Macroalgae and phytoplankton as indicators of ... - Naturstyrelsen
Macroalgae and phytoplankton as indicators of ... - Naturstyrelsen
Macroalgae and phytoplankton as indicators of ... - Naturstyrelsen
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
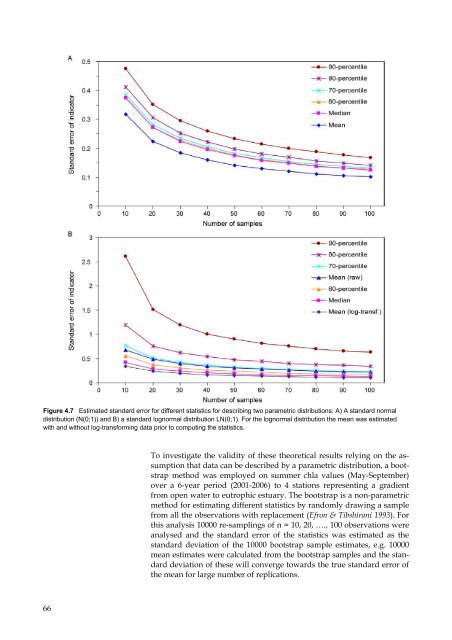
Figure 4.7 Estimated st<strong>and</strong>ard error for different statistics for describing two parametric distributions: A) A st<strong>and</strong>ard normal<br />
distribution (N(0;1)) <strong>and</strong> B) a st<strong>and</strong>ard lognormal distribution LN(0;1). For the lognormal distribution the mean w<strong>as</strong> estimated<br />
with <strong>and</strong> without log-transforming data prior to computing the statistics.<br />
To investigate the validity <strong>of</strong> these theoretical results relying on the <strong>as</strong>sumption<br />
that data can be described by a parametric distribution, a bootstrap<br />
method w<strong>as</strong> employed on summer chla values (May-September)<br />
over a 6-year period (2001-2006) to 4 stations representing a gradient<br />
from open water to eutrophic estuary. The bootstrap is a non-parametric<br />
method for estimating different statistics by r<strong>and</strong>omly drawing a sample<br />
from all the observations with replacement (Efron & Tibshirani 1993). For<br />
this analysis 10000 re-samplings <strong>of</strong> n = 10, 20, …., 100 observations were<br />
analysed <strong>and</strong> the st<strong>and</strong>ard error <strong>of</strong> the statistics w<strong>as</strong> estimated <strong>as</strong> the<br />
st<strong>and</strong>ard deviation <strong>of</strong> the 10000 bootstrap sample estimates, e.g. 10000<br />
mean estimates were calculated from the bootstrap samples <strong>and</strong> the st<strong>and</strong>ard<br />
deviation <strong>of</strong> these will converge towards the true st<strong>and</strong>ard error <strong>of</strong><br />
the mean for large number <strong>of</strong> replications.<br />
66