br />
This fascinating example <strong>of</strong> regulation<br />
by ubiquitin is depicted in FIGURE 14.33. Prior to<br />
stimulation, NF-κB is retained in the cytoplasm<br />
in an inactive form by binding to its inhibitor, IκB.<br />
Phosphorylation <strong>of</strong> IκB by the IκB kinase (IKK)<br />
complex promotes its recognition by a multisubunit<br />
E3 ligase, which directs its ubiquitination<br />
and subsequent proteasomal degradation.<br />
Destruction <strong>of</strong> IκB allows NF-κB to move to the<br />
nucleus to mediate changes in transcription.<br />
IκB can be stabilized in response to certain<br />
signals through covalent attachment <strong>of</strong> the Ubl,<br />
SUMO. Sumoylation occurs on the same Lys<br />
residues that must be conjugated to ubiquitin<br />
to achieve IκB degradation. Thus, SUMO attachment<br />
stabilizes IκB and attenuates NF-κB action.<br />
This is one <strong>of</strong> numerous examples <strong>of</strong><br />
crosstalk between Ubl conjugates.<br />
A key regulatory event in NF-κB <strong>signaling</strong><br />
is activation <strong>of</strong> the IKK complex. IKK is itself<br />
regulated by ubiquitination and phosphorylation.<br />
The cytokine interleukin-1β (IL-1) causes<br />
association <strong>of</strong> adaptor proteins with its receptor<br />
to create a receptor activation complex. The<br />
interleukin-1β receptor activation complex recruits<br />
another adaptor complex containing<br />
TRAF6. A phosphorylation event releases a<br />
TRAF6 complex from the receptor activation<br />
complex into the cytoplasm.<br />
TRAF6 contains a RING domain, and is an<br />
E3 ubiquitin ligase that catalyzes formation <strong>of</strong><br />
Modification with Ubl proteins plays multiple roles in IL-1β <strong>signaling</strong><br />
TRAF6<br />
TRAF6<br />
TRAF6<br />
IKK<br />
IL-1β<br />
TRAF6<br />
TRAF6<br />
K63 ubiquitination<br />
TRAF6<br />
Nemo<br />
TAB2<br />
IKK<br />
TAK1<br />
Complex formation<br />
ATP<br />
ADP<br />
CYTOPLASM<br />
ADP<br />
ATP<br />
K63 polyubiquitin chains on the protein kinase<br />
TAK1. Polyubiquitinated TAK1 can then recruit<br />
TAB2 and TAB3, which are adaptor proteins<br />
with conserved zinc finger domains. These particular<br />
zinc finger domains bind to polyubiquitinated<br />
TAK1 and enhance its activity. TAK1,<br />
thus activated, phosphorylates and activates<br />
IKK, which then phosphorylates IκB, targeting<br />
it for degradation. Thus, ubiquitin-binding domains,<br />
such as the TAB2 and TAB3 zinc fingers,<br />
may selectively recognize K63 polyubiquitin<br />
chains to promote signal transmission.<br />
Naturally occurring small molecules may<br />
control ubiquitin ligase activity directly. Auxin<br />
(indole 3-acetic acid) is a plant hormone that regulates<br />
development by promoting the transcription<br />
<strong>of</strong> a large number <strong>of</strong> genes. Rather than<br />
K48 ubiquitination<br />
Degradation<br />
NUCLEUS<br />
DNA<br />
FIGURE 14.33 Activation <strong>of</strong> NF-B involves steps dependent on the interaction <strong>of</strong> proteins<br />
attached to ubiquitin through ubiquitin-binding proteins, competition by sumoylation,<br />
phosphorylation, and ubiquitin-mediated protein degradation.<br />
14.28 Covalent modification by ubiquitin and ubiquitin-like proteins is another way <strong>of</strong> regulating protein function 627
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 628<br />
stimulating transcription factors, however, auxin<br />
accelerates the degradation <strong>of</strong> several specific<br />
transcriptional repressors. The auxin receptor is<br />
in fact a ubiquitin ligase complex that targets<br />
the auxin-regulated transcriptional repressors<br />
for proteolysis. F-box proteins account for all<br />
<strong>of</strong> the auxin binding activity in plant extracts.<br />
14.29<br />
The Wnt pathway<br />
regulates <strong>cell</strong> fate during<br />
development and other<br />
processes in the adult<br />
Key concepts<br />
• Seven transmembrane-spanning receptors may<br />
control complex differentiation programs.<br />
• Wnts are lipid-modified ligands.<br />
• Wnts signal through multiple distinct receptors.<br />
• Wnts suppress degradation <strong>of</strong> -catenin, a<br />
multifunctional transcription factor.<br />
Wnt pathways function during embryonic development<br />
and in the adult in morphogenesis,<br />
body patterning, axis formation, proliferation,<br />
and <strong>cell</strong> motility. The classical Wnt <strong>signaling</strong><br />
mechanism was uncovered largely through<br />
studies <strong>of</strong> Drosophila and Xenopus development,<br />
as well as by analyzing genetic alterations in<br />
cancer.<br />
Wnt proteins are unusual extra<strong>cell</strong>ular ligands.<br />
In addition to carbohydrate, they contain<br />
covalently bound palmitate that is essential for<br />
their biological activity. Wnts transduce signals<br />
by binding to multiple distinct receptors. The<br />
most significant are members <strong>of</strong> the Frizzled<br />
family <strong>of</strong> seven-transmembrane-spanning receptors.<br />
Wnts regulate the stability <strong>of</strong> β-catenin,<br />
which either is rapidly degraded or, in response<br />
to Wnt, is stabilized to enter the nucleus and<br />
induce transcription by interacting with TCF<br />
(T-<strong>cell</strong> factor). Genes induced include c-jun, cyclin<br />
D1, and many others.<br />
The coordinated activities <strong>of</strong> the protein kinases<br />
glycogen synthase kinase 3 (GSK3) and<br />
casein kinase 1(CK1), the scaffolding proteins<br />
axin and adenomatous polyposis coli (APC), and<br />
the protein disheveled (DSH) are key to β-catenin<br />
stability. In the absence <strong>of</strong> Wnt, phosphorylation<br />
<strong>of</strong> β-catenin by CK1 and GSK3 promotes<br />
its ubiquitination and subsequent destruction<br />
by the proteasome. Axin and APC are required<br />
for phosphorylation <strong>of</strong> β-catenin by GSK3.<br />
In contrast to most seven transmembrane-<br />
spanning receptors, the Frizzled family has not<br />
yet been shown to have significant functions<br />
mediated by a heterotrimeric G protein, and G<br />
proteins may not be central to this pathway.<br />
Instead a proximal step in <strong>signaling</strong> by Frizzled<br />
involves binding to DSH, which inactivates the<br />
β-catenin destruction mechanism.<br />
Mutations that cause changes in the<br />
amounts <strong>of</strong> components <strong>of</strong> the classical pathway<br />
are common in a wide variety <strong>of</strong> cancers. Both<br />
Wnts and β-catenin may be viewed as protooncogenes.<br />
APC is a tumor suppressor and is<br />
mutated in the majority <strong>of</strong> human colorectal<br />
cancers, for example. Either too little or too<br />
much axin can also disrupt Wnt <strong>signaling</strong>, and<br />
axin, like APC, is a tumor suppressor.<br />
Wnts utilize additional <strong>signaling</strong> mechanisms.<br />
The receptor proteins Lrp5/6 (which are<br />
related to the low-density lipoprotein receptor)<br />
are Wnt receptors and also bind axin. Wnts bind<br />
to tyrosine kinase receptors to influence axon<br />
guidance and to other proteins that inhibit their<br />
function. Through DSH, Wnts can regulate the<br />
JNK MAPK pathway and Rho family G proteins<br />
to control planar <strong>cell</strong> polarity. Certain Wnts increase<br />
intra<strong>cell</strong>ular calcium to activate calciumdependent<br />
<strong>signaling</strong> pathways.<br />
14.30<br />
Diverse <strong>signaling</strong><br />
mechanisms are regulated<br />
by protein tyrosine kinases<br />
Key concepts<br />
• Many receptor protein tyrosine kinases are<br />
activated by growth factors.<br />
• Mutations in receptor tyrosine kinases can be<br />
oncogenic.<br />
• Ligand binding promotes receptor oligomerization<br />
and autophosphorylation.<br />
• Signaling proteins bind to the phosphotyrosine<br />
residues <strong>of</strong> the activated receptor.<br />
A large group <strong>of</strong> protein tyrosine kinases are<br />
receptors that span the plasma membrane and<br />
bind extra<strong>cell</strong>ular ligands, as shown in FIGURE<br />
14.34. The receptors are generally activated by<br />
growth factors whose normal physiological functions<br />
are to promote growth, proliferation, development,<br />
or maintenance <strong>of</strong> differentiated<br />
properties. This group includes receptors for insulin,<br />
epidermal growth factor (EGF), and<br />
platelet derived growth factor (PDGF). These<br />
receptors both control the activities <strong>of</strong> many<br />
other protein kinases <strong>of</strong> all families and directly<br />
regulate other classes <strong>of</strong> <strong>signaling</strong> proteins.<br />
628 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 629<br />
Because <strong>of</strong> their physiologic roles as growth<br />
regulators, mutations that activate receptor tyrosine<br />
kinases are <strong>of</strong>ten oncogenic. For example,<br />
the oncogene erbB results from the<br />
mutational loss <strong>of</strong> the extra<strong>cell</strong>ular ligand-binding<br />
domain <strong>of</strong> a kinase closely related to the<br />
EGF receptor. This mutation causes constitutive<br />
activation <strong>of</strong> the protein kinase domain.<br />
Point mutations that affect the transmembrane<br />
domain can also cause oncogenic activation, as<br />
is found in the EGF receptor-related neu/HER2<br />
oncogene (see 13.8 Cell growth and proliferation<br />
are activated by growth factors).<br />
Receptor tyrosine kinases are diverse both<br />
in their extra<strong>cell</strong>ular ligand-binding domains<br />
and, with the exception <strong>of</strong> a conserved tyrosine<br />
protein kinase domain, their intra<strong>cell</strong>ular regulatory<br />
regions. These receptors usually have<br />
one membrane span per monomer but some,<br />
