DISPATCHESTable. Alignment of residues involved receptor binding of hemagglutinin of influenza A viruses*Amino acid position (H3 numbering)Origin/subtype Isolate name 131 135 137 138 152 186 190 193 200 222 224 225 226 227 228Human/H3N2 A/Panama/2007/1999 A T S A N S D S G W R G V S SHuman/H3N2 A/Texas/50/2012 T T S A N G D F G R R N I P SHuman/H3N2 A/Brisbane/10/2007 T T S A N V N F G R R N I P SHuman/H1N1 A/California/04/2009 D V A A I S D S T K R D Q E GHuman/H1N1 A/Texas/36/1991 V V T S L S D A A K R G Q E GHuman/H1N1 A/Brisbane/59/2007 T V A S L P D A A K R D Q E GAvian/H1N1 A/duck/Alberta/1976 T V A A L P E S A E R G Q A GAvian/H7N1 A/rhea/North R A S A K G E K T F S G R I DCarolina/39482/1993Avian/H6N1 A/mallard/Sweden/81/ D V K A L P E T R A N G Q R G2002Avian (human A/Vietnam/1203/2004 A V S A V N E K T K N G Q S Gisolate)/H5N1Avian (human A/Anhui/1/2013 R A S A K V E K K Q N G L S Gisolate)/H7N9Avian/H10N7 A/shorebird/Delaware N T R A K S E D L Q N G Q S GBay/10/2004Avian/H10N7 A/mallard/Interior N T K A K S E D L Q N G Q S GAlaska/10BM01929/2010Avian (sealA/harbor N T K A K S E D L Q N G Q S Gisolate)/H10N7 seal/Germany/1/2014Avian (human A/Jiangxi- N T R A K S E D L Q N G Q S Gisolate)/H10N8 Donghu/346–1/2013*Residues found in human H1 or H3 and in H10 hemagglutinin but not in other avian hemagglutinin sequences are shown in bold.MDCK epithelial cells with H10-JD346 virus (6:2 re-assortantwith the backbone of laboratory strain A/PuertoRico/8/1934 [PR8], which was generated as described)(9,10); H10-mallard (wild-type); human isolate H3-P99(wild-type); and H5-Viet 6:2 (low pathogenicity reassortantwith the backbone of PR8) (9,10) at a multiplicityof infection of 1. Cells were harvested 24-h postinfectionand incubated with antibody against matrix protein 2(E10), which was detected by using an antibody againstIgG (Alexa 647 antibody; Invitrogen, Carlsbad, CA, USA)as a control of infection and with the sialyl-glycans (detectedwith streptavidin–fluorescein isothiocyanate; JacksonLaboratories, Bar Harbor, ME, USA). We determinedthe percentage of infected cells in each sample and gatedthe infected population to determine the SA binding profile(Figure 1, panel B).H3-P99 showed high levels of binding to SAα2,6 andH5-Viet bound more efficiently to SAα2,3 than to SAα2,6,which is similar to observations with recombinant HAsin the solid-phase binding assay. H10-mallard and H10-JD346 showed similar binding profiles with preferentialbinding for SAα2,3 and binding to SAα2,6 slightly higherthan that for the negative control.Analysis of receptor binding of H10-JD346 and of H10-mallard with 2 independent assays indicated that the H10subtype influenza virus interacts slightly with human-likereceptors and maintains preferential binding to avian-like receptors.Consequently, these data suggest that H10 subtypeinfluenza virus might have the ability to interact with the upperhuman respiratory tract, which is rich in SAα2,6 (3).To test this hypothesis, we precomplexed H3-P99and H10-JD346 with primary antibody (mouse anti-Histag) and secondary fluorescent antibody, then incubatedthe complex with 2 human tracheal samples (12). As expected,H3-P99 HA bound to the surface of respiratoryepithelia (Figure 2). Recombinant H10-JD346 HA alsointeracted with respiratory epithelia (Figure 2), whichsuggested that the virus might be able to attach and replicatein the human upper respiratory tract. However, the6:2 reassortant virus H10-JD346 virus showed markedlydecreased replication compared with that of an H3N2subtype virus (PR8 6:2 reassortant) in a human lung epithelialcell line (Figure 3, http://wwwnc.cdc.gov/EID/article/21/7/14-1755-F3.htm).ConclusionsHA of novel influenza A(H10N8) virus interacts withSAα2,3 and slightly with SAα2,6, at levels similar to thatfor an avian H10 subtype HA, and binds to cells in the humanupper respiratory tract. Our findings are consistentwith those of Vachiery et al. (8) but show some differencesfrom those of Yang et al. (13) and Wang et al. (14), whodid not detect interaction with SAα2,6 or human trachea.Variations in the experimental settings and protocols (e.g.,concentration of HA or glycans used) might account forthese dissimilarities.Only 3 cases of human infections with influenzaA(H10N8) viruses have been reported. However, H10N7subtype viruses have caused conjunctivitis or mild respiratorysymptoms in humans. An epidemic among seals1198 Emerging Infectious Diseases • www.cdc.gov/eid • Vol. 21, No. 7, July 2015
