- Page 1 and 2: http://researchspace.auckland.ac.nz
- Page 3 and 4: Abstract Abstract Olfaction or the
- Page 5 and 6: Acknowledgements Acknowledgements I
- Page 7 and 8: Contents v Contents Abstract……
- Page 9 and 10: Contents 4.2.5 Degenerate PCR .....
- Page 11 and 12: List of Figures List of Figures Fig
- Page 13 and 14: List of Tables List of Tables Table
- Page 15 and 16: List of Abbreviations gDNA Genomic
- Page 17 and 18: 1.1 General overview 1 General Intr
- Page 19 and 20: General Introduction 3 moth in loca
- Page 21 and 22: General Introduction 5 Figure 1.1:
- Page 23 and 24: General Introduction 7 1.5 Componen
- Page 25 and 26: General Introduction 9 OBPs have al
- Page 27 and 28: General Introduction 11 A model has
- Page 29 and 30: General Introduction 13 Figure 1.3:
- Page 31 and 32: General Introduction 15 Tanaka et a
- Page 33 and 34: General Introduction 17 Table 1.1:
- Page 35 and 36: General Introduction 19 Figure 1.4:
- Page 37 and 38: General Introduction 21 et al., 200
- Page 39 and 40: General Introduction 23 the attract
- Page 41 and 42: General Introduction 25 leaves and
- Page 43 and 44: General Introduction 27 Figure 1.7:
- Page 45: General Introduction 29 Expression
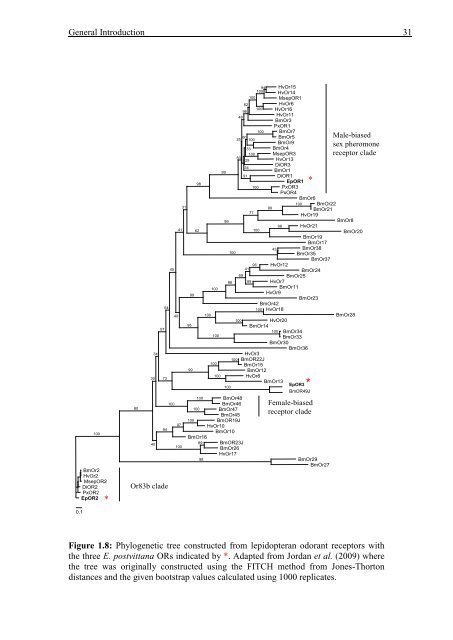
- Page 49 and 50: General Introduction 33 for lepidop
- Page 51 and 52: Functional characterisation of Epip
- Page 53 and 54: Functional characterisation of Epip
- Page 55 and 56: Functional characterisation of Epip
- Page 57 and 58: Functional characterisation of Epip
- Page 59 and 60: Functional characterisation of Epip
- Page 61 and 62: Functional characterisation of Epip
- Page 63 and 64: Functional characterisation of Epip
- Page 65 and 66: Functional characterisation of Epip
- Page 67 and 68: Functional characterisation of Epip
- Page 69 and 70: Functional characterisation of Epip
- Page 71 and 72: Functional characterisation of Epip
- Page 73 and 74: 3.1 Introduction 3 The roles of Epi
- Page 75 and 76: The roles of Epiphyas postvittana G
- Page 77 and 78: The roles of Epiphyas postvittana G
- Page 79 and 80: The roles of Epiphyas postvittana G
- Page 81 and 82: The roles of Epiphyas postvittana G
- Page 83 and 84: The roles of Epiphyas postvittana G
- Page 85 and 86: The roles of Epiphyas postvittana G
- Page 87 and 88: The roles of Epiphyas postvittana G
- Page 89 and 90: The roles of Epiphyas postvittana G
- Page 91 and 92: The roles of Epiphyas postvittana G
- Page 93 and 94: The roles of Epiphyas postvittana G
- Page 95 and 96: 4.1 Introduction 4 Identification o
- Page 97 and 98:
Identification of putative odorant
- Page 99 and 100:
Identification of putative odorant
- Page 101 and 102:
Identification of putative odorant
- Page 103 and 104:
Identification of putative odorant
- Page 105 and 106:
Identification of putative odorant
- Page 107 and 108:
Identification of putative odorant
- Page 109 and 110:
Microarray OR 454 Transcriptome OR
- Page 111 and 112:
Identification of putative odorant
- Page 113 and 114:
Identification of putative odorant
- Page 115 and 116:
Identification of putative odorant
- Page 117 and 118:
Identification of putative odorant
- Page 119 and 120:
Identification of putative odorant
- Page 121 and 122:
Identification of putative odorant
- Page 123 and 124:
Identification of putative odorant
- Page 125 and 126:
Identification of putative odorant
- Page 127 and 128:
Identification of putative odorant
- Page 129 and 130:
Identification of putative odorant
- Page 131 and 132:
Identification of putative odorant
- Page 133 and 134:
Identification of putative odorant
- Page 135 and 136:
Identification of putative odorant
- Page 137 and 138:
Identification of putative odorant
- Page 139 and 140:
Identification of putative odorant
- Page 141 and 142:
5.1 Introduction 5 Concluding Discu
- Page 143 and 144:
Concluding Discussion 127 VOBA sett
- Page 145 and 146:
Concluding Discussion 129 redundant
- Page 147 and 148:
Concluding Discussion 131 against t
- Page 149 and 150:
Concluding Discussion 133 sequencin
- Page 151 and 152:
Appendix B 135 Buffered TB (Sambroo
- Page 153 and 154:
Appendix C 137 C. Appendix C - Clus
- Page 155 and 156:
Appendix D 139 D. Appendix D - Solu
- Page 157 and 158:
Appendix E 141 I II
- Page 159 and 160:
Appendix E 143 V
- Page 161 and 162:
Appendix E 145 VII continued Figure
- Page 163 and 164:
References 147 Ansebo, L., Ignell,
- Page 165 and 166:
References 149 Butenandt, A., Beckm
- Page 167 and 168:
References 151 Fedurco, M., Romieu,
- Page 169 and 170:
References 153 Große-Wilde, E., St
- Page 171 and 172:
References 155 Hrdy, I., F. Kuldova
- Page 173 and 174:
References 157 Kaissling, K. E. (19
- Page 175 and 176:
References 159 Krieger, J., Große-
- Page 177 and 178:
References 161 Lopez-Gomez, R. and
- Page 179 and 180:
References 163 Miura, N., Nakagawa,
- Page 181 and 182:
References 165 Pell, J. K., Macaula
- Page 183 and 184:
References 167 Røstelien, T., Stra
- Page 185 and 186:
References 169 Stowers, L. and Loga
- Page 187 and 188:
References 171 Turner, C. T., Davy,
- Page 189 and 190:
References 173 Vogt, R. G., Riddifo
- Page 191 and 192:
References 175 Widmayer, P., Heifet