Annual Scientific Report 2015
EMBL_EBI_ASR_2015_DigitalEdition
EMBL_EBI_ASR_2015_DigitalEdition
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Ugis Sarkans<br />
Functional Genomics Development<br />
PhD in Computer Science, University of Latvia,<br />
1998. Postdoctoral research at the University of<br />
Wales, Aberystwyth, 2000.<br />
At EMBL-EBI since 2000.<br />
Future plans<br />
In 2016 we will release the BioStudies submission tool<br />
and, in the interests of providing users with a consistent<br />
experience, we will work with teams throughout<br />
EMBL-EBI to align further developments with those of<br />
other submission systems.<br />
To make BioStudies an appealing destination for<br />
supplementary data files in the life sciences, we will<br />
work on the presentation of content. In particular, we<br />
will further improve the BioStudies ‘project’ feature, by<br />
which information on studies can be rendered in such<br />
a way that they reflect the project/domain to which<br />
they belong.<br />
We will work with large projects that generate data on<br />
diverse ‘omics platforms to ensure that the data ends up<br />
in the correct structured repositories. We will use the<br />
BioStudies database to tie various modalities together,<br />
and provide study metadata and robust links to the<br />
BioSamples database. This work will support to the<br />
institute’s efforts to build a multi-omics atlas.<br />
The BioStudies and BioSamples databases represent<br />
two orthogonal components for dealing with<br />
multi-omics data at EMBL-EBI. BioStudies contains<br />
overall study descriptions and grouping assays, and<br />
BioSamples groups samples and provides annotation.<br />
We will continue integrating the two systems, such<br />
that information on samples in BioSamples is readily<br />
available for BioStudy submissions.<br />
We will further develop the Annotare tool for data<br />
submissions and, as for the BioStudies submission<br />
tool, will align it with other submission tools. We will<br />
also further simplify and automate the ArrayExpress<br />
data flow.<br />
Our continued participation in medical informatics<br />
projects will provide us with a better understanding of<br />
the data types and data-management patterns in a<br />
range of life-science communities, and this knowledge<br />
will be essential for our work in developing the<br />
BioStudies database.<br />
Selected publications<br />
Hendrickx DM, Aerts HJ, Caiment F, et al. (<strong>2015</strong>) diXa:<br />
a data infrastructure for chemical safety assessment.<br />
Bioinformatics (Oxford, England) 31:1505-1507<br />
Kirsanova C, Brazma A, Rustici G, Sarkans U (<strong>2015</strong>)<br />
Cellular phenotype database: a repository for systems<br />
microscopy data. Bioinformatics (Oxford, England)<br />
31:2736-2740<br />
Kolesnikov N, Hastings E, Keays M, et al. (<strong>2015</strong>)<br />
ArrayExpress update--simplifying data submissions.<br />
Nucleic Acids Res. 43:d1113-d1116<br />
McEntyre J, Sarkans U, Brazma A (<strong>2015</strong>) The BioStudies<br />
database. Mol. Syst.Biol. 11:847<br />
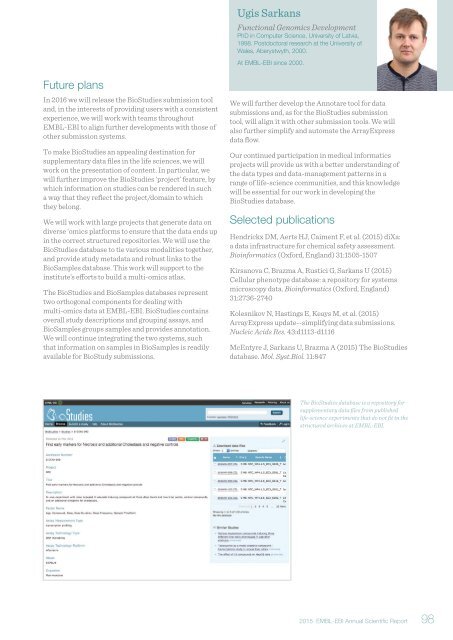
The BioStudies database is a repository for<br />
supplementary data files from published<br />
life-science experiments that do not fit in the<br />
structured archives at EMBL-EBI.<br />
<strong>2015</strong> EMBL-EBI <strong>Annual</strong> <strong>Scientific</strong> <strong>Report</strong> 98