- Page 6: Retrotransposons of rice as a tool
- Page 11 and 12: First, the Rice Genetics Cooperativ
- Page 13 and 14: OverviewRice genetics from Mendel t
- Page 15 and 16: Rice genetics from Mendelto functio
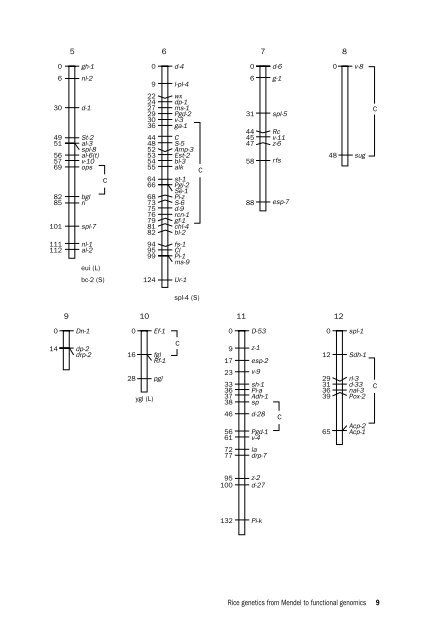
- Page 17 and 18: nese cultivar Nipponbare. The chrom
- Page 19: Gene symbolization in riceIn the ab
- Page 27 and 28: Table 2 continued.Gene Trait Chromo
- Page 29 and 30: plant resistance genes but is quite
- Page 31 and 32: and the information is freely avail
- Page 33: Hayano-Saito Y, Saito K, Nakamura S
- Page 36 and 37: Toriyama K, Arimoto Y, Uchimiya H,
- Page 39 and 40: Application of Mendelian geneticsin
- Page 41 and 42: In temperate regions, breeders have
- Page 43 and 44: Elongated uppermost internodeA gene
- Page 45 and 46: ecent years, Katy has been used as
- Page 47 and 48: Childs N. 1999. Rice situation and
- Page 49 and 50: Rutger JN, Carnahan HL. 1981. A fou
- Page 51 and 52: The Rockefeller Foundation’sInter
- Page 53 and 54: generated from a protoplast in 1986
- Page 55 and 56: Table 1. The Rockefeller Foundation
- Page 57 and 58: Box no. 1The saga of Xa21THE STORY
- Page 59 and 60: viruses, molecular tools made possi
- Page 61 and 62: BSLLBTETBTBox no. 2Rice—the pivot
- Page 63 and 64: Table 2. Types of institutions supp
- Page 65 and 66: Table 3. Breakdown of number of ins
- Page 67 and 68: ●●●Transformation of chloropl
- Page 69 and 70: Lander ES, Weinberg RA. 2000. Genom
- Page 71:
Zhang J, Zheng HG, Ali ML, Tripathy
- Page 74 and 75:
62 Morishima
- Page 76 and 77:
defined as the diploid species havi
- Page 78 and 79:
Interspecific relatedness was also
- Page 80 and 81:
Within the geographical range of O.
