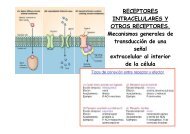
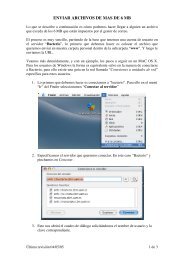
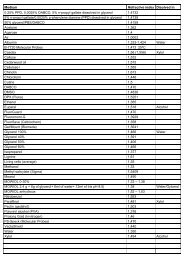
Jefe <strong>de</strong> Línea / Group Lea<strong>de</strong>r: Angel Ramírez Ortiz Unidad <strong>de</strong> bioinformática Bioinformatics unit Publicaciones Publications Resumen <strong>de</strong> investigación Research summary Murcia, M., Morreale, A. and Ortiz, A.R. (2006). Comparative binding energy analysis consi<strong>de</strong>ring multiple receptors: a step toward 3D- QSAR mo<strong>de</strong>ls for multiple targets. J Med Chem. 49, 6241-6253. Durante estos dos años hemos continuado nuestras investigaciones en el campo <strong>de</strong> la bioinformática estructural, en los siguientes aspectos: During these two years we have continued our research in the field of the Structural Bioinformatics, in the following aspects: Martín, V., Perales, C., Abia, D., Ortiz, A.R., Domingo E. and Briones, C. (2006). Microarray-based i<strong>de</strong>ntification of antigenic variants of foot-and-mouth disease virus: a bioinformatics quality assessment. BMC Genomics. 7, 117. Personal Científico / Scientific Staff: Paulino Gómez Postdoctorales / Postdoctoral: Antonio Morreale Rongsheng Han Ugo Bastolla Ester Marco Becarios Predoctorales / Predoctoral Fellows: David Abia Alberto Pascual Técnicos <strong>de</strong> Investigación / Technical Assistance: Rubén Muñoz Rubén Gil Redondo Carmen Arroyo Unidad <strong>de</strong> Bioinformática Bioinformatics unit 1. Estudio <strong>de</strong> los mecanismos <strong>de</strong> evolución estructural en proteínas: La estructura <strong>de</strong> las proteínas homólogas cambia durante el proceso evolutivo, lo que permite su adaptación a nuevas funciones. Cómo este proceso tiene lugar es gran parte <strong>de</strong>sconocido. Mediante análisis <strong>de</strong> estructuras homólogas hemos observado que éstas experimentan cambios concertados que en buena medida pue<strong>de</strong>n pre<strong>de</strong>cirse y explicarse como transiciones entre conformaciones en un espacio <strong>de</strong> baja dimensionalidad. En <strong>de</strong>finitiva, es la topología estructural la que <strong>de</strong>termina el patrón evolutivo <strong>de</strong> la familia. Similarmente, hemos continuado el estudio <strong>de</strong> cómo la topología <strong>de</strong> la proteína <strong>de</strong>termina la distribución <strong>de</strong> secuencias compatibles con un plegamiento dado y mejorando los métodos <strong>de</strong> la predicción estructural. 2. Diseño <strong>de</strong> fármacos basado en la estructura <strong>de</strong> proteínas: Hemos prestado gran atención al estudio <strong>de</strong> los mecanismos que <strong>de</strong>terminan la selectividad <strong>de</strong> la unión <strong>de</strong> una familia <strong>de</strong> ligandos a un conjunto <strong>de</strong> estructuras homologas, y hemos <strong>de</strong>sarrollado métodos para <strong>de</strong>terminar cuales son los residuos más importantes para explicar el grado <strong>de</strong> selectividad en la unión. 