View - DSpace UniPR
View - DSpace UniPR
View - DSpace UniPR
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
Arg-PNA Microarrays<br />
PNA-microarray platforms were built by using glass slides derivatised with N-<br />
hydroxysuccinimide active groups, which were allowed to react with the PNA terminal amino<br />
groups. The solutions for PNA spotting were prepared by dissolving the PNA probes (50 µM)<br />
in carbonate buffer (0.1 M, pH=9) containing SDS (0.001%). The deposition of the probes on<br />
the surface was done by using a pin system spotter.<br />
Since PNAs are known to be strongly absorbed on many types of surfaces, three so called<br />
“washing-spotting” cycles had to be introduced after each PNA deposition, in order to avoid<br />
false positive signals due to the contamination of the pin. The optimization of the washing<br />
step was done by spotting a PNA, followed by two or three spotting of washing solutions, and<br />
finally spotting the same buffer used to spot the probe, in order to check if any residue<br />
contamination was present in the pin. The essays allowed to evaluate the “washing power” of<br />
different mixtures, looking for the one able to completely remove the undeposited PNA from<br />
the pin. Washing solutions were prepared using organic solvents (such as acetonitrile and<br />
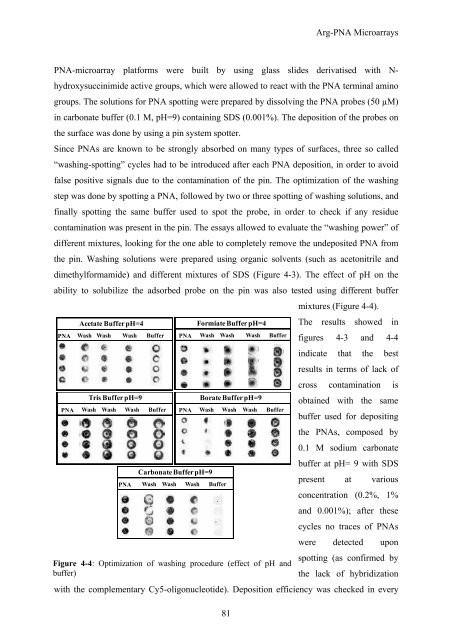
dimethylformamide) and different mixtures of SDS (Figure 4-3). The effect of pH on the<br />
ability to solubilize the adsorbed probe on the pin was also tested using different buffer<br />
mixtures (Figure 4-4).<br />
Acetate Buffer pH=4<br />
Formiate Buffer pH=4 The results showed in<br />
PNA Wash Wash Wash Buffer PNA Wash Wash Wash Buffer<br />
figures 4-3 and 4-4<br />
indicate that the best<br />
results in terms of lack of<br />
cross contamination is<br />
Tris Buffer pH=9<br />
BorateBuffer pH=9<br />
obtained with the same<br />
PNA Wash Wash Wash Buffer PNA Wash Wash Wash Buffer<br />
buffer used for depositing<br />
the PNAs, composed by<br />
0.1 M sodium carbonate<br />
buffer at pH= 9 with SDS<br />
Carbonate Buffer pH=9<br />
present at various<br />
PNA Wash Wash Wash Buffer<br />
concentration (0.2%, 1%<br />
and 0.001%); after these<br />
cycles no traces of PNAs<br />
were detected upon<br />
spotting (as confirmed by<br />
Figure 4-4: Optimization of washing procedure (effect of pH and<br />
buffer)<br />
the lack of hybridization<br />
with the complementary Cy5-oligonucleotide). Deposition efficiency was checked in every<br />
81