- Page 82 and 83:
Search for the immediate ancestor o
- Page 84 and 85:
gene pool, we performed QTL analysi
- Page 86 and 87:
Table 3. Changes in population para
- Page 88 and 89:
Kadowaki K, Yazaki K, Osumi T, Hara
- Page 91 and 92:
Rice: a central genomefor the genet
- Page 93 and 94:
Gibberellicacid-insensitiveRiceFoxt
- Page 95 and 96:
parative organization of the Pooide
- Page 97 and 98:
clear alignment of major factors co
- Page 99 and 100:
Gale MD, Devos KM. 1998. Plant comp
- Page 101 and 102:
Phylogeny of the genusOryza as reve
- Page 103 and 104:
Table 1. Species recognized in Oryz
- Page 105 and 106:
However, the subdivisional classifi
- Page 107 and 108:
Multiple gene trees: new insights i
- Page 109 and 110:
AABBBBCCCCCCDDDDKKHHKKFFHHHHJJJJGGF
- Page 111 and 112:
group with the diploid E genome spe
- Page 113 and 114:
genomes occupy the basal positions
- Page 115 and 116:
Donoghue MJ. 1994. Progress and pro
- Page 117:
Vaughan DA. 1994. The wild relative
- Page 120 and 121:
The existence of TEs also demonstra
- Page 122 and 123:
S. Wessler, unpublished data). Othe
- Page 124 and 125:
parent but not in the other can be
- Page 126 and 127:
ReferencesBaker B, Schell J, Lorz H
- Page 128 and 129:
Walker EL, Eggleston WB, Demopulos
- Page 130 and 131:
hypervariability make them valuable
- Page 132 and 133:
periments tended to detect only the
- Page 134 and 135:
Distribution of SSR motifsSignifica
- Page 136:
Association between SSRs and transp
- Page 140 and 141:
Molecular wt (bp)15 cultivars241 cu
- Page 142 and 143:
20222426283032343638404244464850525
- Page 144 and 145:
Blair MW, Hedetale V, McCouch SR. 2
- Page 146 and 147:
Semon M, Coburn J, Cadalen T, Jones
- Page 149 and 150:
Molecular mapping and markerassiste
- Page 151 and 152:
symposium (Li, this volume), here w
- Page 153 and 154:
Table 2 continued.Trait Gene Chromo
- Page 155 and 156:
markers were used for the other two
- Page 157 and 158:
volume). Those SSRs that can be sco
- Page 159 and 160:
Fukuoka S, Namai H, Okuno K. 1998.
- Page 161 and 162:
Naqvi NI, Chattoo BB. 1996. Molecul
- Page 163:
Zhang JS, Xie C, Li ZY, Chen SY. 19
- Page 166 and 167:
1980s have had far-reaching effects
- Page 168 and 169:
Table 2. The number of detected mai
- Page 170 and 171:
Table 3. Relative importance of mai
- Page 172 and 173:
Table 6. Some nonenvironment-specif
- Page 174 and 175:
On average, QTL main effects were d
- Page 176 and 177:
Table 9. Main-effect QTLs affecting
- Page 178 and 179:
Gene action of QTLsIn QTL mapping s
- Page 180 and 181:
tion of the parental lines; (3) enf
- Page 182 and 183:
Li ZK, Paterson AH, Pinson SRM, Sta
- Page 185 and 186:
Genetic and molecular basisof heter
- Page 187 and 188:
in Table 1 for the four traits were
- Page 189 and 190:
Table 2. A summary of the significa
- Page 191 and 192:
hybrids rather than progenies of se
- Page 193 and 194:
Correlation between gene expression
- Page 195 and 196:
Table 7. Numbers of cDNAs showing v
- Page 197:
Saghai Maroof MA, Yang GP, Zhang Q,
- Page 200 and 201:
188 Sasaki et al
- Page 202 and 203:
The boom in rice genome analysis is
- Page 204 and 205:
Selected PAC/BAC clones are culture
- Page 206 and 207:
an overestimation of the gene numbe
- Page 208 and 209:
Saji S, Umehara Y, Antonio BA, Yama
- Page 210 and 211:
BackgroundIn the random shotgun pha
- Page 212 and 213:
200 de la Bastide et alFig. 2. Scre
- Page 214 and 215:
a different sequence or contig. Fig
- Page 216 and 217:
ing annotation must be tagged accor
- Page 218 and 219:
slip in these regions, producing se
- Page 220 and 221:
sequencing method for us on cDNA cl
- Page 222 and 223:
Direct repeats Inverted repeats Tan
