Annual Scientific Report 2015
EMBL_EBI_ASR_2015_DigitalEdition
EMBL_EBI_ASR_2015_DigitalEdition
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Birney Group<br />
Nucleotide Data<br />
DNA sequence remains at the heart of molecular biology and bioinformatics.<br />
The Birney research group focuses on developing sequence algorithms and using<br />
intra-species variation to explore elements of basic biology.<br />
Dr Birney’s group has a long-standing interest in<br />
developing sequencing algorithms. Over the past few<br />
years they have focused on data compression, developing<br />
first theoretical then practical implementations of<br />
compression techniques. Dr Birney’s ‘blue skies’<br />
research includes collaborating with Dr Nick Goldman<br />
on a method to store digital data in DNA molecules. The<br />
Birney group continues to be involved in this area as<br />
new opportunities arise, for example the application of<br />
new sequencing technologies (in particular with Oxford<br />
Nanopore) and the integration of these methods with<br />
imaging techniques.<br />
The other of strand of the group’s research involves<br />
using natural DNA sequence variation to understand<br />
basic biology. Since 2010 there has been a tremendous<br />
increase in the use of genome-wide association studies<br />
to understand human diseases. This approach is very<br />
general and can be applied outside the arena of human<br />
disease. Association analysis can be applied to nearly<br />
any measureable phenotype in a cellular or organismal<br />
system where an accessible, outbred population is<br />
available. We are pursuing association analysis for<br />
a number of molecular (e.g. RNA expression levels,<br />
chromatin levels) and basic biology traits in a number<br />
of species where favourable populations are available,<br />
including human and Drosophila.<br />
We are also exploring molecular and physiological traits<br />
in human, for example 4D resolution of human hearts in<br />
healthy volunteers. We hope to expand this to a variety<br />
of basic biological phenotypes in other species, including<br />
establishing the first vertebrate near-isogenic wild panel<br />
in Japanese Rice Paddy fish (Medaka, Oryzias latipes).<br />
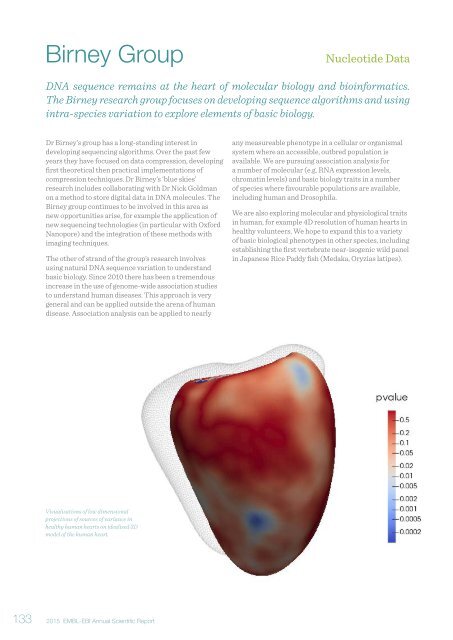
Visualisations of low dimensional<br />
projections of sources of variance in<br />
healthy human hearts on idealised 3D<br />
model of the human heart.<br />
133<br />
<strong>2015</strong> EMBL-EBI <strong>Annual</strong> <strong>Scientific</strong> <strong>Report</strong>