Redesigning Animal Agriculture
Redesigning Animal Agriculture
Redesigning Animal Agriculture
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
128 T. Doran and L. Lambeth<br />
ATP<br />
ADP + Pi<br />
5�-P<br />
3�-HO<br />
ATP<br />
ADP + Pi<br />
dsRNA<br />
(Bernstein et al., 2001), and is evolutionarily<br />
conserved in worms, flies, fungi, plants<br />
and mammals. A proposed model for Dicer<br />
involves the ATP-dependent translocation of<br />
the enzyme along its target prior to dsRNA<br />
cleavage, the efficiency of which has been<br />
shown to be directly proportional to the<br />
length of the target (Bernstein et al., 2001).<br />
Dicer has four distinct domains: an aminoterminal<br />
helicase domain, dual RNAse III<br />
motifs, a dsRNA binding domain, and a PAZ<br />
domain (named after its component proteins<br />
Piwi, Argo, and Zwile/Pinhead) (Tabara<br />
et al., 1999; Catalanotto et al., 2000).<br />
Following the cleavage of dsRNA into<br />
siRNAs by Dicer the second important stage<br />
OH-3�<br />
P-5� siRNA duplex<br />
RISC<br />
RISC activation and<br />
siRNA unwinding<br />
RISC<br />
Dicer<br />
RISC-mediated<br />
target recognition<br />
mRNA cleavage<br />
and degradation<br />
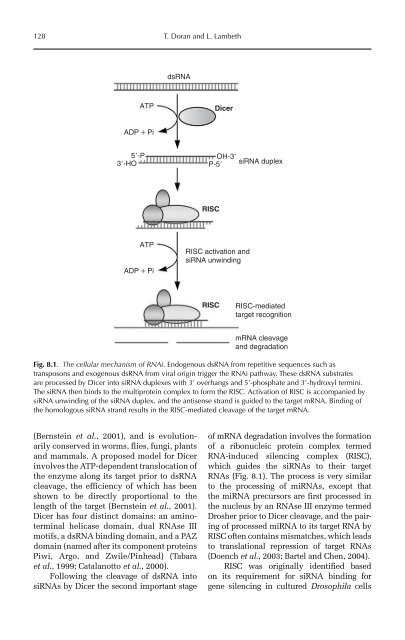
Fig. 8.1. The cellular mechanism of RNAi. Endogenous dsRNA from repetitive sequences such as<br />
transposons and exogenous dsRNA from viral origin trigger the RNAi pathway. These dsRNA substrates<br />
are processed by Dicer into siRNA duplexes with 3’ overhangs and 5’-phosphate and 3’-hydroxyl termini.<br />
The siRNA then binds to the multiprotein complex to form the RISC. Activation of RISC is accompanied by<br />
siRNA unwinding of the siRNA duplex, and the antisense strand is guided to the target mRNA. Binding of<br />
the homologous siRNA strand results in the RISC-mediated cleavage of the target mRNA.<br />
of mRNA degradation involves the form ation<br />
of a ribonucleic protein complex termed<br />
RNA-induced silencing complex (RISC),<br />
which guides the siRNAs to their target<br />
RNAs (Fig. 8.1). The process is very similar<br />
to the processing of miRNAs, except that<br />
the miRNA precursors are first processed in<br />
the nucleus by an RNAse III enzyme termed<br />
Drosher prior to Dicer cleavage, and the pairing<br />
of processed miRNA to its target RNA by<br />
RISC often contains mismatches, which leads<br />
to translational repression of target RNAs<br />
(Doench et al., 2003; Bartel and Chen, 2004).<br />
RISC was originally identified based<br />
on its requirement for siRNA binding for<br />
gene silencing in cultured Drosophila cells