1. Study of the mechanisms of structural protein evolution. The structure of homologous proteins change during the evolutionary process, allowing an adaptation to new functions. How this process takes place is almost unknown. By means of structural analysis of homologous structures, we have observed that homologous proteins experience concerted changes that can be predicted and largely explained as transitions between energetically favourable conformations. In fact, we conclu<strong>de</strong> that the protein topology <strong>de</strong>termines the evolutionary pattern of a protein family. A second field of study is the analysis of how the topology of the protein <strong>de</strong>termines the distribution of compatible sequences with the protein fold in or<strong>de</strong>r to improve structure predition methods. Structure-based drug <strong>de</strong>sign: We have paid great attention to the study of the mechanisms that <strong>de</strong>termine the selectivity of the binding of a family of ligands to a set of homologous structures, and we have <strong>de</strong>veloped methods to <strong>de</strong>termine which are the most important residues able to explain the <strong>de</strong>gree of binding selectivity. Schamel, W.W., Risueno, R.M., Minguet, S., Ortiz, A.R. and Alarcón, B. (2006). A conformation- and avidity-based proofreading mechanism for the TCR-CD3 complex. Trends Immunol. 27, 176-182. Ortiz, A.R., Gomez-Puertas, P., Leo-Macias, A., López-Romero, P., López-Vinas, E., Morreale, A., Murcia, M. and Wang, K. (2006). Computational approaches to mo<strong>de</strong>l ligand selectivity in drug <strong>de</strong>sign. Curr Top Med Chem. 6, 41-55. Lozano, J. J., Soler, M., Bermuda, R., Abia, D., Fernán<strong>de</strong>z, P.L., Thomson, T.M. and Ortiz, A. R. (2005). Dual activation of pathways regulated by steroid receptors and pepti<strong>de</strong> growth factors in primary prostate cancer revealed by Factor Analysis of microarray data. BMC Genomics. 6, 109. Lupyan, D., Leo-Macias, A and Ortiz, A. R. (2005). A new progressiveiterative algorithm for multiple structure alignment. Bioinformatics. 21, 3255-3263. Leo-Macias, A., López-Romero, P., Lupyan, D., Zerbino, D. and Ortiz, A.R. (2005). Core <strong>de</strong>formations in protein families: a physical perspective. Biophys Chem. 115, 125-128. Leo-Macias, A., López-Romero, P., Lupyan, D., Zerbino, D. and Ortiz, A. R. (2004). An analysis of core <strong>de</strong>formations in protein superfamilies. Biophys J. 88, 1291-1299. Bastolla, U., Porto, M., Roman, H.E. and Vendruscolo, M. (2006). A protein evolution mo<strong>de</strong>l with in<strong>de</strong>pen<strong>de</strong>nt sites that reproduces sitespecific amino acid distributions from the Protein Data Bank. BMC Evol Biol. 6, 43. Bastolla, U. and Demetrius, L. (2005). Stability constraints and protein evolution: the role of chain length. Protein Eng. Des Sel. 18, 405-415. Bastolla, U., Porto, M., Roman, H. E.and Vendruscolo, M. (2005). Principal eigenvector of contact matrices and hydrophobicity profiles in proteins. Proteins. 58, 22-30. Talavera, D., Morreale, A., Meye, T., Hospital, A., Ferrer-Costa, C., Gelpi, J.L., <strong>de</strong> la Cruz, X., Soliva, R., Luque, F.J. and Orozco, M. (2006). A fast method for the <strong>de</strong>termination of fractional contributions to solvation in proteins. Protein Sci. 15, 2525-2533. CBM 2005/2006 148 Figura 2. Espacio evolutivo muestreado por diferentes familias <strong>de</strong> proteínas. Figure 2. Evolutionary subspace spanned by different protein families. Figura 1. Algoritmo implementado para la optimización <strong>de</strong> series congenéricas en contra familias <strong>de</strong> proteínas. Figure 1. Implemented algorithm <strong>de</strong>veloped to optimize the activity of congeneric series against specific family members of a protein family. 149
Cultura y divulgación científica Science and society program Semana <strong>de</strong> la Ciencia 2007. Science week 2007. 152 Oficina <strong>de</strong> Cultura Científica <strong>de</strong>l CBMSO CBMSO Science and Society Programme José Antonio López Guerrero