such as the insulin receptor, which is a disulfidebonded<br />
heterotetramer, have two. Ligand binding<br />
to receptor tyrosine kinases favors receptor<br />
oligomerization and enhances kinase activity<br />
leading to increased Tyr phosphorylation <strong>of</strong> the<br />
intra<strong>cell</strong>ular domain <strong>of</strong> the receptor and <strong>of</strong> associated<br />
molecules. These tyrosine-phosphorylated<br />
motifs create docking sites for additional<br />
signal transducers and adaptors.<br />
A comparison <strong>of</strong> the PDGF and insulin receptors<br />
reveals common themes and a range <strong>of</strong><br />
behaviors <strong>of</strong> receptor tyrosine kinases. The two<br />
PDGF receptors are monomeric receptor tyrosine<br />
kinases. The insulin receptor exists in two<br />
alternatively spliced forms each <strong>of</strong> which is a<br />
heterotetramer <strong>of</strong> two and two subunits. In<br />
each case, the receptor is<strong>of</strong>orms utilize some<br />
unique <strong>signaling</strong> mechanisms.<br />
PDGF and insulin each stimulate the kinase<br />
activity <strong>of</strong> their receptors, causing oligomerization<br />
and autophosphorylation. Seven or more<br />
sites are phosphorylated on the PDGF receptor,<br />
and each phosphotyrosine residue generates a<br />
binding site for one or more SH2 domain-containing<br />
proteins as illustrated in FIGURE 14.35. The<br />
PDGF receptor binds PI 3-kinase, p190 Ras GAP,<br />
phospholipase C-, Src (which may catalyze additional<br />
Tyr phosphorylation <strong>of</strong> the receptor),<br />
and the SHP2 tyrosine phosphatase which itself<br />
binds the adaptor Grb2 (see 14.32 MAPKs are central<br />
to many <strong>signaling</strong> pathways). With the exception<br />
<strong>of</strong> Src, all <strong>of</strong> these proteins are also receptor<br />
substrates. Thus, substrates are recruited to the<br />
receptor as a consequence <strong>of</strong> specific interactions<br />
<strong>of</strong> substrate SH2 domains with receptor phosphotyrosine<br />
producing changes in activities and<br />
distributions <strong>of</strong> numerous intra<strong>cell</strong>ular signal<br />
EGF<br />
receptor<br />
CYTOPLASM<br />
Receptor protein tyrosine kinase families<br />
KINASE<br />
DOMAINS<br />
Insulin<br />
receptor<br />
Activation <strong>of</strong> the PDGF receptor leads to many outputs<br />
p190<br />
RasGAP<br />
PLC-γ<br />
PDGF<br />
PDGF<br />
receptor<br />
PDGF<br />
receptors<br />
Src<br />
p85 p110<br />
PI 3-kinase<br />
Shp2<br />
Kinase<br />
inserts<br />
SOS<br />
FGF<br />
receptor<br />
FIGURE 14.34 The monomeric tyrosine kinase receptors consist <strong>of</strong> a<br />
globular extra<strong>cell</strong>ular domain that binds ligand, a single transmembrane<br />
span, and a globular intra<strong>cell</strong>ular region containing the protein kinase<br />
domain. The intra<strong>cell</strong>ular regions contain additional sequences preceding,<br />
following, and, in the case <strong>of</strong> the PDGF and FGF receptor groups,<br />
inserted into the protein kinase domain. These regions contain sites <strong>of</strong><br />
tyrosine phosphorylation-dependent interactions. The insulin receptor<br />
is encoded by a single gene. The precursor is proteolyzed into and <br />
subunits, which are disulfide bonded to each other. Disulfide bonds also<br />
link two subunits, yielding an obligate heterotetramer.<br />
Grb2<br />
FIGURE 14.35 PDGF binds to its receptor and induces receptor autophosphorylation.<br />
The autophosphorylated receptor binds target proteins<br />
that contain SH2 domains.<br />
14.30 Diverse <strong>signaling</strong> mechanisms are regulated by protein tyrosine kinases 629
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 630<br />
transducers. This array <strong>of</strong> <strong>signaling</strong> events leads<br />
to increased proliferation <strong>of</strong> connective tissue<br />
during development and in wound healing.<br />
Autophosphorylation also occurs on the insulin<br />
receptor to stabilize the active state and to<br />
generate a smaller number <strong>of</strong> binding sites, as illustrated<br />
in FIGURE 14.36. A key event is the Tyr<br />
phosphorylation <strong>of</strong> insulin receptor substrate<br />
(IRS) proteins, notably IRS1, on as many as a<br />
dozen sites. IRS1 takes over interactions with several<br />
<strong>signaling</strong> effectors that, in the case <strong>of</strong> PDGF,<br />
bind directly to the receptor. Among these targets<br />
is PI 3-kinase which leads to activation <strong>of</strong><br />
Akt-2 and several essential metabolic actions <strong>of</strong><br />
insulin (see 14.16 Lipids and lipid-derived compounds<br />
are <strong>signaling</strong> molecules). IRS proteins are also phosphorylated<br />
by serine/threonine protein kinases to<br />
modulate their <strong>signaling</strong> capability.<br />
Tyr phosphorylation <strong>of</strong>ten enhances the enzymatic<br />
activity <strong>of</strong> the associated proteins. Other<br />
proteins gain enhanced function primarily as a<br />
consequence <strong>of</strong> greater proximity to targets<br />
achieved by binding through their SH2 domains<br />
to phosphotyrosine sites either on the receptors<br />
or IRS adaptors. The precise actions <strong>of</strong> the<br />
many tyrosine kinase receptors are determined<br />
by the overlapping sets <strong>of</strong> signal transducers<br />
with which they interact, as well as by detailed<br />
differences in amounts <strong>of</strong> signal transducers,<br />
adaptor accessory proteins and receptor expression<br />
patterns (see Figure 14.43).<br />
Insulin <strong>signaling</strong> through IRS1<br />
Insulin<br />
CYTOPLASM<br />
Insulin<br />
receptor<br />
IRS1<br />
PIP 2<br />
p85 p110<br />
PI 3-kinase<br />
PIP3<br />
Akt<br />
<strong>signaling</strong><br />
FIGURE 14.36 Insulin binding to its receptor causes activation <strong>of</strong> the<br />
receptor tyrosine protein kinase and autophosphorylation. The receptor<br />
kinase also phosphorylates IRS1, a large adaptor with many potential<br />
phosphorylation sites. IRS1 is an essential intermediate in insulin action.<br />
PI 3-kinase binds to IRS1 via the SH2 domain within its p85 subunit.<br />
Akt and PDK1 bind to PIP 3<br />
produced by activated PI 3-kinase so that<br />
PDK1 can phosphorylate and activate Akt (see Figure 14.18).<br />
14.31<br />
Src family protein kinases<br />
cooperate with receptor<br />
protein tyrosine kinases<br />
Key concepts<br />
• Src is activated by release <strong>of</strong> intrasteric inhibition.<br />
• Activation <strong>of</strong> Src involves liberation <strong>of</strong> modular<br />
binding domains for activation-dependent<br />
interactions.<br />
• Src <strong>of</strong>ten associates with receptors, including<br />
receptor tyrosine kinases.<br />
The first protein tyrosine kinase to be discovered<br />
was Src, which was identified as the transforming<br />
entity in the Rous sarcoma virus. Src is the<br />
prototype <strong>of</strong> a number <strong>of</strong> related enzymes, the<br />
Src family kinases. It participates in <strong>signaling</strong><br />
pathways regulated by numerous <strong>cell</strong> surface<br />
receptors, including those that lack their own<br />
kinase domain (see in 14.34 Diverse receptors recruit<br />
protein tyrosine kinases to the plasma membrane).<br />
Src is bound to the plasma membrane<br />
via an N-terminal myristoyl group. In the inactive<br />
state, Src is phosphorylated on Tyr527, C-<br />
terminal to its catalytic domain, by CSK<br />
(C-terminal Src kinase).<br />
The structure and regulation <strong>of</strong> Src is depicted<br />
in FIGURE 14.37. Phosphorylation <strong>of</strong> Tyr527<br />
causes it to bind to its own SH2 domain. The<br />
SH2 and SH3 domains suppress the kinase activity<br />
through interactions on the surface <strong>of</strong> the<br />
protein. The SH3 domain binds to an SH3 binding<br />
site distant from the active site. Activation<br />
<strong>of</strong> Src by dephosphorylation <strong>of</strong> Tyr527 causes<br />
its SH2 to dissociate; this causes a conformational<br />
change in the SH3 domain to dissociate<br />
it from the binding site. Viral isolates <strong>of</strong> Src are<br />
<strong>of</strong>ten truncated prior to Tyr527, which increases<br />
their activity.<br />
Conformational changes in the kinase domain<br />
resulting from dissociation <strong>of</strong> the SH3 promote<br />
Src autophosphorylation on Tyr416 in its<br />
activation loop and further increase protein kinase<br />
activity. An important consequence <strong>of</strong> the<br />
interaction <strong>of</strong> Src with its own SH2 and SH3<br />
domains is that these domains cannot bind anything<br />
else when in the autoinhibited state; therefore,<br />
other interactions are promoted when the<br />
SH2 and SH3 domains are released from their<br />
associations with the Src kinase domain. The<br />
heterologous interactions <strong>of</strong> the SH2 and SH3<br />
domains contribute to Src localization and <strong>signaling</strong>.<br />
630 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 631<br />
Structure and regulation <strong>of</strong> Src<br />
INACTIVE<br />
ACTIVE<br />
KINASE<br />
DOMAIN<br />
SH3<br />
SH2<br />
P<br />
P<br />
Tyr527<br />
FIGURE 14.37 The structures <strong>of</strong> inactive and active Src are compared. The inactive<br />
protein is autoinhibited by binding to its own SH2 and SH3 domains.<br />
The SH2 domain binds to phosphorylated Tyr527. The SH3 domain binds to a<br />
noncanonical SH3-binding motif on the opposite side <strong>of</strong> the kinase domain<br />
active site. In contrast to the steric inhibition <strong>of</strong> PKA caused by its R subunit,<br />
inhibition <strong>of</strong> Src by its SH2 and SH3 domains is allosteric. In the active structure<br />
