in vitro PHARMACOLOGY 2011 CATALOG - Cerep
in vitro PHARMACOLOGY 2011 CATALOG - Cerep
in vitro PHARMACOLOGY 2011 CATALOG - Cerep
You also want an ePaper? Increase the reach of your titles
YUMPU automatically turns print PDFs into web optimized ePapers that Google loves.
173<br />
proteases [metalloproteases] ❚<br />
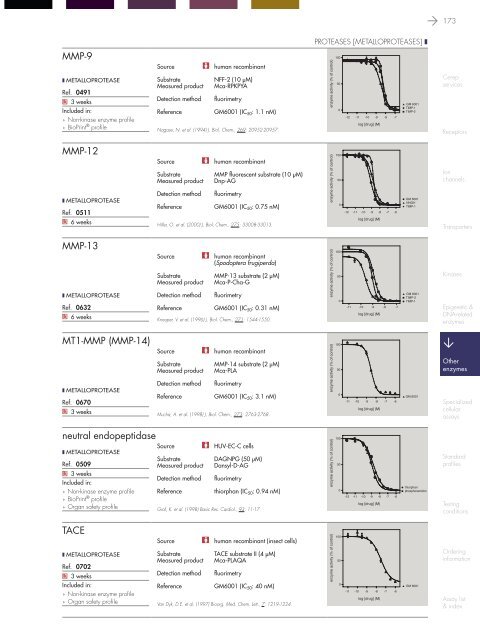
MMP-9<br />
❚ metalloprotease<br />
Ref. 0491<br />
Q 3 weeks<br />
Included <strong>in</strong>:<br />
Non-k<strong>in</strong>ase enzyme profile<br />
BioPr<strong>in</strong>t ® profile<br />
Source<br />
human recomb<strong>in</strong>ant<br />
Substrate<br />
NFF-2 (10 µM)<br />
Measured product Mca-RPKPYA<br />
Detection method fluorimetry<br />
Reference<br />
GM6001 (IC 50 : 1.1 nM)<br />
Nagase, N. et al. (1994) J. Biol. Chem., 269: 20952-20957.<br />
enzyme activity (% of control)<br />
100<br />
50<br />
0<br />
-12 -11 -10 -9 -8 -7<br />
log [drug] (M)<br />
GM 6001<br />
TIMP-1<br />
TIMP-2<br />
<strong>Cerep</strong><br />
services<br />
Receptors<br />
MMP-12<br />
❚ metalloprotease<br />
Ref. 0511<br />
Q 6 weeks<br />
Source<br />
Substrate<br />
Measured product<br />
Detection method<br />
Reference<br />
human recomb<strong>in</strong>ant<br />
MMP fluorescent substrate (10 µM)<br />
Dnp-AG<br />
fluorimetry<br />
GM6001 (IC 50 : 0.75 nM)<br />
Hiller, O. et al. (2000) J. Biol. Chem., 275: 33008-33013.<br />
enzyme activity (% of control)<br />
100<br />
-10 -9 -8 -7 -6 -5 -4<br />
50<br />
-11 -10 -9 -8 -7 -6<br />
GM 6001<br />
NNGH<br />
0<br />
TIMP-1<br />
-12 -11 -10 -9 -8 -7 -6<br />
log [drug] (M)<br />
Ion<br />
channels<br />
Transporters<br />
MMP-13<br />
❚ metalloprotease<br />
Ref. 0632<br />
Q 6 weeks<br />
Source<br />
Substrate<br />
Measured product<br />
Detection method<br />
Reference<br />
human recomb<strong>in</strong>ant<br />
(Spodoptera frugiperda)<br />
MMP-13 substrate (2 µM)<br />
Mca-P-Cha-G<br />
fluorimetry<br />
GM6001 (IC 50 : 0.31 nM)<br />
Knauper, V. et al. (1996) J. Biol. Chem., 271: 1544-1550.<br />
<br />
<br />
-10 -9 -8 -7 -6 -5 -4<br />
<br />
-11 -10 -9 -8 -7 -6<br />
-12 -11 -10 -6<br />
-9 -8 -7<br />
<br />
<br />
<br />
<br />
<br />
<br />
K<strong>in</strong>ases<br />
Epigenetic &<br />
DNA-related<br />
enzymes<br />
MT1-MMP (MMP-14)<br />
❚ metalloprotease<br />
Ref. 0670<br />
Q 3 weeks<br />
Source<br />
human recomb<strong>in</strong>ant<br />
Substrate<br />
MMP-14 substrate (2 µM)<br />
Measured product Mca-PLA<br />
Detection method fluorimetry<br />
Reference<br />
GM6001 (IC 50 : 3.1 nM)<br />
Mucha, A. et al. (1998) J. Biol. Chem., 273: 2763-2768.<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
Other<br />
enzymes<br />
Specialized<br />
cellular<br />
assays<br />
neutral endopeptidase<br />
❚ metalloprotease<br />
Ref. 0509<br />
Q 3 weeks<br />
Included <strong>in</strong>:<br />
Non-k<strong>in</strong>ase enzyme profile<br />
BioPr<strong>in</strong>t ® profile<br />
Organ safety profile<br />
Source<br />
Substrate<br />
Measured product<br />
Detection method<br />
Reference<br />
HUV-EC-C cells<br />
DAGNPG (50 µM)<br />
Dansyl-D-AG<br />
fluorimetry<br />
thiorphan (IC 50 : 0.94 nM)<br />
Graf, K. et al. (1998) Basic Res. Cardiol., 93: 11-17.<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
<br />
Standard<br />
profiles<br />
Test<strong>in</strong>g<br />
conditions<br />
TACE<br />
❚ metalloprotease<br />
Ref. 0702<br />
Q 3 weeks<br />
Included <strong>in</strong>:<br />
Non-k<strong>in</strong>ase enzyme profile<br />
Organ safety profile<br />
Source<br />
human recomb<strong>in</strong>ant (<strong>in</strong>sect cells)<br />
Substrate<br />
TACE substrate II (4 µM)<br />
Measured product Mca-PLAQA<br />
Detection method fluorimetry<br />
Reference<br />
GM6001 (IC 50 : 40 nM)<br />
Van Dyk, D.E. et al. (1997) Bioorg. Med. Chem. Lett., 7: 1219-1224.<br />
enzyme activity (% of control)<br />
100<br />
50<br />
0<br />
-11 -10 -9 -8 -7 -6<br />
log [drug] (M)<br />
GM 6001<br />
Order<strong>in</strong>g<br />
<strong>in</strong>formation<br />
Assay list<br />
& <strong>in</strong>dex