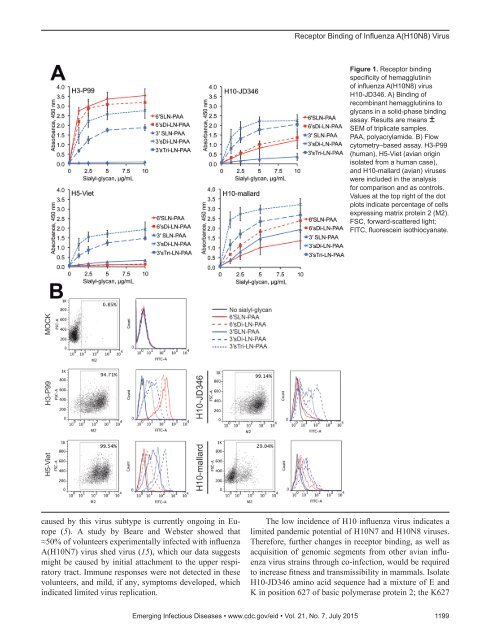
Receptor Binding of Influenza A(H10N8) VirusFigure 1. Receptor bindingspecificity of hemagglutininof influenza A(H10N8) virusH10-JD346. A) Binding ofrecombinant hemagglutinins toglycans in a solid-phase bindingassay. Results are means ±SEM of triplicate samples.PAA, polyacrylamide. B) Flowcytometry–based assay. H3-P99(human), H5-Viet (avian originisolated from a human case),and H10-mallard (avian) viruseswere included in the analysisfor comparison and as controls.Values at the top right of the dotplots indicate percentage of cellsexpressing matrix protein 2 (M2).FSC, forward-scattered light;FITC, fluorescein isothiocyanate.caused by this virus subtype is currently ongoing in Europe(5). A study by Beare and Webster showed that≈50% of volunteers experimentally infected with influenzaA(H10N7) virus shed virus (15), which our data suggestsmight be caused by initial attachment to the upper respiratorytract. Immune responses were not detected in thesevolunteers, and mild, if any, symptoms developed, whichindicated limited virus replication.The low incidence of H10 influenza virus indicates alimited pandemic potential of H10N7 and H10N8 viruses.Therefore, further changes in receptor binding, as well asacquisition of genomic segments from other avian influenzavirus strains through co-infection, would be requiredto increase fitness and transmissibility in mammals. IsolateH10-JD346 amino acid sequence had a mixture of E andK in position 627 of basic polymerase protein 2; the K627Emerging Infectious Diseases • www.cdc.gov/eid • Vol. 21, No. 7, July 2015 1199
- Page 3 and 4:
July 2015SynopsisOn the CoverMarian
- Page 5 and 6:
1240 Gastroenteritis OutbreaksCause
- Page 7 and 8:
SYNOPSISDisseminated Infections wit
- Page 9 and 10:
Disseminated Infections with Talaro
- Page 11 and 12:
Disseminated Infections with Talaro
- Page 13 and 14:
Macacine Herpesvirus 1 inLong-Taile
- Page 15 and 16:
Macacine Herpesvirus 1 in Macaques,
- Page 17 and 18:
Macacine Herpesvirus 1 in Macaques,
- Page 19:
Macacine Herpesvirus 1 in Macaques,
- Page 23:
Malaria among Young Infants, Africa
- Page 26 and 27:
RESEARCHFigure 3. Dynamics of 19-kD
- Page 28 and 29:
Transdermal Diagnosis of MalariaUsi
- Page 30 and 31:
RESEARCHFigure 2. A) Acoustic trace
- Page 32 and 33:
RESEARCHof malaria-infected mosquit
- Page 34 and 35:
Lack of Transmission amongClose Con
- Page 36 and 37:
RESEARCH(IFA) and microneutralizati
- Page 38 and 39:
RESEARCHoropharyngeal, and serum sa
- Page 40 and 41:
RESEARCH6. Assiri A, McGeer A, Perl
- Page 42 and 43:
RESEARCHadvanced genomic sequencing
- Page 44 and 45:
RESEARCHTable 2. Next-generation se
- Page 46 and 47:
RESEARCHTable 3. Mutation analysis
- Page 48 and 49:
RESEARCHReferences1. Baize S, Panne
- Page 50 and 51:
Parechovirus Genotype 3 Outbreakamo
- Page 52 and 53:
RESEARCHFigure 1. Venn diagramshowi
- Page 54 and 55: RESEARCHTable 2. HPeV testing of sp
- Page 56 and 57: RESEARCHFigure 5. Distribution of h
- Page 58 and 59: RESEARCHReferences1. Selvarangan R,
- Page 60 and 61: RESEARCHthe left lobe was sampled b
- Page 62 and 63: RESEARCHTable 2. Middle East respir
- Page 64 and 65: RESEARCHseroprevalence in domestic
- Page 66 and 67: RESEARCHmeasure their current surve
- Page 68 and 69: RESEARCHTable 2. States with labora
- Page 70 and 71: RESEARCHFigure 2. Comparison of sur
- Page 72 and 73: RESEARCH9. Centers for Disease Cont
- Page 74 and 75: RESEARCHthe analyses. Cases in pers
- Page 76 and 77: RESEARCHTable 3. Sampling results (
- Page 78 and 79: RESEARCHpresence of Legionella spp.
