Colletotrichum: complex species or species ... - CBS - KNAW
Colletotrichum: complex species or species ... - CBS - KNAW
Colletotrichum: complex species or species ... - CBS - KNAW
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Damm et al.<br />
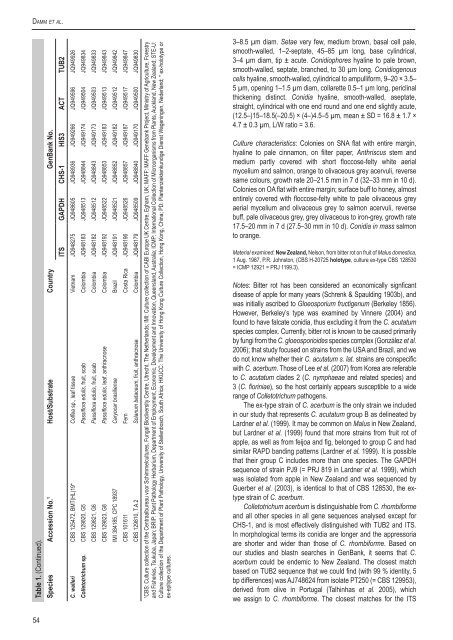
Table 1. (Continued).<br />
Species Accession No. 1 Host/Substrate Country GenBank No.<br />
ITS GAPDH CHS-1 HIS3 ACT TUB2<br />
C. walleri <strong>CBS</strong> 125472, BMT(HL)19* Coffea sp., leaf tissue Vietnam JQ948275 JQ948605 JQ948936 JQ949266 JQ949596 JQ949926<br />
<strong>Colletotrichum</strong> sp. <strong>CBS</strong> 129820, G5 Passifl<strong>or</strong>a edulis, fruit, scab Colombia JQ948183 JQ948513 JQ948844 JQ949174 JQ949504 JQ949834<br />
<strong>CBS</strong> 129821, G6 Passifl<strong>or</strong>a edulis, fruit, scab Colombia JQ948182 JQ948512 JQ948843 JQ949173 JQ949503 JQ949833<br />
<strong>CBS</strong> 129823, G8 Passifl<strong>or</strong>a edulis, leaf, anthracnose Colombia JQ948192 JQ948522 JQ948853 JQ949183 JQ949513 JQ949843<br />
IMI 384185, CPC 18937 Caryocar brasiliense Brazil JQ948191 JQ948521 JQ948852 JQ949182 JQ949512 JQ949842<br />
<strong>CBS</strong> 101611 Fern Costa Rica JQ948196 JQ948526 JQ948857 JQ949187 JQ949517 JQ949847<br />
<strong>CBS</strong> 129810, T.A.2 Solanum betaceum, fruit, anthracnose Colombia JQ948179 JQ948509 JQ948840 JQ949170 JQ949500 JQ949830<br />
1 <strong>CBS</strong>: Culture collection of the Centraalbureau vo<strong>or</strong> Schimmelcultures, Fungal Biodiversity Centre, Utrecht, The Netherlands; IMI: Culture collection of CABI Europe UK Centre, Egham, UK; MAFF: MAFF Genebank Project, Ministry of Agriculture, F<strong>or</strong>estry<br />
and Fisheries, Tsukuba, Japan; BRIP: Plant Pathology Herbarium, Department of Employment, Economic, Development and Innovation, Queensland, Australia; ICMP: International Collection of Micro<strong>or</strong>ganisms from Plants, Auckland, New Zealand; STE-U:<br />
Culture collection of the Department of Plant Pathology, University of Stellenbosch, South Africa; HKUCC: The University of Hong Kong Culture Collection, Hong Kong, China; PD: Plantenziektenkundige Dienst Wageningen, Nederland; * ex-holotype <strong>or</strong><br />
ex-epitype cultures.<br />
3–8.5 µm diam. Setae very few, medium brown, basal cell pale,<br />
smooth-walled, 1–2-septate, 45–85 µm long, base cylindrical,<br />
3–4 µm diam, tip ± acute. Conidioph<strong>or</strong>es hyaline to pale brown,<br />
smooth-walled, septate, branched, to 30 µm long. Conidiogenous<br />
cells hyaline, smooth-walled, cylindrical to ampullif<strong>or</strong>m, 9–20 × 3.5–<br />
5 µm, opening 1–1.5 µm diam, collarette 0.5–1 µm long, periclinal<br />
thickening distinct. Conidia hyaline, smooth-walled, aseptate,<br />
straight, cylindrical with one end round and one end slightly acute,<br />
(12.5–)15–18.5(–20.5) × (4–)4.5–5 µm, mean ± SD = 16.8 ± 1.7 ×<br />
4.7 ± 0.3 µm, L/W ratio = 3.6.<br />
Culture characteristics: Colonies on SNA flat with entire margin,<br />
hyaline to pale cinnamon, on filter paper, Anthriscus stem and<br />
medium partly covered with sh<strong>or</strong>t floccose-felty white aerial<br />
mycelium and salmon, <strong>or</strong>ange to olivaceous grey acervuli, reverse<br />
same colours, growth rate 20–21.5 mm in 7 d (32–33 mm in 10 d).<br />
Colonies on OA flat with entire margin; surface buff to honey, almost<br />