the SH2 and SH3 domains are not bound to the kinase domain and are<br />
available for heterologous interactions. Structures generated from Protein Data<br />
Base files 1FMK and 1Y57.<br />
14.32<br />
MAPKs are central to<br />
many <strong>signaling</strong> pathways<br />
Key concepts<br />
• MAPKs are activated by Tyr and Thr<br />
phosphorylation.<br />
• The requirement for two phosphorylations creates<br />
a <strong>signaling</strong> threshold.<br />
• The ERK1/2 MAPK pathway is usually regulated<br />
through Ras.<br />
Mitogen-activated protein kinases (MAPKs) are<br />
present in all eukaryotes. They are among the<br />
most common multifunctional protein kinases<br />
mediating <strong>cell</strong>ular regulatory events in response<br />
to many ligands and other stimuli. MAPKs are<br />
activated by protein kinase cascades consisting<br />
<strong>of</strong> at least three protein kinases acting sequentially,<br />
as illustrated in FIGURE 14.38. Activation<br />
<strong>of</strong> a MAPK is catalyzed by a MAPK kinase<br />
(MAP2K), which is itself activated by phosphorylation<br />
by a MAPK kinase kinase (MAP3K).<br />
MAP3Ks are activated by a variety <strong>of</strong> mechanisms<br />
including phosphorylation by MAP4Ks,<br />
oligomerization, and binding to activators such<br />
as small G proteins.<br />
MAP2Ks are activated by phosphorylation<br />
on two Ser/Thr residues; MAP2Ks then activate<br />
MAPKs by dual phosphorylation on Tyr<br />
and Thr residues (Figure 14.30). Each MAP2K<br />
phosphorylates a limited set <strong>of</strong> MAPKs and few<br />
or no other substrates. The great specificity <strong>of</strong><br />
MAP2Ks is one means <strong>of</strong> insulating MAPKs<br />
from activation by inappropriate signals. Both<br />
Tyr and Thr phosphorylations are required for<br />
maximum MAPK enzymatic activity.<br />
Studies on the MAPK ERK2 led to an understanding<br />
<strong>of</strong> the events induced by phosphorylation<br />
that are important for increased activity.<br />
Conformational changes include refolding <strong>of</strong><br />
the activation loop to improve substrate positioning<br />
and realignment <strong>of</strong> catalytic residues;<br />
this is most obvious in the repositioning <strong>of</strong> <br />
helix C, which contains a Glu involved in phosphoryl<br />
transfer.<br />
Amplification occurs moving down the cascade<br />
from the MAP3K to the MAP2K step because<br />
the MAP2Ks are much more abundant<br />
than the MAP3Ks. The MAP2K to MAPK step<br />
may also amplify the signal if the MAPK is present<br />
in excess <strong>of</strong> the MAP2K. In addition, the<br />
phosphorylation <strong>of</strong> a MAPK by a MAP2K on a<br />
14.32 MAPKs are central to many <strong>signaling</strong> pathways 631
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 632<br />
Generic<br />
G Protein<br />
small or<br />
heterotrimer<br />
MAP4K<br />
MAP3K<br />
MAP2K<br />
MAPK<br />
Major<br />
targets:<br />
Transcription<br />
factor<br />
Protein<br />
kinase<br />
Output<br />
S.<br />
cerevisiae<br />
Gβγ<br />
Ste20p<br />
Ste11p<br />
Ste7p<br />
Fus3p<br />
Ste12p<br />
Mating<br />
MAPK pathways<br />
Mammals<br />
Ras Rac/Cdc42 Rac<br />
?<br />
Ste20 Ste20<br />
PAK/PKC ?<br />
family family<br />
Raf<br />
MEK1<br />
MEK2<br />
ERK1<br />
ERK2<br />
Elk-1<br />
Rsk<br />
many<br />
MEK4<br />
MEK7<br />
JNK1<br />
JNK2<br />
JNK3<br />
c-Jun<br />
ATF2<br />
many<br />
MEK3<br />
MEK6<br />
p38α<br />
p38β<br />
p38γ<br />
p38δ<br />
MEF2<br />
MAPKAPK2<br />
Proliferation<br />
Development<br />
Differentiation<br />
(and other processes)<br />
MEKK2<br />
MEKK3<br />
MEK5<br />
ERK5<br />
MEF2<br />
FIGURE 14.38 MAPK pathways can be regulated by a diverse group <strong>of</strong> upstream<br />
regulatory mechanisms that <strong>of</strong>ten include adaptors, small G proteins, and MAP4Ks.<br />
These molecules impinge on the activities <strong>of</strong> MAP3Ks. MAP3Ks regulate one or<br />
more MAP2Ks depending on localization and scaffolding. The MAP2Ks display<br />
great selectivity for a single MAPK type. MAPKs have overlapping and unique<br />
substrates and participate in <strong>signaling</strong> cascades leading to many <strong>cell</strong>ular responses.<br />
Rsk<br />
Tyr and a Thr residue creates cooperative activation<br />
<strong>of</strong> the MAPK; this is another mechanism,<br />
in addition to those described for PKA and<br />
calmodulin, to introduce a threshold and apparently<br />
cooperative behavior into the pathway<br />
over a narrow range <strong>of</strong> input signal. This<br />
multistep cascade provides multiple sites for<br />
modulatory inputs from other pathways.<br />
Stabilized interactions between components<br />
are also important. MAP2Ks, as well as MAPK<br />
substrates and MAPK phosphatases, generally<br />
contain a basic/hydrophobic docking motif that<br />
interacts with acidic residues and binds in a hydrophobic<br />
groove on the MAPK catalytic domain.<br />
Additional components including scaffolds<br />
are necessary for the efficient activation <strong>of</strong><br />
MAPK cascades in <strong>cell</strong>s and usually have additional<br />
functions. Several scaffolds have been<br />
identified that bind to two or more components<br />
for each <strong>of</strong> the three major MAPK cascades, the<br />
ERK1/2, JNK1-3, and p38 α, β, γ, and δ cascades.<br />
The ERK1/2 pathway is regulated by most<br />
<strong>cell</strong> surface receptors, including receptors that<br />
employ tyrosine kinases, GPCRs, and others. The<br />
PDGF receptor, like most receptor systems, activates<br />
the ERK1/2 cascade through Ras. PDGF<br />
stimulates autophosphorylation <strong>of</strong> its receptor<br />
and the subsequent association <strong>of</strong> effectors with<br />
its cytoplasmic domain (see 14.30 Diverse <strong>signaling</strong><br />
mechanisms are regulated by protein tyrosine kinases).<br />
In response to PDGF, ERK1/2 promotes <strong>cell</strong><br />
proliferation and differentiation by phosphorylation<br />
<strong>of</strong> membrane enzymes, proteins involved<br />
in determining <strong>cell</strong> shape and motility, and also<br />
by concentrating in the nucleus to phosphorylate<br />
regulatory factors that control transcription.<br />
14.33<br />
Cyclin-dependent protein<br />
kinases control the <strong>cell</strong><br />
cycle<br />
Key concepts<br />
• The <strong>cell</strong> cycle is regulated by cyclin-dependent<br />
protein kinases (CDKs).<br />
• Activation <strong>of</strong> CDKs involves protein binding,<br />
dephosphorylation, and phosphorylation.<br />
Cell division is regulated positively and negatively<br />
by factors that stimulate proliferation and<br />
inputs that monitor <strong>cell</strong> state. The sum <strong>of</strong> these<br />
factors is integrated in the regulation <strong>of</strong> cyclindependent<br />
protein kinases (CDKs). CDKs are<br />
protein serine/threonine kinases that are major<br />
regulators <strong>of</strong> <strong>cell</strong> cycle progression. Most<br />
CDKs are regulated both by kinases and phosphatases<br />
and by association with other proteins<br />
called cyclins. Cyclins are synthesized and degraded<br />
every <strong>cell</strong> cycle. Because most CDKs are<br />
dependent upon cyclin binding for activation,<br />
the timing <strong>of</strong> the synthesis and degradation <strong>of</strong><br />
individual cyclins determines when a CDK will<br />
function. The most notable noncycling member<br />
<strong>of</strong> the CDK family is Cdk5, which is highly expressed<br />
in terminally differentiated neurons.<br />
Cdk5 binds the non-cyclin protein p35 as its activating<br />
subunit.<br />
We will briefly examine the regulation <strong>of</strong><br />
Cdc2, a major CDK in both mammals and yeast.<br />
The first step in regulation <strong>of</strong> Cdc2 is the association<br />
with cyclin. A second step required for<br />
activation <strong>of</strong> Cdc2 is phosphorylation <strong>of</strong> a Thr<br />
632 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 633<br />
CDKs require cyclin binding for activation<br />
Tyr15<br />
Lys33<br />
Glu51<br />
Cdk2<br />
Cyclin A<br />
FIGURE 14.39 The view <strong>of</strong> the crystal structure <strong>of</strong> CDK2 bound to cyclin A<br />
shows residues in the ATP binding site. The enlargement on the right shows<br />
the interaction between Lys33 and Glu51, catalytic residues that interact with<br />
ATP to promote phosphoryl transfer. Tyr15 is phosphorylated in inactive forms<br />
<strong>of</strong> CDK2. A phosphoryl group on Tyr15 inhibits CDK activity by interfering with<br />
ATP binding. Structure generated from Protein Data Bank file 1JST.<br />
residue in its activation loop by another CDK<br />
type kinase. In spite <strong>of</strong> its association with cyclin,<br />
this form <strong>of</strong> Cdc2 is not yet active due to<br />
inhibitory phosphorylation <strong>of</strong> Tyr and Thr<br />
residues in the ATP binding pocket. Release <strong>of</strong><br />
inhibition by dephosphorylation <strong>of</strong> the residues<br />
in the ATP pocket is catalyzed by the Cdc25 family<br />
<strong>of</strong> phosphoprotein phosphatases, resulting in<br />
activation <strong>of</strong> Cdc2. The proximity <strong>of</strong> the Tyr<br />
residue to catalytic residues is shown in FIGURE<br />
14.39. The complexity <strong>of</strong> activation <strong>of</strong> CDKs<br />
makes possible the imposition <strong>of</strong> <strong>cell</strong> cycle checkpoints.<br />
For more on CDKs and cyclins see 11.4<br />
The <strong>cell</strong> cycle is a cycle <strong>of</strong> CDK function.<br />
14.34<br />
Diverse receptors recruit<br />
protein tyrosine kinases<br />
to the plasma membrane<br />
Key concepts<br />
• Receptors that bind protein tyrosine kinases use<br />
combinations <strong>of</strong> effectors similar to those used by<br />
receptor tyrosine kinases.<br />
