34Molecular s<strong>im</strong>ulation studies of the interaction of weakly coordinating ions withbiological interfacesD. Gabel, K. Karki, D. Roccatano, Department ofChemistry, University of Bremen, School of Engineeringand Science, Jacobs University BremenAbstract• Weakly coordinating ions show properties whichcannot be explained on purely electrostaticgrounds. Such ions are large, and prominent examplesare ionic boron clusters (IBC).• We want to study the interaction of IBC withmembranes and with proteins using moleculardynamics s<strong>im</strong>ulations, in order to have a bettermolecular understanding of the exper<strong>im</strong>ental results.• A force field for molecular dynamics s<strong>im</strong>ulationof IBC must be developed and opt<strong>im</strong>ized againstexper<strong>im</strong>ental data.• With this force field, the interaction of IBC withlipid bilayers and proteins (acetylcholine esteraseand histones) will be s<strong>im</strong>ulated.Weakly coordinating ions (WCI), both cations andanions, are a broad class of substances known fortheir peculiar properties in solution. Among them,icosahedral ionic boron cluster compounds (IBC)of the type YB 12 X 12 (2-) (X = H, I, CH 3 and Y = SH,OH) and XB 12 H 11 (-) (X=H, NH 3 , N(CH 3 ) 3 ), see Figure1, are particularly interesting. They influencethe structure and tightness of lipid bilayers, andthey inhibit enzyme functions. The structure, size,and charge, of anions are known to affect their coordinatingproperties, and this is especially true forthe IBC. We do, however, not yet have an understandingof their structural, dynamics and thermodynamicsproperties in water and in other solvents.Furthermore, despite their long use in the designand synthesis of therapeutics for Boron NeutronCapture Therapy of cancer and, more recently, inother areas of drug design, the molecular mechanismof their interaction with the components ofliving cells (protein and biological membranes) islargely unknown both exper<strong>im</strong>entally and theoretically.The a<strong>im</strong> of this project is to study molecularproperties of these anions and the details of theirinteractions with biological systems, using moleculardynamics (MD) techniques. We will focus onthe interaction of IBC with two different systems.1. Ion-Membrane Interactions. It is known thatWCI interact with membranes. We have shownthat this is especially true for boron cluster anions[1],[2]. Membrane integrity is influenced, andmorphological changes (from minor to drastic) arefound. Understanding the interaction mechanismson a molecular level will greatly <strong>im</strong>prove predictivemodels and the description of toxicologic andpharmacologic effects of the anions. The interactionof boron clusters with membranes will be analyzedusing MD s<strong>im</strong>ulations. We first have to developa force field model for the IBC, mandatory forthe subsequent MD calculations. The parametersfor the force field (equilibrium geometries, forceconstants, partial charges) are derived from quantummechanics calculations, which provide a detaileddescription of the IBC and of the interactionwith solvent molecules. The new force field will bechecked against exper<strong>im</strong>ental data which we havegathered. In particular, thermodynamics properties,transport properties in water and in other solventswill be considered. Free energies will beest<strong>im</strong>ated using thermodynamics integration methods.Diffusion coefficients will be est<strong>im</strong>ated fromexper<strong>im</strong>ental conductivity measurements using theNernst-Einstein equation. The IBC model will beused to perform MD s<strong>im</strong>ulations in the presenceof a phosphatidylcholine lipid bilayer sheet. Thecalculations will provide information on free energybarriers for percolation, changes of lipid surfacearea, orientation of asymmetric molecules, stoichiometricratios of lipid to clusters, and inducedpore formation. Different types of lipid membranewill be used for these purposes. The free energybarrier of diffusion of anions through membraneswill then be calculated using equilibriumand non-equilibrium approaches such as umbrellasampling, thermodynamics integration and steeredmolecular dynamics (SMD). The potential of meanforce necessary to penetrate the lipid bilayer will becalculated using SMD s<strong>im</strong>ulations. SMD appliesan external force that lowers the energy barrier,and thus allows to study long t<strong>im</strong>e scale processeson the nanosecond t<strong>im</strong>escale. The trajectories obtainedfrom SMD will be used to obtain accuratefree energy barriers of starting conformation fromumbrella sampling or from constrained potential ofmean force calculations.2. Ion-Protein Interactions. The recent discoveryof another ICB, metallacarborane, as effectiveinhibitor of HIV-1 protease has demonstrated thatICB can interact strongly with proteins. Few modelingstudies have been performed, and none ofthem used MD s<strong>im</strong>ulations. In this project, twomodel proteins, acetylcholinesterase (AChE) andChemie
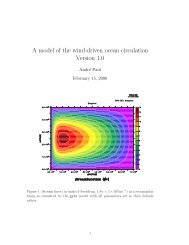
35Figure 1: Structure of the icosahedral ionic boron clusters used (top) and their interactions with membranes (bottomleft) and proteins (bottom center and right). The approach to SMD is shown in the bottom left: The ICPis pulled into the lipid bilayer by an externally applied field, represented by the spring. The center showsIBC ions interacting with AChE, and the right is an NCP ensemble.the nucleosome core particle (NCP), will be used.AChE is <strong>im</strong>portant for the transmission of electricsignals in the neuron cells and it is a target enzymefor the treatment of Alzhe<strong>im</strong>er disease. We foundthat AChE is inhibited by ICB [1], which is surprising,as the ICB are negatively charged and the naturalsubstrate is positively charged. NCP is themolecular building block of the chromatin presentin the nuclei of eukaryotic cells. We have found thatIBC accumulate in the nuclei of the cells, probablyon NCP. It likely that the anions interact preferentiallywith the positively charged histones. We haveperformed MD s<strong>im</strong>ulations of NCP [3], but dockingstudies with IBC have not yet been conductedon this system. We plan to investigate the bindingof IBC to AChE and with the histone proteinsin NCP by using MD s<strong>im</strong>ulations. MD will be usedto analyze preferential binding of the ICB to specificsites on the protein surface, in order to learnabout the specific propensity of the ions to bind tospecific amino acids. Subsequently, the diffusionof the boron cluster into the AchE active site willbe analyzed by calculating the free energy barrierof diffusion and free energy of binding. We expectto obtain information on binding constants andconformational changes, which can be comparedwith exper<strong>im</strong>ental data from enzyme inhibition andNMR spectroscopy.More Information1. Awad, D., L. Damian, M. Winterhalter, G. Gabel,D., D. Awad, T. Schaffran, D. Radovan, D. Daraban,L. Damian, M. Winterhalter, G. Karlsson,and K. Edwards. 2007. The Anionic BoronCluster (B 12 H 11 SH(2-) as a Means To TriggerRelease of Liposome Contents. ChemMed-Chem 2:51-53.2. Schaffran, T., E. Justus, M. Elfert, T. Chen, andD. Gabel. 2009. Toxicity of N,N,N- trialkylammoniododecaboratesas new anions of ionic liquidsin cellular, liposomal and enzymatic testsystems. Green Chem 11:1458-1464.3. Roccatano D., A. Barthel, M. Zacharias. 2007.Structural flexibility of the nucleosome core particleat atomic resolution studied by moleculardynamics s<strong>im</strong>ulations. Biopolymers, 85, 401-421.FundingUniversity of BremenChemie