View - DSpace UniPR
View - DSpace UniPR
View - DSpace UniPR
Create successful ePaper yourself
Turn your PDF publications into a flip-book with our unique Google optimized e-Paper software.
PNA Molecular Beacons<br />
The sample thus obtained was<br />
hybridizedd with the 13 mer PNA<br />
beacon 100nM and analyzed<br />
by AE-HPLC using<br />
the same conditions optimized with synthetic oligonucleotides.<br />
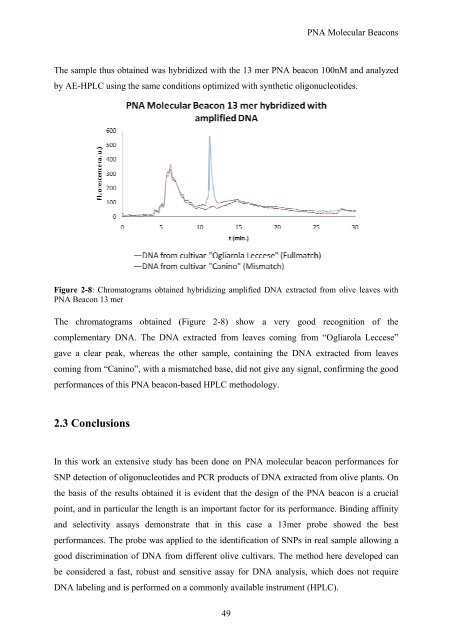
Figure 2-8: Chromatograms obtained hybridizing amplified DNA<br />
PNA Beacon 13 mer<br />
extracted from olive leaves with<br />
The chromatograms obtained (Figure<br />
2-8) show a very<br />
good recognition<br />
of the<br />
complementary DNA. The DNA extracted from leaves coming from “ Ogliarola Leccese”<br />
gave a clear peak, whereas the other sample, containing the<br />
DNA extracted from<br />
leaves<br />
coming from “Canino”, with a mismatched base, didd not give any signal, confirming the good<br />
performances of this PNA beacon-based HPLC methodology.<br />
2.3 Conclusions<br />
In this work an extensive study has been<br />
done on PNA molecular beacon performances for<br />
SNP detection of oligonucleotides and PCR products of DNA extracted from olive plants. On<br />
the basis of the results obtained it is evident that the design of the PNA beacon is a crucial<br />
point, and in particular the length is an important factor for its performance. Binding<br />
affinity<br />
and selectivity assays demonstrate that in this case a 13mer probe<br />
showed the best<br />
performances. The<br />
probe was applied to the identification of SNPs in real<br />
sample allowing a<br />
good discrimination of DNA from different olive cultivars. The method here developed can<br />
be considered a fast, robust and sensitive assay for DNA analysis, which does not<br />
require<br />
DNA labeling and is performed on a commonly available instrument (HPLC).<br />
49