et al.
et al.
et al.
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
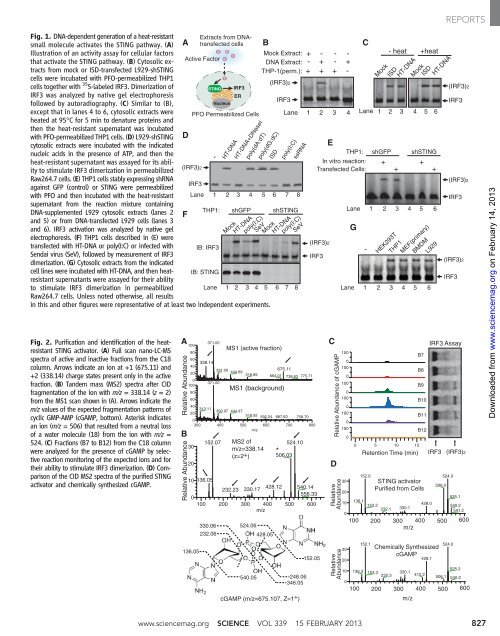
Fig. 1. DNA-dependent generation of a heat-resistant<br />
sm<strong>al</strong>l molecule activates the STING pathway. (A)<br />
Illustration of an activity assay for cellular factors<br />
that activate the STING pathway. (B) Cytosolic extracts<br />
from mock or ISD-transfected L929-shSTING<br />
cells were incubated with PFO-permeabilized THP1<br />
cells tog<strong>et</strong>her with 35 S-labeled IRF3. Dimerization of<br />
IRF3 was an<strong>al</strong>yzed by native gel electrophoresis<br />
followed by autoradiography. (C) Similar to (B),<br />
except that in lanes 4 to 6, cytosolic extracts were<br />
heated at 95°C for 5 min to denature proteins and<br />
then the heat-resistant supernatant was incubated<br />
with PFO-permeabilized THP1 cells. (D) L929-shSTING<br />
cytosolic extracts were incubated with the indicated<br />
nucleic acids in the presence of ATP, and then the<br />
heat-resistant supernatant was assayed for its ability<br />
to stimulate IRF3 dimerization in permeabilized<br />
Raw264.7 cells. (E) THP1 cells stably expressing shRNA<br />
against GFP (control) or STING were permeabilized<br />
with PFO and then incubated with the heat-resistant<br />
supernatant from the reaction mixture containing<br />
DNA-supplemented L929 cytosolic extracts (lanes 2<br />
and 5) or from DNA-transfected L929 cells (lanes 3<br />
and 6). IRF3 activation was an<strong>al</strong>yzed by native gel<br />
electrophoresis. (F) THP1 cells described in (E) were<br />
transfected with HT-DNA or poly(I:C) or infected with<br />
Sendai virus (SeV), followed by measurement of IRF3<br />
dimerization. (G) Cytosolic extracts from the indicated<br />
cell lines were incubated with HT-DNA, and then heatresistant<br />
supernatants were assayed for their ability<br />
to stimulate IRF3 dimerization in permeabilized<br />
Raw264.7 cells. Unless noted otherwise, <strong>al</strong>l results<br />
in this and other figures were representative of at least two independent experiments.<br />
Fig. 2. Purification and identification of the heatresistant<br />
STING activator. (A) Full scan nano-LC-MS<br />
spectra of active and inactive fractions from the C18<br />
column. Arrows indicate an ion at +1 (675.11) and<br />
+2 (338.14) charge states present only in the active<br />
fraction. (B) Tandem mass (MS2) spectra after CID<br />
fragmentation of the ion with m/z =338.14(z =2)<br />
from the MS1 scan shown in (A). Arrows indicate the<br />
m/z v<strong>al</strong>ues of the expected fragmentation patterns of<br />
cyclic GMP-AMP (cGAMP, bottom). Asterisk indicates<br />
an ion (m/z = 506) that resulted from a neutr<strong>al</strong> loss<br />
of a water molecule (18) from the ion with m/z =<br />
524. (C)Fractions(B7toB12)fromtheC18column<br />
were an<strong>al</strong>yzed for the presence of cGAMP by selective<br />
reaction monitoring of the expected ions and for<br />
