PDF Download - Laborwelt
PDF Download - Laborwelt
PDF Download - Laborwelt
Erfolgreiche ePaper selbst erstellen
Machen Sie aus Ihren PDF Publikationen ein blätterbares Flipbook mit unserer einzigartigen Google optimierten e-Paper Software.
B L I T Z L I C H T<br />
Sensorchips<br />
XNA on Gold TM Microarrays:<br />
Sensor Platform for Genomic,<br />
Proteomic and Glycomic Research<br />
DEV KAMBHAMPATI*, WEIDONG DU, MICHAEL SCHÄFERLING, MARGIT KRUSCHINA,MICHAEL CIEPLIK<br />
& FLAVIO ORTIGAO, THERMO HYBAID, ULM<br />
One of the outcomes of the colossal Human Genome Project was the widespread use of microarrays in<br />
biological research. Microarrays have revolutionized the way biotech and pharmaceutical research is<br />
conducted today. Large-scale gene expression and genotyping experiments can be conducted on a<br />
single chip, thus reducing the overall time and cost of the entire research study. Now that the euphoria<br />
over the sequencing of the Human Genome has finally settled, researchers are increasingly finding it<br />
challenging how to extract useful information from the raw code of massive data that was generated<br />
during the sequencing process. In addition, understanding the genome is only the first step in the long<br />
process of comprehending how the various aspects of cellular machinery function. The greatest<br />
challenge at the moment is to understand gene expression, polymorphisms and the functioning of<br />
proteins within the human body. Besides the genes and proteins, it is also important to realize the<br />
function and contribution of some of the most important, and often neglected, molecules which are of<br />
the carbohydrate family (saccharide, glycans, etc.). These molecules constitute the structural components<br />
of cells and are very critical in regulating inter-cellular molecular traffic across the cell membrane.<br />
Key Words: DNA Chips, Protein Chips, Glycochips, Microarrays<br />
To understand the genetic causation of<br />
diseases and how preventive steps on that<br />
basis can be taken, it is important to solve<br />
the mysteries of the above genomic, proteomic<br />
and glycomic challenges. Almost all of<br />
the microarrays that are currently available<br />
in the market are primarily designed to<br />
address only genomic problems and are<br />
highly inflexible when they are applied for<br />
addressing proteomic and glycomic issues.<br />
The biggest bottleneck for the currently<br />
available microarrays is their immobilization<br />
chemistries which are designed only for<br />
a limited range of applications. In addition,<br />
these microarrays suffer from a lot of nonspecific<br />
binding problems which greatly<br />
hampers the overall quality of the microarray<br />
data.<br />
The XNA on Gold TM microarray is a sensor<br />
platform that can currently address (Table 1)<br />
all aspects of genomic, proteomic and glycomic<br />
research. This sensor platform is universal<br />
in nature and can be used directly without<br />
changing the immobilization procedures<br />
for the various biomolecules (DNA, PNA,<br />
proteins, peptides, glycans, saccharides, lectins,<br />
etc.). Data with high signal-to-noise<br />
ratios can be achieved on this sensor platform,<br />
and since this is an universal microarray,<br />
it provides enormous flexibility for researchers<br />
to conducts a wide range of experiments<br />
that can have profound significance<br />
in the area of therapeutics, drug discovery<br />
and diagnostics.<br />
In the following sub-sections, the sensor strategy<br />
along with selected examples from the<br />
genomic (Single Nucleotide Polymor-<br />
phisms), proteomic (immunostatus), and<br />
glycomic (blood serology) applications will<br />
be provided. Finally, the various aspects of<br />
the proteomic research landscape will also<br />
be briefly addressed.<br />
Universal Sensor Platform<br />
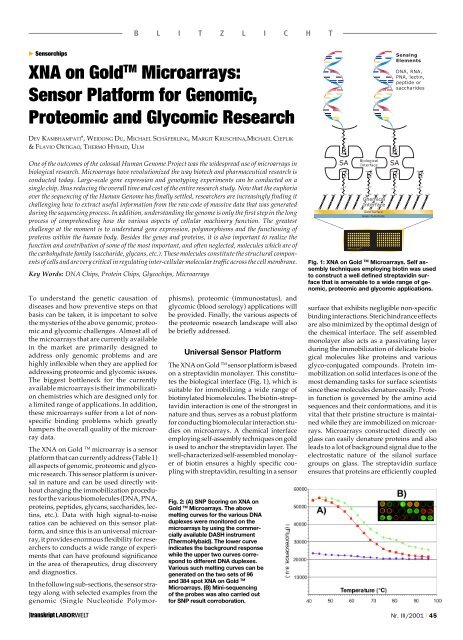
The XNA on Gold TM sensor platform is based<br />
on a streptavidin monolayer. This constitutes<br />
the biological interface (Fig. 1), which is<br />
suitable for immobilizing a wide range of<br />
biotinylated biomolecules. The biotin-streptavidin<br />
interaction is one of the strongest in<br />
nature and thus, serves as a robust platform<br />
for conducting biomolecular interaction studies<br />
on microarrays. A chemical interface<br />
employing self-assembly techniques on gold<br />
is used to anchor the streptavidin layer. The<br />
well-characterized self-assembled monolayer<br />
of biotin ensures a highly specific coupling<br />
with streptavidin, resulting in a sensor<br />
Fig. 2: (A) SNP Scoring on XNA on<br />
Gold TM Microarrays. The above<br />
melting curves for the various DNA<br />
duplexes were monitored on the<br />
microarrays by using the commercially<br />
available DASH instrument<br />
(ThermoHybaid). The lower curve<br />
indicates the background response<br />
while the upper two curves correspond<br />
to different DNA duplexes.<br />
Various such melting curves can be<br />
generated on the two sets of 96<br />
and 384 spot XNA on Gold TM<br />
Microarrays. (B) Mini-sequencing<br />
of the probes was also carried out<br />
for SNP result corroboration.<br />
NH<br />
O O O- NH<br />
Fig. 1: XNA on Gold TM Microarrays. Self assembly<br />
techniques employing biotin was used<br />
to construct a well defined streptavidin surface<br />
that is amenable to a wide range of genomic,<br />
proteomic and glycomic applications.<br />
surface that exhibits negligible non-specific<br />
binding interactions. Steric hindrance effects<br />
are also minimized by the optimal design of<br />
the chemical interface. The self assembled<br />
monolayer also acts as a passivating layer<br />
during the immobilization of delicate biological<br />
molecules like proteins and various<br />
glyco-conjugated compounds. Protein immobilization<br />
on solid interfaces is one of the<br />
most demanding tasks for surface scientists<br />
since these molecules denature easily. Protein<br />
function is governed by the amino acid<br />
sequences and their conformations, and it is<br />
vital that their pristine structure is maintained<br />
while they are immobilized on microarrays.<br />
Microarrays constructed directly on<br />
glass can easily denature proteins and also<br />
leads to a lot of background signal due to the<br />
electrostatic nature of the silanol surface<br />
groups on glass. The streptavidin surface<br />
ensures that proteins are efficiently coupled<br />
|transkript LABORWELT Nr. III/2001 | 45