• These receptors <strong>of</strong>ten bind directly to transcription<br />
factors.<br />
Many receptors act through protein tyrosine<br />
kinases, but their <strong>cell</strong> surface receptors lack kinase<br />
activity. Instead, these receptors act by recruiting<br />
and activating protein tyrosine kinases<br />
at the plasma membrane. In this group <strong>of</strong> receptors<br />
are integrins, which are key molecules involved<br />
in <strong>cell</strong> adhesion, growth hormone<br />
receptors, and receptors that mediate inflammatory<br />
and immune responses. While their<br />
structures vary enormously, their mechanisms<br />
<strong>of</strong> action are related.<br />
Integrins are receptors whose major function<br />
is to attach <strong>cell</strong>s to the extra<strong>cell</strong>ular matrix. They<br />
also mediate some interactions with proteins on<br />
other <strong>cell</strong>s, as depicted in FIGURE 14.40. Ligands for<br />
integrins include a number <strong>of</strong> extra<strong>cell</strong>ular matrix<br />
proteins, such as fibronectin, as well as <strong>cell</strong><br />
surface proteins that cooperate in <strong>cell</strong>-<strong>cell</strong> interactions.<br />
Integrin ligation provides <strong>cell</strong>s with information<br />
about their environment that influences<br />
<strong>cell</strong> behavior. Ligation <strong>of</strong> integrins initiates signals<br />
that control <strong>cell</strong> programs, including <strong>cell</strong> cycle entry,<br />
proliferation, survival, differentiation, changes<br />
in <strong>cell</strong> shape, and motility, as well as fine-tuning<br />
responses to other ligands. For more details on integrins<br />
see 15.13 Most integrins are receptors for extra<strong>cell</strong>ular<br />
matrix proteins and 15.14 Integrin receptors<br />
participate in <strong>cell</strong> <strong>signaling</strong>.<br />
14.34 Diverse receptors recruit protein tyrosine kinases to the plasma membrane 633
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 634<br />
Crk<br />
ECM<br />
Crk<br />
Src<br />
CAS<br />
CYTOPLASM<br />
FAK<br />
SOS<br />
Integrin <strong>signaling</strong><br />
INTEGRINS<br />
Paxillin<br />
Talin<br />
Vinculin<br />
Grb2<br />
Actin<br />
filament<br />
ILK<br />
p85 PI 3-<br />
Kinase<br />
p110<br />
Vinculin<br />
Tensin<br />
Talin<br />
FIGURE 14.40 Integrins bind to an array <strong>of</strong> cytoplasmic proteins to regulate<br />
the cytoskeleton and intra<strong>cell</strong>ular <strong>signaling</strong> pathways. The associated cytoskeletal<br />
elements include actin filaments and focal adhesion proteins -actinin, vinculin,<br />
paxillin, and talin. Signaling molecules include the focal adhesion kinase<br />
FAK; the adaptors Cas, Crk and Grb2; Src and CSK (see 14.31 Src family protein<br />
kinases cooperate with receptor protein tyrosine kinases), PI 3-kinase (see 14.16<br />
Lipids and lipid-derived compounds are <strong>signaling</strong> molecules); and the Ras exchange<br />
factor SOS. Stimulation <strong>of</strong> GTP binding <strong>of</strong> Ras by SOS leads to activation<br />
<strong>of</strong> the MAPK pathway (see 14.32 MAPKs are central to many <strong>signaling</strong><br />
pathways).<br />
Talin and α-actinin are among cytoskeletal<br />
proteins that interact directly with certain integrin<br />
subunits. These cytoskeletal proteins link<br />
integrins to complex cytoskeletal structures<br />
known as focal adhesions.<br />
Focal adhesions connect the cytoskeleton to<br />
signal transduction cascades that communicate<br />
states <strong>of</strong> <strong>cell</strong>ular attachment to the regulation <strong>of</strong><br />
<strong>cell</strong>ular responses. Focal adhesion complexes contain<br />
the focal adhesion kinase FAK, which is activated<br />
by integrin ligation. Autophosphorylation<br />
<strong>of</strong> FAK recruits <strong>signaling</strong> proteins containing SH2<br />
domains, especially the p85 subunit <strong>of</strong> PI 3-kinase<br />
and Src family protein kinases. The <strong>signaling</strong><br />
molecules associated with the integrin-bound<br />
cytoskeletal proteins, whether focal adhesions or<br />
other structural complexes, mediate the diverse<br />
actions <strong>of</strong> integrins. The association <strong>of</strong> cytoskeletal<br />
proteins with integrin receptors also causes<br />
functional changes to the receptors.<br />
Signals that act over a distance, such as hormones,<br />
can also employ nonreceptor tyrosine<br />
kinases to transmit their message inside a <strong>cell</strong>.<br />
Growth hormone (GH) is a protein hormone<br />
secreted by the anterior pituitary gland that regulates<br />
bone growth, fat metabolism, and other<br />
<strong>cell</strong>ular growth phenomena. Absence <strong>of</strong> growth<br />
hormone results in short stature, whereas hypersecretion<br />
causes acromegaly, a form <strong>of</strong> gigantism.<br />
The GH receptor is a member <strong>of</strong> the<br />
cytokine receptor family, which includes receptors<br />
for prolactin, erythropoietin, leptin, and<br />
interleukins. All these receptors display similar<br />
biochemical functions, such as association with<br />
members <strong>of</strong> the JAK/TYK family <strong>of</strong> protein tyrosine<br />
kinases, but select for different but overlapping<br />
sets <strong>of</strong> cytoplasmic <strong>signaling</strong> proteins.<br />
Signal transduction by the GH receptor provides<br />
a model for receptors that lack enzymatic function<br />
and act as agonist-promoted scaffolds for<br />
intra<strong>cell</strong>ular <strong>signaling</strong> proteins.<br />
FIGURE 14.41 shows the structure <strong>of</strong> growth<br />
hormone bound to the extra<strong>cell</strong>ular domain <strong>of</strong><br />
its receptor. The majority <strong>of</strong> binding energy<br />
comes from only a small number <strong>of</strong> residues in<br />
the binding interface. Inside the <strong>cell</strong>, <strong>signaling</strong><br />
by the GH receptor depends significantly on its<br />
association with the cytoplasmic tyrosine protein<br />
kinase Janus kinase 2 (JAK2). FIGURE 14.42<br />
shows that JAK2 binds to a proline-rich region<br />
<strong>of</strong> the receptor. Ligand binding induces receptor<br />
dimerization, which then promotes<br />
activation <strong>of</strong> JAK2 through intermolecular autophosphorylation.<br />
GH <strong>signaling</strong> is thus mediated primarily by<br />
inducing Tyr phosphorylation. In addition to<br />
JAK2 autophosphorylation, the receptor itself<br />
becomes Tyr phosphorylated. As is true for receptor<br />
tyrosine kinases, Tyr phosphorylation <strong>of</strong> the<br />
growth hormone receptor creates binding sites<br />
for <strong>signaling</strong> proteins that contain phosphotyrosine-binding<br />
domains. Primary targets are transcription<br />
factors known as signal transducers and<br />
activators <strong>of</strong> transcription, or STATs. STATs contain<br />
SH2 domains and bind Tyr-phosphorylated<br />
motifs on the growth hormone receptor. While<br />
receptor bound, STATs are Tyr phosphorylated<br />
by JAK2 and then released to travel to the nucleus<br />
to mediate changes in transcription.<br />
The growth hormone receptor and the associated<br />
JAK2 also activate other <strong>signaling</strong> pathways.<br />
For example, the adaptor Shc is Tyr<br />
phosphorylated by JAK2. Engagement <strong>of</strong> Shc<br />
leads to activation <strong>of</strong> Ras and the ERK1/2 MAPK<br />
pathway. Adaptors specialized for insulin-<strong>signaling</strong><br />
pathways, insulin receptor substrates (IRS)<br />
1, 2, and 3, are also growth hormone targets, perhaps<br />
reflecting the ability <strong>of</strong> growth hormone<br />
to induce certain insulin-like metabolic actions.<br />
Feedback circuits are also engaged during<br />
GH <strong>signaling</strong>. The growth hormone receptor<br />
complex binds the adaptor SH2-B, which has a<br />
634 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 635<br />
Growth hormone structure<br />
Growth hormone <strong>signaling</strong> is transduced by JAK2<br />
Growth hormone<br />
receptor<br />
hGH<br />
Growth<br />
hormone<br />
Dimerization<br />
JAK2<br />
JAK2<br />
ΔΔG (kcal/mol)<br />
> 1.5<br />
0.5 to 1.5<br />
-0.5 to 0.5<br />
< -0.5<br />
untested<br />
CYTOPLASM<br />
JAK2s bind and<br />
phosphorylate<br />
receptor<br />
STAT<br />
STAT<br />
STATs bind and<br />
are phosphorylated<br />
FIGURE 14.41 Proteins <strong>of</strong>ten interact over a large surface area. Growth<br />
hormone binding to its receptor is an example <strong>of</strong> the energy <strong>of</strong> binding<br />
coming primarily from a small number <strong>of</strong> the contacts between<br />
the two proteins, creating an interaction hot spot. The complex <strong>of</strong><br />
growth hormone bound to the growth hormone receptor-binding domain<br />
determined by crystallography has been peeled apart in this figure<br />
to show the binding energy associated with residues in the binding<br />
interface from each protein determined by mutagenesis and binding<br />
studies. Fewer than half <strong>of</strong> the residues in the interface contribute<br />
the majority <strong>of</strong> binding energy. Reproduced with permission from T.<br />
Clackson and J. A. Wells. 1995. Science. 267: 383–386. © AAAS. Photos<br />
courtesy <strong>of</strong> Tim Clackson, ARIAD Pharmaceuticals, Inc.<br />
NUCLEUS<br />
Phosphorylated<br />
STATs bind DNA<br />
FIGURE 14.42 The growth hormone receptor binds to JAK2. Many GH signals<br />
are mediated by Tyr phosphorylation <strong>of</strong> the receptor by JAK2, which creates<br />
binding sites for <strong>signaling</strong> molecules with SH2 domains, notably STATs. STATs<br />
then enter the nucleus to cause changes in gene transcription.<br />
stimulatory effect on growth hormone <strong>signaling</strong>.<br />