- Page 80 and 81: Seroprevalence for Hepatitis Eand O
- Page 82 and 83: RESEARCHTable 1. Description of stu
- Page 84 and 85: RESEARCHTable 3. Crude and adjusted
- Page 86 and 87: RESEARCHrates by gender or HIV stat
- Page 88 and 89: RESEARCH25. Taha TE, Kumwenda N, Ka
- Page 90 and 91: POLICY REVIEWDutch Consensus Guidel
- Page 92 and 93: POLICY REVIEWTable 3. Comparison of
- Page 94 and 95: POLICY REVIEW6. Botelho-Nevers E, F
- Page 96 and 97: DISPATCHESFigure 1. Phylogenetic tr
- Page 98 and 99: DISPATCHESSevere Pediatric Adenovir
- Page 100 and 101: DISPATCHESTable 1. Demographics and
- Page 102 and 103: DISPATCHES13. Kim YJ, Hong JY, Lee
- Page 106 and 107: DISPATCHESFigure 2. Interaction of
- Page 108 and 109: DISPATCHESSchmallenberg Virus Recur
- Page 110 and 111: DISPATCHESFigure 2. Detection of Sc
- Page 112 and 113: DISPATCHESFigure 1. Histopathologic
- Page 114: DISPATCHESFigure 2. Detection of fo
- Page 117 and 118: Influenza Virus Strains in the Amer
- Page 119 and 120: Novel Arenavirus Isolates from Nama
- Page 121 and 122: Novel Arenaviruses, Southern Africa
- Page 123 and 124: Readability of Ebola Informationon
- Page 125 and 126: Readability of Ebola Information on
- Page 127 and 128: Patients under investigation for ME
- Page 129 and 130: Patients under investigation for ME
- Page 131 and 132: Wildlife Reservoir for Hepatitis E
- Page 133 and 134: Asymptomatic Malaria and Other Infe
- Page 135 and 136: Asymptomatic Malaria in Children fr
- Page 137 and 138: Bufavirus in Wild Shrews and Nonhum
- Page 139 and 140: Bufavirus in Wild Shrews and Nonhum
- Page 141 and 142: Range Expansion for Rat Lungworm in
- Page 143 and 144: Slow Clearance of Plasmodium falcip
- Page 145 and 146: Slow Clearance of Plasmodium falcip
- Page 147 and 148: Gastroenteritis Caused by Norovirus
- Page 149 and 150: Ebola Virus Stability on Surfaces a
- Page 151 and 152: Ebola Virus Stability on Surfaces a
- Page 153 and 154: Outbreak of Ciprofloxacin-Resistant
- Page 155 and 156:
Outbreak of S. sonnei, South KoreaT
- Page 157 and 158:
Rapidly Expanding Range of Highly P
- Page 159 and 160:
Cluster of Ebola Virus Disease, Bon
- Page 161 and 162:
Cluster of Ebola Virus Disease, Lib
- Page 163 and 164:
ANOTHER DIMENSIONThe Past Is Never
- Page 165 and 166:
Measles Epidemic, Boston, Massachus
- Page 167 and 168:
LETTERSInfluenza A(H5N6)Virus Reass
- Page 169 and 170:
LETTERSsystem (8 kb-span paired-end
- Page 171 and 172:
LETTERS3. Van Hong N, Amambua-Ngwa
- Page 173 and 174:
LETTERSTable. Prevalence of Bartone
- Page 175 and 176:
LETTERSavian influenza A(H5N1) viru
- Page 177 and 178:
LETTERSprovinces and a total of 200
- Page 179 and 180:
LETTERS7. Manian FA. Bloodstream in
- Page 181 and 182:
LETTERSforward projections. N Engl
- Page 183 and 184:
LETTERS3. Guindon S, Gascuel OA. Si
- Page 185 and 186:
BOOKS AND MEDIAin the port cities o
- Page 187 and 188:
ABOUT THE COVERNorth was not intere
- Page 189 and 190:
Earning CME CreditTo obtain credit,
- Page 191:
Emerging Infectious Diseases is a p