Annual Scientific Report 2015
EMBL_EBI_ASR_2015_DigitalEdition
EMBL_EBI_ASR_2015_DigitalEdition
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
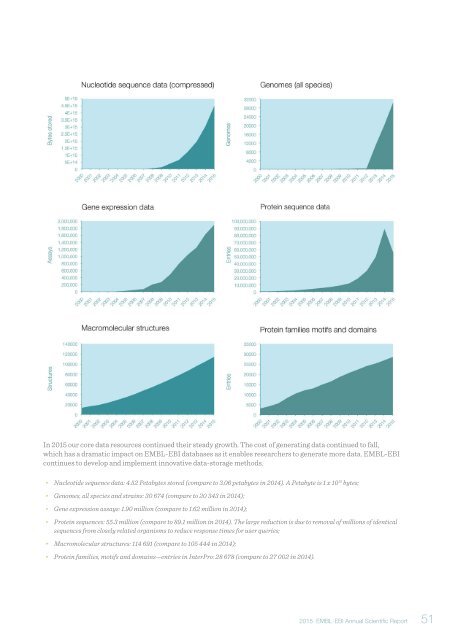
In <strong>2015</strong> our core data resources continued their steady growth. The cost of generating data continued to fall,<br />
which has a dramatic impact on EMBL-EBI databases as it enables researchers to generate more data. EMBL-EBI<br />
continues to develop and implement innovative data-storage methods.<br />
• Nucleotide sequence data: 4.52 Petabytes stored (compare to 3.06 petabytes in 2014). A Petabyte is 1 x 10 15 bytes;<br />
• Genomes, all species and strains: 30 674 (compare to 20 343 in 2014);<br />
• Gene expression assays: 1.90 million (compare to 1.62 million in 2014);<br />
• Protein sequences: 55.3 million (compare to 89.1 million in 2014). The large reduction is due to removal of millions of identical<br />
sequences from closely related organisms to reduce response times for user queries;<br />
• Macromolecular structures: 114 691 (compare to 105 444 in 2014);<br />
• Protein families, motifs and domains—entries in InterPro: 28 678 (compare to 27 002 in 2014).<br />
<strong>2015</strong> EMBL-EBI <strong>Annual</strong> <strong>Scientific</strong> <strong>Report</strong> 51