entirely covered with floccose-felty white to pale olivaceous grey<br />
aerial mycelium and olivaceous grey to salmon acervuli, reverse<br />
buff, pale olivaceous grey, grey olivaceous to iron-grey, growth rate<br />
17.5–20 mm in 7 d (27.5–30 mm in 10 d). Conidia in mass salmon<br />
to <strong>or</strong>ange.<br />
Material examined: New Zealand, Nelson, from bitter rot on fruit of Malus domestica,<br />
1 Aug. 1987, P.R. Johnston, (<strong>CBS</strong> H-20725 holotype, culture ex-type <strong>CBS</strong> 128530<br />
= ICMP 12921 = PRJ 1199.3).<br />
Notes: Bitter rot has been considered an economically signficant<br />
disease of apple f<strong>or</strong> many years (Schrenk & Spaulding 1903b), and<br />
was initially ascribed to Gloeosp<strong>or</strong>ium fructigenum (Berkeley 1856).<br />
However, Berkeley’s type was examined by Vinnere (2004) and<br />
found to have falcate conidia, thus excluding it from the C. acutatum<br />
<strong>species</strong> <strong>complex</strong>. Currently, bitter rot is known to be caused primarily<br />
by fungi from the C. gloeosp<strong>or</strong>ioides <strong>species</strong> <strong>complex</strong> (González et al.<br />
2006); that study focused on strains from the USA and Brazil, and we<br />
do not know whether their C. acutatum s. lat. strains are conspecific<br />
with C. acerbum. Those of Lee et al. (2007) from K<strong>or</strong>ea are referable<br />
to C. acutatum clades 2 (C. nymphaeae and related <strong>species</strong>) and<br />
3 (C. fi<strong>or</strong>iniae), so the host certainly appears susceptible to a wide<br />
range of <strong>Colletotrichum</strong> pathogens.<br />
The ex-type strain of C. acerbum is the only strain we included<br />
in our study that represents C. acutatum group B as delineated by<br />
Lardner et al. (1999). It may be common on Malus in New Zealand,<br />
but Lardner et al. (1999) found that m<strong>or</strong>e strains from fruit rot of<br />
apple, as well as from feijoa and fig, belonged to group C and had<br />
similar RAPD banding patterns (Lardner et al. 1999). It is possible<br />
that their group C includes m<strong>or</strong>e than one <strong>species</strong>. The GAPDH<br />
sequence of strain PJ9 (= PRJ 819 in Lardner et al. 1999), which<br />
was isolated from apple in New Zealand and was sequenced by<br />
Guerber et al. (2003), is identical to that of <strong>CBS</strong> 128530, the extype<br />
strain of C. acerbum.<br />
<strong>Colletotrichum</strong> acerbum is distinguishable from C. rhombif<strong>or</strong>me<br />
and all other <strong>species</strong> in all gene sequences analysed except f<strong>or</strong><br />
CHS-1, and is most effectively distinguished with TUB2 and ITS.<br />
In m<strong>or</strong>phological terms its conidia are longer and the appress<strong>or</strong>ia<br />
are sh<strong>or</strong>ter and wider than those of C. rhombif<strong>or</strong>me. Based on<br />
our studies and blastn searches in GenBank, it seems that C.<br />
acerbum could be endemic to New Zealand. The closest match<br />
based on TUB2 sequence that we could find (with 99 % identity, 5<br />
bp differences) was AJ748624 from isolate PT250 (= <strong>CBS</strong> 129953),<br />
derived from olive in P<strong>or</strong>tugal (Talhinhas et al. 2005), which<br />
we assign to C. rhombif<strong>or</strong>me. The closest matches f<strong>or</strong> the ITS<br />
54