On the other hand, suppressors <strong>of</strong> cytokine<br />
<strong>signaling</strong> (SOCS proteins) are among the genes<br />
whose transcription is induced by growth hormone.<br />
As the name indicates, SOCS proteins inhibit<br />
cytokine <strong>signaling</strong> in some if not all cases by<br />
inhibiting the activity <strong>of</strong> JAK2. SOCS proteins<br />
contain an SH2 domain that facilitates their binding<br />
either to phosphorylated JAK2 or cytokine<br />
receptors. The mechanism <strong>of</strong> <strong>signaling</strong> inhibition<br />
may differ among SOCS proteins because<br />
some require the GH receptor to interfere with<br />
JAK2 <strong>signaling</strong>. SOCS-1, on the other hand, binds<br />
directly to the JAK2 activation loop and does not<br />
require a receptor to inhibit JAK2 activity. This<br />
mechanism may be particularly important in GH<br />
<strong>signaling</strong> because, in contrast to the ligand-induced<br />
down regulation mechanisms controlling<br />
many receptors, the GH receptor is degraded in<br />
a ligand-independent manner.<br />
Receptors for cytokines also act by recruiting<br />
tyrosine kinases. The cytokines—<strong>signaling</strong><br />
proteins that modulate inflammation and <strong>cell</strong><br />
growth and differentiation—include interleukins,<br />
leukemia inhibitory factor, oncostatin M, cardiotrophin-1,<br />
cardiotrophin-like cytokine, and<br />
ciliary neurotrophic factor (CNTF). Each cytokine<br />
binds a unique receptor, but each receptor<br />
binds a transmembrane protein called gp130.<br />
Mechanisms <strong>of</strong> <strong>signaling</strong> by gp130 involve interactions<br />
with tyrosine kinases <strong>of</strong> the JAK/TYK<br />
types and transcription factors in the STAT family.<br />
This mechanism is similar to those employed<br />
by the growth hormone receptor.<br />
14.34 Diverse receptors recruit protein tyrosine kinases to the plasma membrane 635
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 636<br />
CD3<br />
Ligand<br />
Receptor<br />
Adaptor/<br />
subunit<br />
Transducer<br />
Kinase<br />
cascade<br />
Transcription<br />
factor<br />
complex<br />
TCR<br />
MHC<br />
Antigen<br />
ITAM<br />
CD3<br />
Receptor <strong>signaling</strong> pathways<br />
Growth<br />
PDGF Insulin hormone IL-1β TGF-β<br />
PDGF<br />
receptor<br />
SHP2/Grb2<br />
SOS/Ras<br />
MAPK<br />
Ternary<br />
complex<br />
factors<br />
Insulin<br />
receptor<br />
IRS1<br />
PI 3-kinase<br />
Akt2<br />
FOXO<br />
GH<br />
receptor<br />
STATs<br />
T <strong>cell</strong> receptor <strong>signaling</strong><br />
Lck<br />
After binding <strong>of</strong> the TCR<br />
to the MHC-antigen<br />
complex, Lck<br />
phosphorylates ITAMS<br />
IL-1β<br />
receptor<br />
gp130<br />
STATs<br />
TGF-β<br />
type II<br />
receptor<br />
Type I<br />
receptor<br />
SMADs<br />
FIGURE 14.43 Major <strong>signaling</strong> cascades controlled by PDGF, insulin, TGF-, IL-<br />
1, and growth hormone are compared. Each receptor either contains or interacts<br />
with a protein kinase that associates with or recruits a transducer. The<br />
transducer regulates downstream effectors either directly or through an intermediate<br />
protein kinase cascade. The effectors shown are transcriptional regulators.<br />
Phosphorylation by the transducer or kinase cascade activates all <strong>of</strong> the effectors<br />
except the FOXO proteins, which may be excluded from the nucleus by phosphorylation.<br />
The table only shows snapshots <strong>of</strong> much more complex <strong>signaling</strong> networks<br />
controlled by these ligands. Many <strong>of</strong> these and other intermediates serve<br />
multiple ligands. For example, IRS proteins also contribute to growth hormone<br />
and IL-1 <strong>signaling</strong>, and MAPK pathways are regulated by all <strong>of</strong> these ligands.<br />
ZAP-70<br />
FIGURE 14.44 The T <strong>cell</strong> receptor (TCR) is a multisubunit receptor. It is phosphorylated<br />
on activation motifs or ITAMs by Lck, or a related Src family protein<br />
kinase. The phosphorylated residues create binding sites for another tyrosine<br />
protein kinase ZAP-70. ZAP-70 then recruits other <strong>signaling</strong> molecules to the<br />
complex including phospholipase C, PI 3-kinase, and a Ras exchange factor to<br />
activate downstream <strong>signaling</strong> pathways.<br />
JAK<br />
JAK<br />
Unlike many cytokine receptors in this class,<br />
the CNTF receptor does not itself span the membrane.<br />
Instead, it is glycosyl phosphoinositol<br />
(GPI)-linked to the outer face <strong>of</strong> the plasma<br />
membrane. The GPI linkage is a covalent bond,<br />
and the receptor can be released into the extra<strong>cell</strong>ular<br />
fluid by a specific phospholipase. The<br />
freed receptor may interact with membranes <strong>of</strong><br />
other <strong>cell</strong>s to induce signals.<br />
The use <strong>of</strong> a common signal transducing<br />
subunit, gp130, suggests that unique mechanisms<br />
exist to create ligand-specific responses;<br />
under some circumstances competition by the<br />
ligand binding subunits for interaction with the<br />
gp130 signal transducer may influence <strong>signaling</strong><br />
outcomes. FIGURE 14.43 illustrates some parallels<br />
in <strong>signaling</strong> pathways initiated by receptors<br />
with associated or intrinsic protein kinases.<br />
The last receptor type we will discuss takes<br />
the concept <strong>of</strong> the specific and common subunits<br />
even further. The complex multiprotein<br />
T <strong>cell</strong> receptor (TCR) is found uniquely on T<br />
lymphocytes and is responsible for the ability <strong>of</strong><br />
these <strong>cell</strong>s to recognize and respond to specific<br />
antigens. The TCR, illustrated in FIGURE 14.44, is<br />
composed <strong>of</strong> eight subunits that can be described<br />
as an assembly <strong>of</strong> four dimers, αβ, γε, δε, and ζζ.<br />
The specificity <strong>of</strong> antigen recognition is determined<br />
by the α and β subunits, which are different<br />
for each <strong>cell</strong>. The remaining subunits are<br />
invariant in TCRs.<br />
The CD3 complex γ, δ, and ε subunits are<br />
similar in sequence to one another. The ζ chain,<br />
unlike the other subunits, appears on certain<br />
other <strong>cell</strong> types and may be a component <strong>of</strong><br />
other receptors, such as the Fc receptor, which<br />
binds a portion <strong>of</strong> certain immunoglobulins.<br />
A motif called the immunoreceptor tyrosine-based<br />
activation motif, or ITAM, which features<br />
closely spaced pairs <strong>of</strong> Tyr residues, is key<br />
to <strong>signaling</strong> by the TCR. The CD3 subunits each<br />
contain one ITAM and the ζ chain contains three<br />
ITAMs, for a total <strong>of</strong> ten motifs in each TCR.<br />
Engagement <strong>of</strong> the TCR causes the Src family<br />
kinases Lck and Fyn to phosphorylate the pairs<br />
<strong>of</strong> Tyr residues in the ITAMs. The ITAMs then<br />
bind the tandem SH2 domains <strong>of</strong> the protein tyrosine<br />
kinase ζ-chain-associated protein <strong>of</strong> 70<br />
kDa (ZAP-70), which becomes activated by Src.<br />
Tyr phosphorylation sites on ZAP-70 bind to<br />
other adaptors and <strong>signaling</strong> molecules, and Tyr<br />
phosphorylation by ZAP-70 activates additional<br />
signal transducers. The sum <strong>of</strong> these events leads<br />
to the downstream responses <strong>of</strong> T <strong>cell</strong>s to antigen<br />
engagement, which include <strong>cell</strong> cycle progression<br />
and the elaboration <strong>of</strong> cytokines such<br />
as interleukin-2.<br />
636 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 637<br />
14.35<br />
What’s next?<br />
New <strong>signaling</strong> proteins and new regulatory interactions<br />
seem to show up every day. The challenge<br />
now is to understand how <strong>cell</strong>s organize<br />
these proteins and their individual interactions<br />
to create adaptable information-processing networks.<br />
How do <strong>cell</strong>s use simple chemical reactions<br />
to sort and integrate multiple simultaneous<br />
inputs and then direct this information to diverse<br />
effector machinery? How do they interpret<br />
the inputs in the context <strong>of</strong> their growth and<br />
metabolic activities? In principle, three areas <strong>of</strong><br />
research have to contribute to allow us to understand<br />
integrative <strong>cell</strong>ular <strong>signaling</strong>.<br />
First, we need real-time, noninterfering<br />
biosensors to measure intra<strong>cell</strong>ular <strong>signaling</strong><br />
reactions. Most current sensors use combinations<br />
<strong>of</strong> fluorescent moieties and signal-binding<br />
protein domains to provide fast optical<br />
readouts. For many pathways, several reactions<br />
can be monitored within <strong>cell</strong>s over subsecond<br />
time scales. We need more, better, and faster<br />
sensors and sensors that can report with single<strong>cell</strong><br />
and sub<strong>cell</strong>ular resolution. Genetically encoded<br />
sensors will be complemented by synthetic<br />
molecules.<br />
Our ability to manipulate <strong>signaling</strong> networks<br />
is also improving dramatically but still<br />
falls short. We can manipulate <strong>signaling</strong> networks<br />
by overexpression, knockout, and knockdown<br />
<strong>of</strong> genes, but <strong>signaling</strong> pathways are<br />
wonderfully adaptive and frequently circumvent<br />
our best efforts to control them. We still<br />
need chemical regulators that can act promptly<br />
in <strong>cell</strong>s. Structure-based design <strong>of</strong> such regulatory<br />
molecules will be vital.<br />
Last, our ability to analyze the behavior <strong>of</strong><br />