- Page 224 and 225:
iopoldat.html). The 2-kb threshold
- Page 226 and 227:
AppendixFinisher’s Clone Submissi
- Page 228 and 229:
The CUGI Rice Genome Framework Proj
- Page 230 and 231:
the development of a fingerprint DB
- Page 232 and 233:
First inriceHexa3%Mono4%Penta12%Di2
- Page 234 and 235:
Contigs2712722738652742755814257782
- Page 236 and 237:
Table 4. CCW sequencing status for
- Page 239 and 240:
Naturally occurring allelic variati
- Page 241 and 242:
in rice. To this end, two high-dens
- Page 243 and 244:
Characterization of genes at QTLs u
- Page 245 and 246:
In summary, in QTL analysis of a po
- Page 247 and 248:
in the response to daylength in ric
- Page 249 and 250:
functional genomics. However, it is
- Page 251 and 252:
Deletion mutants for functionalgeno
- Page 253 and 254:
Although these mutants are useful f
- Page 255 and 256:
Table 1. Frequencies of mutations o
- Page 257 and 258:
ALesion length (cm)252015105IR64D6-
- Page 259 and 260:
However, to detect genetic variabil
- Page 261 and 262:
cific PCR primers to screen a DNA p
- Page 263:
Reardon JT, Liljestrand-Golden CA,
- Page 266 and 267:
insertional mutagenesis an attracti
- Page 268 and 269:
plants transformed with pGA1633 and
- Page 270 and 271:
vealed that the efficiency of GUS s
- Page 272 and 273:
Table 3. GUS assay in flowers of tr
- Page 274 and 275:
Krizkova L, Hrouda M. 1998. Direct
- Page 276 and 277:
Functional genomicsThe field of fun
- Page 278 and 279:
Gene detectionGene detection strate
- Page 280 and 281:
ice. Transgenic plants were also ge
- Page 282 and 283:
ence of a single excision footprint
- Page 284 and 285:
homology to known proteins. The tra
- Page 286 and 287:
Transposon library generation10 sta
- Page 288 and 289:
Krysan PJ, Young JC, Sussman MR. 19
- Page 291 and 292:
Retrotransposons of riceas a tool f
- Page 293 and 294:
that Tos17 is the major insertional
- Page 295 and 296:
Forward and reverse genetic studies
- Page 297 and 298:
mutated genes by isolating the sequ
- Page 299 and 300:
5 kb—G1-T11 kb—5 kb—G1-T21 kb
- Page 301 and 302:
Mutants identified by this method a
- Page 303 and 304:
Ohtsubo H, Kumekawa N, Ohtsubo E. 1
- Page 305 and 306:
Bioinformatics and the rice genomeB
- Page 307 and 308:
total number of sequences submitted
- Page 309 and 310:
Table 2 continued.Database/ URL Inf
- Page 311 and 312:
Specialized databasesIn contrast to
- Page 313 and 314:
more complicated traits such as QTL
- Page 315 and 316:
ESTsGenetic markersMorphologicalMol
- Page 317:
NotesAuthors' address: Rice Genome
- Page 320 and 321:
308 Valent et al
- Page 322 and 323:
product, or they might encode enzym
- Page 324 and 325:
K1 (Kiyosawa 1984) obtained Pi-ta f
- Page 326 and 327:
A B CFig. 2. Rice leaves showing lo
- Page 328 and 329:
M. griseaAVR-PitaPlant cellProcessi
- Page 330 and 331:
The relationship between Pi-ta and
- Page 332 and 333:
ReferencesAtkins JG, Robert AL, Ada
- Page 334 and 335:
NotesAuthors’ addresses: B. Valen
- Page 336 and 337:
Role of the small GTPase Rac in dis
- Page 338 and 339:
Suppression of ROS production and c
- Page 340 and 341:
enging activities at the level of t
- Page 342 and 343:
lesions appeared over the entire le
- Page 344 and 345:
limited amino acid sequence informa
- Page 347 and 348:
Structure, function, and evolution
- Page 349 and 350:
trum, IRBB21 conferred a high level
- Page 351 and 352:
elements. Two of them were located
- Page 353 and 354:
IRBB2134532703506N20-247N18-116-1N1
- Page 355:
Salmeron JM, Oldroyd GED, Rommens C
- Page 358 and 359:
Understanding abiotic stress tolera
- Page 360 and 361:
Table 1. Selected cDNA libraries fr
- Page 362 and 363:
