View - DSpace UniPR
View - DSpace UniPR
View - DSpace UniPR
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
PNA Molecular Beacons<br />
In particular, the 11mer probe could be efficiently set at the concentration of 30 nM (Figure 2-<br />
6 B), while the higher stability offered by the 13mer probe allowed to work at a concentration<br />
of 10 nM (Figure 2-6 D), reaching limits which could not be afforded in previous similar<br />
works 21, 25 .<br />
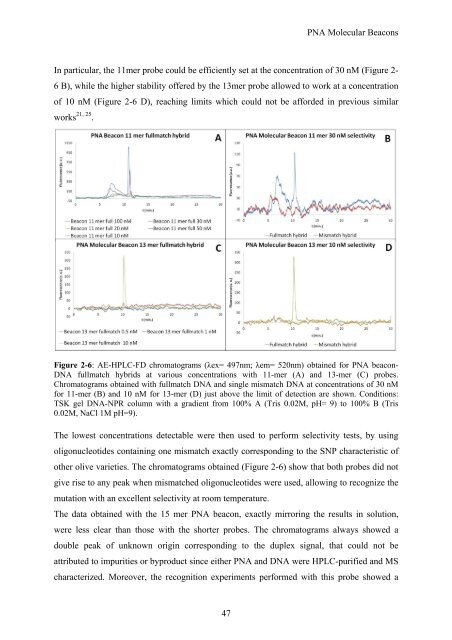
Figure 2-6: AE-HPLC-FD chromatograms (λex= 497nm; λem= 520nm) obtained for PNA beacon-<br />
DNA fullmatch hybrids at various concentrations with 11-mer (A) and 13-mer (C) probes.<br />
Chromatograms obtained with fullmatch DNA and single mismatch DNA at concentrations of 30 nM<br />
for 11-mer (B) and 10 nM for 13-mer (D) just above the limit of detection are shown. Conditions:<br />
TSK gel DNA-NPR column with a gradient from 100% A (Tris 0.02M, pH= 9) to 100% B (Tris<br />
0.02M, NaCl 1M pH=9).<br />
The lowest concentrations detectable were then used to perform selectivity tests, by using<br />
oligonucleotides containing one mismatch exactly corresponding to the SNP characteristic of<br />
other olive varieties. The chromatograms obtained (Figure 2-6) show that both probes did not<br />
give rise to any peak when mismatched oligonucleotides were used, allowing to recognize the<br />
mutation with an excellent selectivity at room temperature.<br />
The data obtained with the 15 mer PNA beacon, exactly mirroring the results in solution,<br />
were less clear than those with the shorter probes. The chromatograms always showed a<br />
double peak of unknown origin corresponding to the duplex signal, that could not be<br />
attributed to impurities or byproduct since either PNA and DNA were HPLC-purified and MS<br />
characterized. Moreover, the recognition experiments performed with this probe showed a<br />
47