Principles of cell signaling - UT Southwestern
Principles of cell signaling - UT Southwestern
Principles of cell signaling - UT Southwestern
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
39057_ch14_<strong>cell</strong>bio.qxd 8/28/06 5:11 PM Page 612<br />
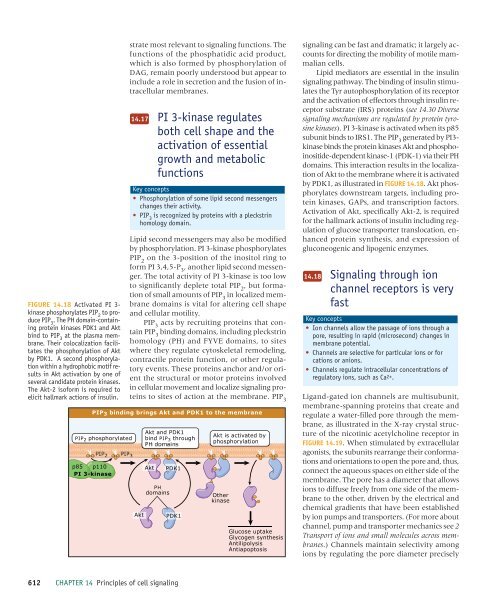
FIGURE 14.18 Activated PI 3-<br />
kinase phosphorylates PIP 2<br />
to produce<br />
PIP 3<br />
. The PH domain-containing<br />
protein kinases PDK1 and Akt<br />
bind to PIP 3<br />
at the plasma membrane.<br />
Their colocalization facilitates<br />
the phosphorylation <strong>of</strong> Akt<br />
by PDK1. A second phosphorylation<br />
within a hydrophobic motif results<br />
in Akt activation by one <strong>of</strong><br />
several candidate protein kinases.<br />
The Akt-2 is<strong>of</strong>orm is required to<br />
elicit hallmark actions <strong>of</strong> insulin.<br />
p85 p110<br />
PI 3-kinase<br />
PIP 2 PIP 3<br />
strate most relevant to <strong>signaling</strong> functions. The<br />
functions <strong>of</strong> the phosphatidic acid product,<br />
which is also formed by phosphorylation <strong>of</strong><br />
DAG, remain poorly understood but appear to<br />
include a role in secretion and the fusion <strong>of</strong> intra<strong>cell</strong>ular<br />
membranes.<br />
14.17<br />
PI 3-kinase regulates<br />
both <strong>cell</strong> shape and the<br />
activation <strong>of</strong> essential<br />
growth and metabolic<br />
functions<br />
Key concepts<br />
• Phosphorylation <strong>of</strong> some lipid second messengers<br />
changes their activity.<br />
• PIP 3<br />
is recognized by proteins with a pleckstrin<br />
homology domain.<br />
Lipid second messengers may also be modified<br />
by phosphorylation. PI 3-kinase phosphorylates<br />
PIP 2<br />
on the 3-position <strong>of</strong> the inositol ring to<br />
form PI 3,4,5-P 3<br />
, another lipid second messenger.<br />
The total activity <strong>of</strong> PI 3-kinase is too low<br />
to significantly deplete total PIP 2<br />
, but formation<br />
<strong>of</strong> small amounts <strong>of</strong> PIP 3<br />
in localized membrane<br />
domains is vital for altering <strong>cell</strong> shape<br />
and <strong>cell</strong>ular motility.<br />
PIP 3<br />
acts by recruiting proteins that contain<br />
PIP 3<br />
binding domains, including pleckstrin<br />
homology (PH) and FYVE domains, to sites<br />
where they regulate cytoskeletal remodeling,<br />
contractile protein function, or other regulatory<br />
events. These proteins anchor and/or orient<br />
the structural or motor proteins involved<br />
in <strong>cell</strong>ular movement and localize <strong>signaling</strong> proteins<br />
to sites <strong>of</strong> action at the membrane. PIP 3<br />
PIP 3 binding brings Akt and PDK1 to the membrane<br />
PIP 2 phosphorylated<br />
Akt<br />
Akt and PDK1<br />
bind PIP3 through<br />
PH domains<br />
Akt<br />
PH<br />
domains<br />
PDK1<br />
PDK1<br />
Akt is activated by<br />
phosphorylation<br />
Other<br />
kinase<br />
Glucose uptake<br />
Glycogen synthesis<br />
Antilipolysis<br />
Antiapoptosis<br />
<strong>signaling</strong> can be fast and dramatic; it largely accounts<br />
for directing the mobility <strong>of</strong> motile mammalian<br />
<strong>cell</strong>s.<br />
Lipid mediators are essential in the insulin<br />
<strong>signaling</strong> pathway. The binding <strong>of</strong> insulin stimulates<br />
the Tyr autophosphorylation <strong>of</strong> its receptor<br />
and the activation <strong>of</strong> effectors through insulin receptor<br />
substrate (IRS) proteins (see 14.30 Diverse<br />
<strong>signaling</strong> mechanisms are regulated by protein tyrosine<br />
kinases). PI 3-kinase is activated when its p85<br />
subunit binds to IRS1. The PIP 3<br />
generated by PI3-<br />
kinase binds the protein kinases Akt and phosphoinositide-dependent<br />
kinase-1 (PDK-1) via their PH<br />
domains. This interaction results in the localization<br />
<strong>of</strong> Akt to the membrane where it is activated<br />
by PDK1, as illustrated in FIGURE 14.18. Akt phosphorylates<br />
downstream targets, including protein<br />
kinases, GAPs, and transcription factors.<br />
Activation <strong>of</strong> Akt, specifically Akt-2, is required<br />
for the hallmark actions <strong>of</strong> insulin including regulation<br />
<strong>of</strong> glucose transporter translocation, enhanced<br />
protein synthesis, and expression <strong>of</strong><br />
gluconeogenic and lipogenic enzymes.<br />
14.18<br />
Signaling through ion<br />
channel receptors is very<br />
fast<br />
Key concepts<br />
• Ion channels allow the passage <strong>of</strong> ions through a<br />
pore, resulting in rapid (microsecond) changes in<br />
membrane potential.<br />
• Channels are selective for particular ions or for<br />
cations or anions.<br />
• Channels regulate intra<strong>cell</strong>ular concentrations <strong>of</strong><br />
regulatory ions, such as Ca2+.<br />
Ligand-gated ion channels are multisubunit,<br />
membrane-spanning proteins that create and<br />
regulate a water-filled pore through the membrane,<br />
as illustrated in the X-ray crystal structure<br />
<strong>of</strong> the nicotinic acetylcholine receptor in<br />
FIGURE 14.19. When stimulated by extra<strong>cell</strong>ular<br />
agonists, the subunits rearrange their conformations<br />
and orientations to open the pore and, thus,<br />
connect the aqueous spaces on either side <strong>of</strong> the<br />
membrane. The pore has a diameter that allows<br />
ions to diffuse freely from one side <strong>of</strong> the membrane<br />
to the other, driven by the electrical and<br />
chemical gradients that have been established<br />
by ion pumps and transporters. (For more about<br />
channel, pump and transporter mechanics see 2<br />
Transport <strong>of</strong> ions and small molecules across membranes.)<br />
Channels maintain selectivity among<br />
ions by regulating the pore diameter precisely<br />
612 CHAPTER 14 <strong>Principles</strong> <strong>of</strong> <strong>cell</strong> <strong>signaling</strong>