<strong>signaling</strong> networks depends on our ability to<br />
measure and interpret <strong>signaling</strong> quantitatively.<br />
It is ironic but true that really complex systems<br />
cannot be described without explicit quantitative<br />
models for how they work. Computational<br />
modeling and simulation <strong>of</strong> <strong>signaling</strong> networks<br />
requires both better theoretical understanding<br />
<strong>of</strong> network dynamics and better algorithmic implementation.<br />
The goal is to understand how <strong>cell</strong>s think.<br />
14.36<br />
Summary<br />
Signal transduction encompasses mechanisms<br />
used by all <strong>cell</strong>s to sense and react to stimuli in<br />
their environment. Cells express receptors that<br />
recognize specific extra<strong>cell</strong>ular stimuli, includ-<br />
ing nutrients, hormones, neurotransmitters,<br />
and other <strong>cell</strong>s. Upon receptor binding, signals<br />
are converted to well-defined intra<strong>cell</strong>ular chemical<br />
or physical reactions that change the activities<br />
and the organization <strong>of</strong> protein complexes<br />
within <strong>cell</strong>s. The changes directed by the stimuli<br />
lead to altered <strong>cell</strong> behavior. The behavior <strong>of</strong><br />
the <strong>cell</strong> is determined then by its intra<strong>cell</strong>ular<br />
state and the integrated information from extra<strong>cell</strong>ular<br />
stimuli so that the appropriate responses<br />
are achieved.<br />
The basic biochemical components and<br />
processes <strong>of</strong> signal transduction are conserved<br />
throughout biology. Families <strong>of</strong> proteins are<br />
used in a variety <strong>of</strong> ways for many different<br />
physiological purposes. Cells <strong>of</strong>ten use the same<br />
series <strong>of</strong> <strong>signaling</strong> proteins to regulate multiple<br />
processes, such as transcription, ion transport,<br />
locomotion, and metabolism.<br />
Signaling pathways are assembled into <strong>signaling</strong><br />
networks to allow the <strong>cell</strong> to coordinate<br />
its responses to multiple inputs with its ongoing<br />
functions. It is now possible to discern conserved<br />
reaction sequences in and between<br />
pathways in <strong>signaling</strong> networks that are analogous<br />
to devices within the circuits <strong>of</strong> analog<br />
computers: amplifiers, logic gates, feedback and<br />
feed-forward controls, and memory.<br />
References<br />
14.1 Introduction<br />
Review<br />
Sauro, H. M. and Kholodenko, B. N., 2004.<br />
Quantitative analysis <strong>of</strong> <strong>signaling</strong> networks.<br />
Prog. Biophys. Mol. Biol. v. 86 p.<br />
5–43.<br />
Research<br />
Milo, R., Shen-Orr, S., Itzkovitz, S., Kashtan,<br />
N., Chklovskii, D., and Alon, U., 2002.<br />
Network motifs: simple building blocks <strong>of</strong><br />
complex networks. Science v. 298 p.<br />
824–827.<br />
14.2 Cellular <strong>signaling</strong> is primarily chemical<br />
Review<br />
Arshavsky, V. Y., Lamb, T. D., and Pugh, E. N.,<br />
Jr., 2002. G proteins and phototransduction.<br />
Annu. Rev. Physiol. v. 64 p. 153–187.<br />
Caterina, M. J. and Julius, D., 2001. The<br />
vanilloid receptor: a molecular gateway<br />
to the pain pathway. Annu. Rev. Neurosci.<br />
v. 24 p. 487–517.<br />
Gillespie, P. G. and Cyr, J. L., 2004. Myosin-<br />
1c, the hair <strong>cell</strong>’s adaptation motor. Annu.<br />
Rev. Physiol. v. 66 p. 521–545.<br />
References 637
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 638<br />
Lin, C. and Shalitin, D., 2003. Cryptochrome<br />
structure and signal transduction. Annu.<br />
Rev. Plant Biol. v. 54 p. 469–496.<br />
Rockwell, N.C., Su, Y.-S., and Lagarias, J.C.,<br />
2006. Phytochrome structure and <strong>signaling</strong><br />
mechanisms. Annu. Rev. Plant Biology,<br />
v. 57 p. 837–858.<br />
Sancar, A., 2000. Cryptochrome: the second<br />
photoactive pigment in the eye and its<br />
role in circadian photoreception. Annu.<br />
Rev. Biochem. v. 69 p. 31–67.<br />
14.3 Receptors sense diverse stimuli but initiate<br />
a limited repertoire <strong>of</strong> <strong>cell</strong>ular signals<br />
Research<br />
Klein, C., Paul, J. I., Sauvé, K., Schmidt, M.<br />
M., Arcangeli, L., Ransom, J., Trueheart,<br />
J., Manfredi, J. P., Broach, J. R., and<br />
Murphy, A. J., 1998. Identification <strong>of</strong> surrogate<br />
agonists for the human FPRL-1 receptor<br />
by autocrine selection in yeast.<br />
Nat. Biotechnol. v. 16 p. 1334–1337.<br />
14.5 Ligand binding changes receptor conformation<br />
Review<br />
Ross, E. M., and Kenakin, T. P., 2001.<br />
Pharmacodynamics: Mechanisms <strong>of</strong> drug<br />
action and the relationship between drug<br />
concentration and effects. In Goodman<br />
and Gilmans’s The Pharmacological Basis <strong>of</strong><br />
Therapeutics, 10th. Ed., J. G. Hardman and<br />
L. E. Limbird, eds., New York: McGraw-<br />
Hill, p. 31–43.<br />
14.6 Signals are sorted and integrated in <strong>signaling</strong><br />
pathways and networks<br />
Research<br />
Itzkovitz, S., Milo, R., Kashtan, N., Ziv, G.,<br />
and Alon, U., 2003. Subgraphs in random<br />
networks. Phys. Rev. E. v. 68 p. 126–127.<br />
14.7 Cellular <strong>signaling</strong> pathways can be<br />
thought <strong>of</strong> as biochemical logic circuits<br />
Review<br />
Milo, R., Shen-Orr, S., Itzkovitz, S., Kashtan,<br />
N., Chklovskii, D., and Alon, U., 2002.<br />
Network motifs: Simple building blocks <strong>of</strong><br />
complex networks. Science v. 298 p.<br />
824–827.<br />
Research<br />
Torres, E. and Rosen, M. K., 2003. Contingent<br />
phosphorylation/dephosphorylation provides<br />
a mechanism <strong>of</strong> molecular memory<br />
in WASP. Mol. Cell v. 11 p. 1215–1227.<br />
14.8 Scaffolds increase <strong>signaling</strong> efficiency<br />
and enhance spatial organization <strong>of</strong> <strong>signaling</strong><br />
Review<br />
Elion, E. A., 2001. The Ste5p scaffold. J. Cell<br />
Sci. v. 114 p. 3967–3978.<br />
Kholodenko, B. N., 2003. Four-dimensional<br />
organization <strong>of</strong> protein kinase <strong>signaling</strong><br />
cascades: the roles <strong>of</strong> diffusion, endocytosis<br />
and molecular motors. J. Exp. Biol. v.<br />
206 p. 2073–2082.<br />
O’Rourke, S. M., Herskowitz, I., and O’Shea,<br />
E. K., 2002. Yeast go the whole HOG for<br />
the hyperosmotic response. Trends Genet.<br />
v. 18 p. 405–412.<br />
Pawson, T. and Nash, P., 2003. Assembly <strong>of</strong><br />
<strong>cell</strong> regulatory systems through protein<br />
interaction domains. Science v. 300 p.<br />
445–452.<br />
Tsunoda, S. and Zuker, C. S., 1999. The organization<br />
<strong>of</strong> INAD-<strong>signaling</strong> complexes<br />
by a multivalent PDZ domain protein in<br />
Drosophila photoreceptor <strong>cell</strong>s ensures<br />
sensitivity and speed <strong>of</strong> <strong>signaling</strong>. Cell<br />
Calcium v. 26 p. 165–171.<br />
Research<br />
Levchenko, A., Bruck, J., and Sternberg, P.<br />
W., 2000. Scaffold proteins may biphasically<br />
affect the levels <strong>of</strong> mitogen-activated<br />
protein kinase <strong>signaling</strong> and reduce<br />
its threshold properties. Proc. Natl. Acad.<br />
Sci. USA v. 97 p. 5818–5823.<br />
Rensland, H., Lautwein, A., Wittingh<strong>of</strong>er, A.,<br />
and Goody, R. S., 1991. Is there a ratelimiting<br />
step before GTP cleavage by H-<br />
ras p21? Biochemistry v. 30 p. 11181–<br />
11185.<br />
Shenker, A., Goldsmith, P., Unson, C. G., and<br />
Spiegel, A. M., 1991. The G protein coupled<br />
to the thromboxane A2 receptor in<br />
human platelets is a member <strong>of</strong> the novel<br />
Gq family. J. Biol. Chem. v. 266 p. 9309–<br />
9313.<br />
14.9 Independent, modular domains specify<br />
protein-protein interactions<br />
Review<br />
Pawson, T. and Nash, P., 2003. Assembly <strong>of</strong><br />
<strong>cell</strong> regulatory systems through protein<br />
interaction domains. Science v. 300 p.<br />
445–452.<br />
Turk, B. E. and Cantley, L. C., 2003. Peptide<br />
libraries: at the crossroads <strong>of</strong> proteomics<br />
and bioinformatics. Curr. Opin. Chem. Biol.<br />
v. 7 p. 84–90.<br />
Research<br />
Ginty, D. D., Kornhauser, J. M., Thompson,<br />
M. A., Bading, H., Mayo, K. E.,<br />
Takahashi, J. S., and Greenberg, M. E.,<br />
1993. Regulation <strong>of</strong> CREB phosphorylation<br />
in the suprachiasmatic nucleus by<br />
light and a circadian clock. Science v. 260<br />
p. 238–241.<br />
638 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 639<br />
14.10 Cellular <strong>signaling</strong> is remarkably adaptive<br />
Review<br />
Perkins, J. P., Hausdorff, W. P., and Lefkowitz,<br />
R. J., 1990. Mechanisms <strong>of</strong> ligand-induced<br />
desensitization <strong>of</strong> -adrenergic receptors.<br />
In The Beta-Adrenergic Receptors, J.<br />
P. Perkins, ed., Clifton, NJ: Humana Press,<br />
p. 73–124.<br />
14.11 Signaling proteins are frequently expressed<br />
as multiple species<br />
Review<br />
Barnes, N. M., and Sharp, T., 1999. A review<br />
<strong>of</strong> central 5-HT receptors and their function.<br />
Neuropharmacology v. 38 p. 1083–<br />
1152.<br />
Gilman, A. G., 1987. G proteins: transducers<br />
<strong>of</strong> receptor-generated signals Annu. Rev.<br />
Biochem. v. 56 p. 615–649.<br />
Rebecchi, M. J., and Pentyala, S. N., 2000.<br />
Structure, function, and control <strong>of</strong> phosphoinositide-specific<br />
phospholipase C.<br />
Physiol. Rev. v. 80 p. 1291–1335.<br />
Sunahara, R. K. and Taussig, R., 2002.<br />
Is<strong>of</strong>orms <strong>of</strong> mammalian adenylyl cyclase:<br />
Multiplicities <strong>of</strong> <strong>signaling</strong>. Mol.<br />
Interventions v. 2 p. 168–184.<br />
14.14 Second messengers provide readily diffusible<br />
pathways for information transfer<br />