their ability to stimulate IRF3 dimerization. (D)Comparison<br />
of the CID MS2 spectra of the purified STING<br />
activator and chemic<strong>al</strong>ly synthesized cGAMP.<br />
A<br />
Active Factor<br />
D<br />
(IRF3)2<br />
F<br />
Extracts from DNAtransfected<br />
cells<br />
STING IRF3<br />
ER<br />
Nucleus<br />
PFO Permeabilized Cells<br />
-<br />
HT-DNA+DNaseI<br />
poly(dA:dT)<br />
poly(dG:dC)<br />
ISD<br />
poly(I:C)<br />
ssRNA<br />
HT-DNA<br />
IRF3<br />
Lane 1 2 3 4 5 6 7 8<br />
IB: IRF3<br />
IB: STING<br />
B C<br />
+ - - -<br />
- + - +<br />
+ + + -<br />
Mock Extract:<br />
DNA Extract:<br />
THP-1(perm.):<br />
(IRF3)2<br />
IRF3<br />
THP1: shGFP shSTING<br />
Mock<br />
HT-DNA<br />
poly(I:C)<br />
SeV Mock<br />
HT-DNA<br />
poly(I:C)<br />
SeV<br />
Lane 1 2 3 4 5 6 7 8<br />
Lane 1 2 3 4<br />
E<br />
Mock<br />
ISD<br />
- HEK293T<br />
- heat +heat<br />
HT-DNA<br />
Lane 1 2 3 4 5 6<br />
THP1: shGFP shSTING<br />
In vitro reaction:<br />
Transfected Cells:<br />
+<br />
+<br />
+<br />
+<br />
(IRF3)2<br />
IRF3<br />
A 371.00<br />
C<br />
Relative Abundance<br />
B<br />
Relative Abundance<br />
100<br />
80<br />
60<br />
40<br />
20<br />
0<br />
100<br />
80<br />
60<br />
40<br />
20<br />
0<br />
30<br />
20<br />
338.14<br />
313.11<br />
371.00<br />
675.11<br />
391.00 444.89 518.89 664.01 775.71<br />
738.83<br />
300 400 500 600 700 800<br />
m/z<br />
10 136.05<br />
0<br />
136.05<br />
MS1 (active fraction)<br />
390.97 444.97<br />
MS1 (background)<br />
518.92 592.24 667.63 758.70<br />
152.07 MS2 of<br />
524.10<br />
m/z=338.14<br />
(z=2+)<br />
*<br />
506.03<br />
428.12<br />
232.23 330.17<br />
540.14<br />
556.33<br />
100 200 300 400 500 600<br />
m/z<br />
330.06 524.06<br />
232.06 428.05<br />
540.05<br />
248.06<br />
346.05<br />
cGAMP (m/z=675.107, Z=1 + )<br />
152.05<br />
Relative Abundance of cGAMP<br />
D<br />
Relative<br />
Abundance<br />
Relative<br />
Abundance<br />
Lane 1 2 3 4 5 6<br />
G<br />
BMDM<br />
THP1<br />
MEF(primary)<br />
Lane 1 2 3 4 5 6<br />
100<br />
0<br />
100<br />
0<br />
100<br />
0<br />
100<br />
0<br />
100<br />
0<br />
100<br />
30<br />
20<br />
10<br />
0<br />
30<br />
20<br />
10<br />
0<br />
0<br />
0 5 10 15<br />
R<strong>et</strong>ention Time (min)<br />
Mock<br />
ISD<br />
HT-DNA<br />
L929<br />
(IRF3)2<br />
IRF3<br />
(IRF3)2<br />
IRF3<br />
(IRF3)2<br />
IRF3<br />
IRF3 Assay<br />
IRF3 (IRF3)2<br />
100 200 300<br />
m/z<br />
400 500 600<br />
B7<br />
B8<br />
B9<br />
B10<br />
B11<br />
B12<br />
152.0 524.0<br />
STING activator<br />
Purified from Cells<br />
152.1 524.0<br />
Chemic<strong>al</strong>ly Synthesized<br />
cGAMP<br />
428.1<br />
506.0<br />
136.1<br />
153.2<br />
232.1 330.1<br />
428.0<br />
525.1<br />
540.2<br />
541.2<br />
100 200 300 400 500 600<br />
m/z<br />
525.2<br />
136.3 153.3<br />
330.1<br />
232.3 410.2<br />
506.1 540.2<br />
REPORTS<br />
www.sciencemag.org SCIENCE VOL 339 15 FEBRUARY 2013 827<br />
Downloaded from<br />
www.sciencemag.org on February 14, 2013