Table 2. Abundance profile of expre
- Page 364 and 365:
Table 3 continued.Gene AbundancePut
- Page 366 and 367:
duction of chips is done commercial
- Page 368 and 369:
Microarray determination of salt st
- Page 370 and 371:
ConclusionsThese analyses must be c
- Page 372 and 373:
Table 5 continued.Open readingframe
- Page 374 and 375:
Lin X, Kaul S, Rounsley S, Shea TP,
- Page 377 and 378:
Molecular dissectionof cell death i
- Page 379 and 380:
lysis appeared to be larger in diam
- Page 381 and 382:
Cell divisionCell elongationand exp
- Page 383 and 384:
extend to the S phase. Furthermore,
- Page 385 and 386:
and, at 6 h, the population of S ph
- Page 387 and 388:
Levine A, Tenhaken R, Dixon R, Lamb
- Page 389 and 390:
Molecular tools for achievingsynthe
- Page 391 and 392:
ANormal sexualpathwayMegasporangium
- Page 393 and 394:
Tripsacum, a close relative of maiz
- Page 395 and 396:
used T-DNA-tagging in Arabidopsis t
- Page 397 and 398:
aposporous apomicts such as H. aura
- Page 399 and 400:
Achieving synthetic apomixisThe min
- Page 401 and 402:
We isolated a rice homologue of the
- Page 403 and 404:
1996), and a putative hydroxyprolin
- Page 405 and 406:
The life cycle of angiosperms is pu
- Page 407 and 408:
Ablation of the sexual embryoIsolat
- Page 409 and 410:
Birchler JA. 1993. Dosage analysis
- Page 411 and 412:
Li JM, Yuan LP. 1999. Hybrid rice:
- Page 413:
Xu FX, Chye ML. 1999. Expression of
- Page 416 and 417:
404 Upadhyaya et al
- Page 418 and 419:
yield losses per annum of about US$
- Page 420 and 421:
Mutant replicase. With replicase ge
- Page 422 and 423:
Antiviral protein-mediated resistan
- Page 424 and 425:
ACRTSV (positive ssRNA)12,433 nt390
- Page 426 and 427:
Table 2. Summary of screening for r
- Page 428 and 429:
esistance obtained was delayed symp
- Page 430 and 431:
Finnegan J, McElroy D. 1994. Transg
- Page 432 and 433:
Stam M, de Bruin R, Kenter S, van d
- Page 435 and 436:
Transgenic approachesfor generating
- Page 437 and 438:
LEA proteins and their effect on st
- Page 439 and 440:
Table 1. Fresh and dry shoot weight
- Page 441 and 442:
Table 2. Growth performance of tran
- Page 443 and 444:
Table 3. The levels of green fluore
- Page 445 and 446:
(Zhang and Wu 1988). Plants were re
- Page 447 and 448:
expression levels in young root tip
- Page 449 and 450:
Kavi Kishor PB, Hong Z, Miao GH, Hu
- Page 451 and 452:
High-level expression ofC 4photosyn
- Page 453 and 454:
(Imaizumi et al 1997). However, the
- Page 455 and 456:
ously. Indeed, earlier attempts to
- Page 457 and 458:
(Hudspeth et al 1992). Not only inc
- Page 459:
Nomura M, Sentoku N, Nishimura A, L
- Page 462 and 463:
fects. Recent data suggest, however
- Page 464 and 465:
AH H H H H HEEEEEEBBBE H B UFig. 1.
- Page 466 and 467:
transcription through one transgene
- Page 468 and 469:
sertion of an adventitious DNA frag
- Page 470 and 471:
tin is folded into loops may result
- Page 472 and 473:
Whole-plasmid DNAClean fragmentBomb
- Page 474 and 475:
Concluding commentsExperiments invo
- Page 476 and 477:
Tinland B. 1996. The integration of
- Page 478 and 479:
immune systems. Thus, restriction e
- Page 480 and 481:
A: p35S-CryIIIA 35SBtt CryIIIA nosB
- Page 482 and 483:
appealing in regard to our efforts
- Page 484 and 485:
1998, Kumpatla and Hall 1999). The
- Page 486 and 487:
HindIIIHindIIIRB 35S hpt 35S 35S in
- Page 488 and 489:
ments (Francastel et al 1999), are
- Page 490 and 491:
ReferencesAmedeo P, Habu Y, Afsar K
- Page 492 and 493:
Matzke MA, Neuhuber F, Matzke AJ. 1
- Page 495 and 496:
Workshop reports483
- Page 497 and 498:
Functional genomics workshopThe obj
- Page 499 and 500:
Bioinformatics workshopThe Bioinfor