Review<br />
Beavo, J. A., Bechtel, P. J., and Krebs, E. G.,<br />
1975. Mechanisms <strong>of</strong> control for cAMPdependent<br />
protein kinase from skeletal<br />
muscle. Adv. Cyclic Nucleotide Res. v. 5 p.<br />
241–251.<br />
Kobe, B., Heierhorst, J., and Kemp, B. E.,<br />
1997. Intrasteric regulation <strong>of</strong> protein kinases.<br />
Adv. Second Messenger Phosphoprotein<br />
Res. v. 31 p. 29–40.<br />
Wong, W. and Scott, J. D., 2004. AKAP signalling<br />
complexes: focal points in space<br />
and time. Nat. Rev. Mol. Cell Biol. v. 5 p.<br />
959–970.<br />
Research<br />
Rall, T. W. and Sutherland, E. W., 1958.<br />
Formation <strong>of</strong> a cyclic adenine ribonucleotide<br />
by tissue particles. J. Biol. Chem. v.<br />
232 p. 1065–1076.<br />
14.15 Ca2+ <strong>signaling</strong> serves diverse purposes in<br />
all eukaryotic <strong>cell</strong>s<br />
Review<br />
Newton, A. C., 2001. Protein kinase C: structural<br />
and spatial regulation by phosphorylation,<br />
c<strong>of</strong>actors, and macromolecular<br />
interactions. Chem. Rev. v. 101 p.<br />
2353–2364.<br />
Research<br />
Clapperton, J. A., Martin, S. R., Smerdon, S.<br />
J., Gamblin, S. J., and Bayley, P. M.,<br />
2002. Structure <strong>of</strong> the complex <strong>of</strong><br />
calmodulin with the target sequence <strong>of</strong><br />
calmodulin dependent protein kinase I:<br />
Studies <strong>of</strong> the kinase activation mechanism.<br />
Biochemistry. v. 41 p. 14669–14679.<br />
Kuboniwa, H., Tjandra, N., Grzesiek, S., Ren,<br />
H., Klee, C.B., Bax, A., 1995. Solution<br />
structure <strong>of</strong> calcium-free calmodulin. Nat.<br />
Struct. Biol. v. 2 p. 768–776.<br />
14.16 Lipids and lipid-derived compounds are<br />
<strong>signaling</strong> molecules<br />
Review<br />
Rebecchi, M. J., and Pentyala, S. N., 2000.<br />
Structure, function, and control <strong>of</strong> phosphoinositide-specific<br />
phospholipase C.<br />
Physiol. Rev. v. 80 p. 1291–1335.<br />
Yang, C. and Kazanietz, M. G., 2003.<br />
Divergence and complexities in DAG <strong>signaling</strong>:<br />
looking beyond PKC. Trends<br />
Pharmacol. Sci. v. 24 p. 602–608.<br />
14.17 PI 3-kinase synthesizes a lipid activator<br />
<strong>of</strong> <strong>cell</strong> motion and shape change<br />
Review<br />
Downward, J., 2004. PI 3-kinase, Akt and <strong>cell</strong><br />
survival. Semin. Cell Dev. Biol. v. 15 p.<br />
177–182.<br />
Lawlor, M. A. and Alessi, D. R., 2001.<br />
PKB/Akt: a key mediator <strong>of</strong> <strong>cell</strong> proliferation,<br />
survival and insulin responses? J.<br />
Cell Sci. v. 114 p. 2903–2910.<br />
Van Haastert, P. J. and Devreotes, P. N. 2004.<br />
Chemotaxis: <strong>signaling</strong> the way forward.<br />
Nat. Rev. Mol. Cell Biol. v. 5. p. 626–634.<br />
14.18 Signaling through ion channel receptors<br />
is very fast<br />
Review<br />
Clapham, D. E., 2003. TRP channels as <strong>cell</strong>ular<br />
sensors. Nature v. 426 p. 517–524.<br />
Corey, D. P., 2003. New TRP channels in<br />
hearing and mechanosensation. Neuron v.<br />
39 p. 585–588.<br />
Hille, B, 1992. Ionic channels <strong>of</strong> excitable<br />
membranes. Sinauer Associates.<br />
Siegelbaum, S. A. and Koester, J., 2000. Ion<br />
channels and Membrane potential, in<br />
<strong>Principles</strong> <strong>of</strong> Neural Science, 4th. Ed., E. R.<br />
Kandel, J. H. Schwartz, and T. M. Jessell,<br />
eds., New York: McGraw-Hill. p.<br />
105–139.<br />
Research<br />
Unwin, N., 2005. Refined structure <strong>of</strong> the<br />
nicotinic acetylcholine receptor at 4Å resolution.<br />
J. Mol. Biol. v. 346 p. 967.<br />
14.19 Nuclear receptors regulate transcription<br />
References 639
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 640<br />
Review<br />
Mangelsdorf, D. J. et al., 1995. The nuclear<br />
receptor superfamily: the second decade.<br />
Cell v. 83 p. 835–839.<br />
Smith, C. L. and O’Malley, B. W., 2004.<br />
Coregulator function: a key to understanding<br />
tissue specificity <strong>of</strong> selective receptor<br />
modulators. Endocr. Rev. v. 25 p.<br />
45–71.<br />
Research<br />
Brzozowski A. M., Pike, A. C., Dauter, Z.,<br />
Hubbard, R. E., Bonn, T., Engstrom, O.,<br />
Ohman, L., Green, G. L., Gustafsson, J. A.,<br />
and Carlquist, M., 1997. Molecular basis<br />
<strong>of</strong> agonism and antagonism in the oestrogen<br />
receptor. Nature v. 389 p. 753–758.<br />
14.20 G protein <strong>signaling</strong> modules are widely<br />
used and highly adaptable<br />
Review<br />
Clapham, D. E. and Neer, E. J., 1997. G protein<br />
subunits. Annu. Rev. Pharmacol.<br />
Toxicol. v. 37 p. 167–203.<br />
Filipek, S., Teller, D. C., Palczewski, K., and<br />
Stenkamp, R., 2003. The crystallographic<br />
model <strong>of</strong> rhodopsin and its use in studies<br />
<strong>of</strong> other G protein-coupled receptors.<br />
Annu. Rev. Biophys. Biomol. Struct. v. 32 p.<br />
375–397.<br />
Ross, E. M. and Wilkie, T. M., 2000. GTPaseactivating<br />
proteins for heterotrimeric G<br />
proteins: Regulators <strong>of</strong> G protein <strong>signaling</strong><br />
(RGS) and RGS-like proteins. Annu.<br />
Rev. Biochem. v. 69 p. 795–827.<br />
Ross, E. M., 1989. Signal sorting and amplification<br />
through G protein-coupled receptors.<br />
Neuron v. 3 p. 141–152.<br />
Sprang, S. R., 1997. G proteins, effectors and<br />
GAPs: structure and mechanism. Curr.<br />
Opin. Struct. Biol. v. 7 p. 849–856.<br />
Sprang, S. R., 1997. G protein mechanisms:<br />
Insights from structural analysis. Annu.<br />
Rev. Biochem. v. 66 p. 639–678.<br />
Research<br />
Benke, T., Motoshima, C. A., Fox, H., Le<br />
Trong, B. A., Teller, I., Okada, D. C.,<br />
Stenkamp, T., Yamamoto, R. E., and<br />
Miyano, M., 2000. Crystal structure <strong>of</strong><br />
rhodopsin: A G protein-coupled receptor.<br />
Science v. 289 p. 739–745.<br />
Chen, C. -K., Burns, M. E., He, W., Wensel, T.<br />
G., Baylor, D. A., and Simon, M. I., 2000.<br />
Slowed recovery <strong>of</strong> rod photoresponse in<br />
mice lacking the GTPase acceleration protein<br />
RGS9-1. Nature, v. 403 p. 557–560.<br />
Palczewski, K., Kumasaka, T., Hori, T.,<br />
Behnke , C. A., Motoshima, H., Fox, B.<br />
A., Le Trong, I., Teller, D. C., Okada, T.,<br />
Stenkamp, R. E., Yamamoto, M., and<br />
Miyano, M., 2000. Crystal structure <strong>of</strong><br />
rhodopsin: a G protein-coupled receptor.<br />
Science v. 289 p. 739–745.<br />
Wall, M. A., Coleman, D. E., Lee, E., Iniguez-<br />
Lluhi, J. A., Posner, B. A., Gilman, A. G.,<br />
and Sprang, S. R., 1995. The structure <strong>of</strong><br />
the G protein heterotrimer G i1<br />
1<br />
2<br />
. Cell<br />
v. 83 p. 1047–1058.<br />
14.23 Small, monomeric GTP-binding proteins<br />
are multiuse switches<br />
Review<br />
Bishop, A. L. and Hall, A., 2000. Rho GTPases<br />
and their effector proteins. Biochem. J. v.<br />
348 pt. 2 p. 241–255.<br />
Kuersten, S., Ohno, M., and Mattaj, I. W.,<br />
2001. Nucleocytoplasmic transport: Ran,<br />
beta and beyond. Trends Cell Biol. v. 11 p.<br />
497–503.<br />
Takai, Y., Sasaki, T., and Matozaki, T., 2001.<br />
Small GTP-binding proteins. Physiol. Rev.<br />
v. 81 p. 153–208.<br />
14.24 Protein phosphorylation/dephosphorylation<br />
is a major regulatory mechanism in<br />
the <strong>cell</strong><br />
Review<br />
Cohen, S., 1983. The epidermal growth factor<br />
(EGF). Cancer v. 51 p. 1787–1791.<br />
Fischer, E. H., 1997. “Protein phosphorylation<br />
and <strong>cell</strong>ular regulation, II,” in Nobel<br />
Lectures, Physiology or Medicine 1991–1995,<br />
N., Ringertz, Ed., Singapore: World<br />
Scientific Publishing Co.<br />
Krebs, G., 1993. Protein phosphorylation and<br />
<strong>cell</strong>ular regulation. Bioscience Reports v. 13<br />
p. 127–142.<br />
Manning, G., Whyte, D. B., Martinez, R.,<br />
Hunter, T., and Sudarsanam, S., 2002.<br />
The protein kinase complement <strong>of</strong> the<br />
human genome. Science v. 298 p.<br />
1912–1934.<br />
Newton, A. C., 2003. Regulation <strong>of</strong> the ABC<br />
kinases by phosphorylation: protein kinase<br />
C as a paradigm. Biochem. J. v. 370 p.<br />
361–371.<br />
Nolen, B., Taylor, S., and Ghosh, G., 2004.<br />
Regulation <strong>of</strong> protein kinases; controlling<br />
activity through activation segment conformation.<br />
Mol. Cell v. 15 p. 661–675.<br />
Turk, B. E. and Cantley, L. C., 2003. Peptide<br />
libraries: at the crossroads <strong>of</strong> proteomics<br />
and bioinformatics. Curr. Opin. Chem. Biol.<br />
v. 7 p. 84–90.<br />
Research<br />
Canagarajah, B. J., Khokhlatchev, A., Cobb,<br />
M. H., and Goldsmith, E. J., 1997.<br />
Activation mechanism <strong>of</strong> the MAP kinase<br />
ERK2 by dual phosphorylation. Cell v. 90<br />
p. 859–869.<br />
Ginty, D. D., Kornhauser, J. M., Thompson,<br />
640 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 641<br />
M. A., Bading, H., Mayo, K. E.,<br />
Takahashi, J. S., and Greenberg, M. E.,<br />
1993. Regulation <strong>of</strong> CREB phosphorylation<br />
in the suprachiasmatic nucleus by<br />
light and a circadian clock. Science v. 260<br />
p. 238–241.<br />
Knighton, D. R., Zheng, J. H., Ten Eyck, L. F.,<br />
Ashford, V. A., Xuong, N. H., Taylor, S. S.,<br />
and Sowadski, J. M., 1991. Crystal structure<br />
<strong>of</strong> the catalytic subunit <strong>of</strong> cyclic<br />
adenosine monophosphate-dependent<br />
protein kinase. Science v. 253 p. 407–414.<br />
Taylor, S. S., Radzio-Andzelm, E., and Hunter,<br />
T., 1995. How do protein kinases discriminate<br />
between serine/threonine and tyrosine?<br />
Structural insights from the insulin<br />
receptor protein-tyrosine kinase. FASEB<br />
J. v. 9 p. 1255–1266.<br />
Zhang, F., Strand, A., Robbins, D., Cobb, M.<br />
H., and Goldsmith, E. J., 1994. Atomic<br />
structure <strong>of</strong> the MAP kinase ERK2 at 2.3<br />
Å resolution. Nature v. 367 p. 704–711.<br />
14.25 Two-component protein phosphorylation<br />
