Mechanisms of Olfaction in Insects - ResearchSpace@Auckland ...
Mechanisms of Olfaction in Insects - ResearchSpace@Auckland ...
Mechanisms of Olfaction in Insects - ResearchSpace@Auckland ...
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
Identification <strong>of</strong> putative odorant receptors from Epiphyas postvittana 99<br />
consensus <strong>of</strong> the different doma<strong>in</strong>s was made manually. A topology map <strong>of</strong> EpOR34<br />
was then drawn <strong>in</strong> TOPO2 (http://www.sacs.ucsf.edu/TOPO-run/wtopo.pl) us<strong>in</strong>g the<br />
consensus sequence above.<br />
4.3 Results<br />
4.3.1 Microarray Analysis<br />
A microarray analysis <strong>of</strong> 1809 ESTs from a male antennal cDNA library <strong>of</strong> E.<br />
postvittana was carried out to identify male-specific genes. From the microarray<br />
screen<strong>in</strong>g, four ESTs had more than two times higher expression <strong>in</strong> male than female<br />
antennae (EST numbers 23893, 25116, 25174 and 25214). The sex-biased expression<br />
<strong>of</strong> two (ESTs 25174 and 25214) <strong>of</strong> the four ESTs were confirmed by qRT-PCR<br />
(Figure 4.1).<br />
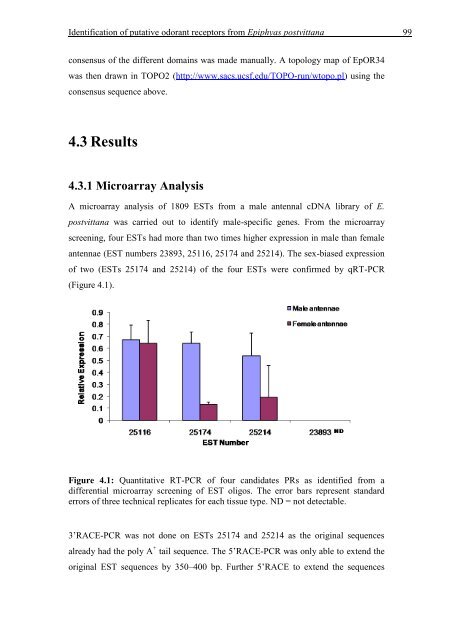
Figure 4.1: Quantitative RT-PCR <strong>of</strong> four candidates PRs as identified from a<br />
differential microarray screen<strong>in</strong>g <strong>of</strong> EST oligos. The error bars represent standard<br />
errors <strong>of</strong> three technical replicates for each tissue type. ND = not detectable.<br />
3‟RACE-PCR was not done on ESTs 25174 and 25214 as the orig<strong>in</strong>al sequences<br />
already had the poly A + tail sequence. The 5‟RACE-PCR was only able to extend the<br />
orig<strong>in</strong>al EST sequences by 350–400 bp. Further 5‟RACE to extend the sequences