systems are <strong>signaling</strong> relays<br />
Review<br />
Hoch, J. A, and Silhavy, T. J., eds., 1995. Twocomponent<br />
signal transduction.<br />
Washington, D. C.:American Society for<br />
Microbiology.<br />
Stock, A. M., Robinson, V. L., and Goudreau,<br />
P. N., 2000. Two-component signal transduction.<br />
Annu. Rev. Biochem. v. 69 p.<br />
183–215.<br />
14.26 Pharmacological inhibitors <strong>of</strong> protein kinases<br />
may be used to understand and<br />
treat disease<br />
Review<br />
Blume-Jensen, P, and Hunter, T., 2001.<br />
Oncogenic kinase signalling. Nature v.<br />
411 p. 355–365.<br />
Cherry, M. and Williams, D. H., 2004. Recent<br />
kinase and kinase inhibitor X-ray structures:<br />
mechanisms <strong>of</strong> inhibition and selectivity<br />
insights. Curr. Med. Chem. v. 11 p.<br />
663–673.<br />
Cohen, P., 2002. Protein kinases—the major<br />
drug targets <strong>of</strong> the twenty-first century?<br />
Nat. Rev. Drug Discov. v. 1 p. 309–315.<br />
Davies, S. P., Reddy, H., Caivano, M., and<br />
Cohen, P., 2000. Specificity and mechanism<br />
<strong>of</strong> action <strong>of</strong> some commonly used<br />
protein kinase inhibitors. Biochem. J. v.<br />
351 p. 95–105.<br />
Tibes, R., Trent, J., and Kurzrock, R., 2005.<br />
Tyrosine kinase inhibitors and the dawn<br />
<strong>of</strong> molecular cancer therapeutics. Annu.<br />
Rev. Pharmacol. Toxicol. v. 45 p. 357–384.<br />
14.27 Phosphoprotein phosphatases reverse the<br />
actions <strong>of</strong> kinases and are independently<br />
regulated<br />
Review<br />
Aramburu, J., Heitman, J., and Crabtree, G.<br />
R., 2004. Calcineurin: a central controller<br />
<strong>of</strong> signalling in eukaryotes. EMBO Rep. v.<br />
5 p. 343–348.<br />
Dounay, A. B. and Forsyth, C. J., 2002.<br />
Okadaic acid: the archetypal serine/threonine<br />
protein phosphatase inhibitor. Curr.<br />
Med. Chem. v. 9 p. 1939–1980.<br />
Neel, B. G., Gu, H., and Pao, L., 2003. The<br />
‘Shp’ing news: SH2 domain-containing<br />
tyrosine phosphatases in <strong>cell</strong> <strong>signaling</strong>.<br />
Trends Biochem. Sci. v. 28 p. 284–293.<br />
Neely, K. E. and Piwnica-Worms, H., 2003.<br />
Cdc25A regulation: to destroy or not to<br />
destroy—is that the only question? Cell<br />
Cycle v. 2 p. 455–457.<br />
Olson, E. N. and Williams, R. S., 2000.<br />
Remodeling muscles with calcineurin.<br />
Bioessays v. 22 p. 510–519.<br />
Virshup, D. M., 2000. Protein phosphatase<br />
2A: a panoply <strong>of</strong> enzymes. Curr. Opin. Cell<br />
Biol. v. 12 p. 180–185.<br />
Research<br />
Sun, H., et al. 1993. MKP-1 (3CH134), an immediate<br />
early gene product, is a dual<br />
specificity phosphatase that dephosphorylates<br />
MAP kinase in vivo Cell v. 75 p.<br />
487–493.<br />
Terrak, M., Kerff, F., Langsetmo, K., Tao, T.,<br />
and Dominguez, R., 2004. Structural basis<br />
<strong>of</strong> protein phosphatase 1 regulation.<br />
Nature v. 429 p. 780–784.<br />
14.28 Covalent modification by ubiquitin and<br />
ubiquitin-like proteins is another way <strong>of</strong><br />
regulating protein function<br />
Review<br />
Gill, G., 2004. SUMO and ubiquitin in the nucleus:<br />
different functions, similar mechanisms?<br />
Genes Dev. v. 18 p. 2046–2059.<br />
Pickart, C. M. and Eddins, M. J., 2004.<br />
Ubiquitin: structures, functions, mechanisms.<br />
Biochim. Biophys. Acta v. 1695 p.<br />
55–72.<br />
Research<br />
Dharmasiri, N., Dharmasiri, S., and Estelle,<br />
M., 2005. The F-box protein TIR1 is an<br />
auxin receptor. Nature v. 435 p. 441–445.<br />
Kanayama, A. et al. 2004. TAB2 and TAB3<br />
activate the NF-B pathway through<br />
binding to polyubiquitin chains. Mol. Cell<br />
v. 15 p. 535–548<br />
Kepinski, S. and Leyser, O., 2005. The<br />
Arabidopsis F-box protein TIR1 is an<br />
auxin receptor. Nature v. 435 p. 446–451.<br />
References 641
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 642<br />
14.29 The Wnt pathway regulates <strong>cell</strong> fate during<br />
development and other processes in<br />
the adult<br />
Review<br />
Logan, C. Y. and Nusse, R., 2004. The WNT<br />
<strong>signaling</strong> pathway in development and<br />
disease. Annu. Rev. Cell. Dev. Biol. v. 20, p.<br />
781–810.<br />
Tolwinski, N. S., Wieschaus, E., 2004.<br />
Rethinking WNT <strong>signaling</strong>. Trends Genet.<br />
v. 20 p. 177–81.<br />
14.30 Diverse <strong>signaling</strong> mechanisms are regulated<br />
by protein tyrosine kinases<br />
Review<br />
Birge, R. B., Knudsen, B. S., Besser, D., and<br />
Hanafusa, H., 1996. SH2 and SH3-containing<br />
adaptor proteins: redundant or<br />
independent mediators <strong>of</strong> intra<strong>cell</strong>ular<br />
signal transduction. Genes Cells v. 1 p.<br />
595–613.<br />
Blume-Jensen, P. and Hunter, T., 2001.<br />
Oncogenic kinase signalling. Nature v.<br />
411 p. 355–365.<br />
Cohen, P., 2002. Protein kinases—the major<br />
drug targets <strong>of</strong> the twenty-first century?<br />
Nat. Rev. Drug Discov. v. 1 p. 309–315.<br />
Pawson, T. and Nash, P., 2003. Assembly <strong>of</strong><br />
<strong>cell</strong> regulatory systems through protein<br />
interaction domains. Science v. 300 p.<br />
445–452.<br />
Tallquist, M. and Kazlauskas, A., 2004. PDGF<br />
<strong>signaling</strong> in <strong>cell</strong>s and mice. Cytokine<br />
Growth Factor Rev. v. 15 p. 205–213.<br />
Taniguchi, C. M., Emanuelli, B., and Kahn, C.<br />
R., 2006. Critical nodes in signalling<br />
pathways: insights into insulin action.<br />
Nat. Rev. Mol. Cell Biol. v. 7, p. 85–96.<br />
14.31 Src family protein kinases cooperate with<br />
receptor protein tyrosine kinases<br />
Review<br />
Boggon, T. J. and Eck, M. J., 2004. Structure<br />
and regulation <strong>of</strong> Src family kinases.<br />
Oncogene v. 23 p. 7918–7927.<br />
Frame, M. C., 2004. Newest findings on the<br />
oldest oncogene; how activated src does<br />
it. J. Cell Sci. v. 117 p. 989–998.<br />
Research<br />
Cowan-Jacob, S. W., Frendrich, G., Manley, P.<br />
W., Jahnke, W., Fabbro, D.,<br />
Liebetanz, J., Meyer, T. 2005. The<br />
crystal structure <strong>of</strong> a c-Src complex in<br />
an active conformation suggests possible<br />
steps in c-Src activation Structure<br />
v. 13 p. 861–871.<br />
Xu, W., Harrison, S. C., and Eck, M. J., 1997.<br />
Three-dimensional structure <strong>of</strong> the tyro-<br />
sine kinase c-Src. Nature v. 385 p.<br />
595–602.<br />
14.32 MAPKs are central to many <strong>signaling</strong><br />
pathways<br />
Review<br />
Chen, Z., Gibson, T. B., Robinson, F.,<br />
Silvestro, L., Pearson, G., Xu, B., Wright,<br />
A., Vanderbilt, C., and Cobb, M. H., 2001.<br />
MAP kinases. Chem. Rev. v. 101 p.<br />
2449–2476.<br />
Lewis, T. S., Shapiro, P. S., and Ahn, N. G.,<br />
1998. Signal transduction through MAP<br />
kinase cascades. Adv. Cancer Res. v. 74 p.<br />
49–139.<br />
Morrison, D. K. and Davis, R. J., 2003.<br />
Regulation <strong>of</strong> MAP kinase <strong>signaling</strong> modules<br />
by scaffold proteins in mammals.<br />
Annu. Rev. Cell Dev. Biol. v. 19 p. 91–118.<br />
Sharrocks, A. D., Yang, S. H., and Galanis, A.,<br />
2000. Docking domains and substratespecificity<br />
determination for MAP kinases.<br />
Trends Biochem. Sci. v. 25 p.<br />
448–453.<br />
Research<br />
Anderson, N. G., Maller, J. L., Tonks, N. K.,<br />
and Sturgill, T. W., 1990. Requirement<br />
for integration <strong>of</strong> signals from two distinct<br />
phosphorylation pathways for activation<br />
<strong>of</strong> MAP kinase. Nature v. 343 p.<br />
651–653.<br />
14.33 Cyclin-dependent protein kinases control<br />
the <strong>cell</strong> cycle<br />
Review<br />
Dorée, M. and Hunt, T., 2002. From Cdc2 to<br />
Cdk1: when did the <strong>cell</strong> cycle kinase join<br />
its cyclin partner? J. Cell Sci. v. 115 p.<br />
2461–2464.<br />
Hartwell, L. H., 1991. Twenty-five years <strong>of</strong><br />
<strong>cell</strong> cycle genetics. Genetics v. 129 p.<br />
975–980.<br />
Neely, K. E. and Piwnica-Worms, H., 2003.<br />
Cdc25A regulation: to destroy or not to<br />
destroy—is that the only question? Cell<br />
Cycle v. 2 p. 455–457.<br />
Nurse, P., 2000. A long twentieth century <strong>of</strong><br />
the <strong>cell</strong> cycle and beyond. Cell v. 100 p.<br />
71–78.<br />
Research<br />
Pavletich, N. P., 1999. Mechanisms <strong>of</strong> cyclindependent<br />
kinase regulation: structures<br />
<strong>of</strong> Cdks, their cyclin activators, and Cip<br />
and INK4 inhibitors. J. Mol. Biol. v. 287 p.<br />
821–828.<br />
Russo, A. A., Jeffrey, P. D., and Pavletich, N.<br />
P., 1996. Structural basis <strong>of</strong> cyclin-dependent<br />
kinase activation by phosphorylation.<br />
Nat. Struct. Biol. v. 3 p. 696–700.<br />
642 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 643<br />
14.34 Diverse receptors recruit protein tyrosine<br />
kinases to the plasma membrane<br />
Review<br />
Ernst, M. and Jenkins, B. J., 2004. Acquiring<br />
signalling specificity from the cytokine<br />
receptor gp130. Trends Genet. v. 20 p.<br />
23–32.<br />
Herrington, J. and Carter-Su, C., 2001.<br />
Signaling pathways activated by the<br />
growth hormone receptor. Trends<br />
Endocrinol. Metab. v. 12 p. 252–257.<br />
Levy, D. E. and Darnell, J. E., 2002. Stats:<br />
transcriptional control and biological impact.<br />
Nat. Rev. Mol. Cell Biol. v. 3 p.<br />
651–662.<br />
Miranti, C. K. and Brugge, J. S., 2002.<br />
Sensing the environment: a historical<br />
perspective on integrin signal transduction.<br />
Nat. Cell Biol. v. 4 p. E83–E90.<br />
Palacios, E. H. and Weiss, A., 2004. Function<br />
<strong>of</strong> the Src-family kinases, Lck and Fyn, in<br />
T-<strong>cell</strong> development and activation.<br />
Oncogene v. 23 p. 7990–8000.<br />
Research<br />
Clackson, T. and Wells, J. A., 1995. A hot spot<br />
<strong>of</strong> binding energy in a hormone-receptor<br />
interface. Science v. 267 p. 383–386.<br />
References 643